External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G14300 148 / 4e-41
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G33410 146 / 2e-40
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G58470 143 / 1e-39
XF41, ATRBP1
RNA-binding protein 1 (.1)
AT4G26650 143 / 6e-39
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT5G55550 136 / 1e-36
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT3G07810 136 / 4e-36
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT5G47620 130 / 2e-34
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT5G40490 127 / 4e-33
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G13224 119 / 9e-31
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT1G17640 118 / 3e-30
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011560
439 / 3e-155
AT4G14300 94 / 5e-28
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10011728
204 / 6e-64
AT5G55550 142 / 3e-39
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
Lus10043124
144 / 1e-39
AT2G33410 334 / 2e-111
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10032624
142 / 9e-39
AT2G33410 335 / 2e-112
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10043168
140 / 1e-37
AT3G07810 456 / 2e-157
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10032579
138 / 5e-37
AT3G07810 447 / 8e-154
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10000537
137 / 2e-36
AT3G07810 420 / 8e-143
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10022889
134 / 3e-36
AT4G14300 315 / 9e-105
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10039054
134 / 8e-36
AT3G07810 428 / 4e-147
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G170400
144 / 2e-39
AT4G14300 317 / 2e-104
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.006G015700
141 / 3e-38
AT3G07810 447 / 8e-154
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.010G067500
140 / 7e-38
AT4G14300 302 / 7e-99
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.002G258600
139 / 1e-37
AT4G14300 277 / 1e-89
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.014G161600
139 / 3e-37
AT3G07810 498 / 3e-174
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.014G195300
136 / 1e-36
AT4G14300 305 / 2e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.002G222400
135 / 5e-36
AT3G07810 439 / 8e-151
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.002G115000
132 / 5e-36
AT1G58470 219 / 7e-69
RNA-binding protein 1 (.1)
Potri.016G010200
134 / 1e-35
AT3G07810 413 / 2e-140
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.001G361300
134 / 2e-35
AT3G07810 433 / 1e-148
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Lus10019276 pacid=23141427 polypeptide=Lus10019276 locus=Lus10019276.g ID=Lus10019276.BGIv1.0 annot-version=v1.0
ATGGAAGGAGGAGGAGGAGTACAAGAAACCATTCAGCAGCTACCTCGCCCAAGGCTGTTCGTAGGCGGACTGAAAATGGAGGCCACGGGGTTATTTCTCC
AGGAGCACTTTAGGGAGTATGGCACATTGAAAGATTGTTTCATCTGTTACAACAAGGAAACTGGACTAAGCCGTACTTTCGGTTTCGTGGAATTCGAAGA
CCCCGCTGTTGCTCAGCGTGTTCTTCAAGATCGGCATTTCATTCTAGGCCAAAAGGCCGATGTGTCCGTAGCTAAAGAGAGGAACCTCCACAATCGCAAT
GACAGCAAGAAGATCTTCGTCGGTGGCTTACCGCCAGCAGTTACACAGCAGGAGCTGAGGAGTTTCTTCGAGAGGTTCGGGACAGTGACTGATGCTGTCC
TGGTTCACGATAGGATTACCAGAGCACTTCGAGGGTTTGGATTCGTAACCTTCGAATCCAAAGACTCGGCAGATGCAGCTCTGCAGTCCAGATATCACGA
ATTTAAAGGCAGGCGGGTGGAGGTGAAGAGAGCAAATAATCCTATAGTCAGGAACAGTACTACTGTTGGTAACAATGTTGGGTACTATGATTTATCAGTC
GACCAGACCGGTTTCTTCGGTTATGGGTCTCCAGATTGTTATTACTGGTACCATCATTGTTTCTGGAACATGAACTACCCATATTATGAATTTCCTGCAA
CAAATTACTATGCTGCTGGCAGCTGGTACAGTGGCTATGGCTATGCTGAGGGTGGGATTGCTGGGGATGAAATCGGGACCGCTGGATGTCAGATTGCTGA
TCATCACGGTACTCAGCATGATACAGAGGATTCAGAAATCGATATCGATTCGTTCGACATGAAGAACTATCTTCTGGATCAAGAAGATGAAGCAGATGGG
TCAGAATCCGACATTGAAAGAAGTCAGCCTAAGTTGATTAAAGGCAAGTCCTTTATTGCAGCCCCACAATGA
AA sequence
>Lus10019276 pacid=23141427 polypeptide=Lus10019276 locus=Lus10019276.g ID=Lus10019276.BGIv1.0 annot-version=v1.0
MEGGGGVQETIQQLPRPRLFVGGLKMEATGLFLQEHFREYGTLKDCFICYNKETGLSRTFGFVEFEDPAVAQRVLQDRHFILGQKADVSVAKERNLHNRN
DSKKIFVGGLPPAVTQQELRSFFERFGTVTDAVLVHDRITRALRGFGFVTFESKDSADAALQSRYHEFKGRRVEVKRANNPIVRNSTTVGNNVGYYDLSV
DQTGFFGYGSPDCYYWYHHCFWNMNYPYYEFPATNYYAAGSWYSGYGYAEGGIAGDEIGTAGCQIADHHGTQHDTEDSEIDIDSFDMKNYLLDQEDEADG
SESDIERSQPKLIKGKSFIAAPQ
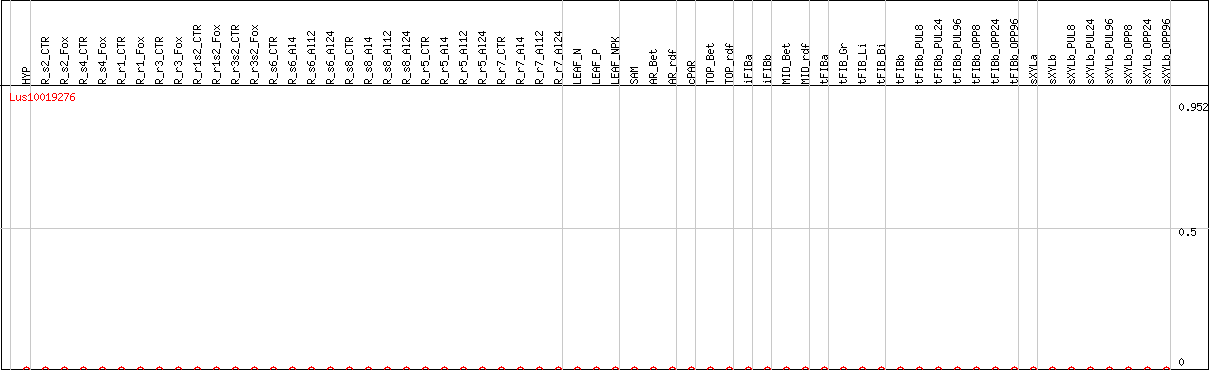
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019276 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.