Lus10019336 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10019336 pacid=23141317 polypeptide=Lus10019336 locus=Lus10019336.g ID=Lus10019336.BGIv1.0 annot-version=v1.0
ATGGCGTCGAAGGCGTCTCCGTCGGCTTCCGCCAGTAGATCGGGGTCTCTCAAGGACACTTCGATGGAAAAAGCGGAAGAGTCAGGTCCAACGAGTGAGG
AGATTGGATCAATTATCGAATCGCTGAAAACGCTTGTGGCGGATGAACGGTTTGTTTCTATAAAGGAAAGGATGAAAGAAAATAGAAACAAACTGGCTAG
TGTCACCAGCCATCTACAGAAACTGTCCATGGAAAGAAGAAACAGCTGGATTAACATTGGCGAAGCTAGTCAAGATCTGCTGAAGAAGAGGCAAGATGAA
GCAGTCGGGATGCATAGTGGGTTTGGTATTAGTAATGGTGATAGAGATAGTGACAGCTATCAAGAAGACTCTCATGGTTCTACGGCTGTTCTTCTGGGAA
CTAGTATTCCTCCAGGGAAGAATTCCGTTAGGCCGATTAAGCTTCCAGGAGTGAAGAAGTTGCCTCCTTATACCACCTGGATATTTCTGGACAGAAATCA
GAGGATGACAGAAGATCAATCAGTGGTGGGCCGGAGAAGGATTTATTATGATAGTGGTGAAGCAGTTATTTGTAGTGACAGTGAGGAGGAAGCTGTTGAT
GAAGAAGAGGAAAAAAGATTTTTTCTGTCGTCAGAAGATTATATTATACGCATGACCCTAGATAAAACTGGATTATCAAATGGTGCGCTCCAATCCTTGG
CGGAATGTCTGTCCAGAAGTAAAGATGAAGTGAAGGCAAGATATGAAAGTCTAACCAAGGAAGAGAAACCTTCAGTGGATTCCATGGATGTGGGCAGTGA
AACACAAACAGTGAATTCTATTCTTGATAAGGATCTGGAAGCTGCTTTGGATTCTTTTGACAACTTATTTTGCCGGCGTTGTCTTGTCTTCGATTGCAGA
TTGCACGGATGCTCACAGGATCTCGTATTTCCAGTTGAGAAACCACTTCCATGGAACCATCCAGACATAGAAAATGTTCCATGTGGTTCCCATTGCTACA
AATCGGTGTCGAAGTCAGGGAAATTGGTTACTTTAAACTCGCCTGAGCAGCTTAGTGATACAAAAGAAAACTCTCTTCAGCCATCTGGCACTGCTGGGAT
TCGCATGTCTGTGAGGAATAAATCTACTTCATCTACTCGAAAGAAAGTCAAGCCCTCCCAAAGTGAAAGTGCCTCTTCAAATGTCAAAAACATGTCAGAG
AGTAGTGACTCAGAGATTGGACCCCAACATGATACTTCTTCCACTGGCCATTTGTCACCCTCCAAGAATAATATTATTGGAAAGGGTGGCATTTCTCACA
GGAACAGCAAGCGAGTGGCTGAACGTGTTCTGAGTTGCATAAAGAAAAGGCAGAAGAAGTTGATAGGATCTGATTCCGATTCTGTCGCTAGTGGAGGTCT
TTTGGGAGGTGATGCAACCAGTCCACATAAGGAAGATGAAGATGCCACTTCTTCTTCCAAAAAGGCAAGAAAATCTTCAAACACTGGGAGGTCTAGGAAA
ACAGAGTTTTCAGCTCATCAAGTGCAAGGCGAAGTATCAAGCCAGATGTGCACTGATCCACCTGGAAGTAGTAGTGATGACACCTTAAGGAAAGATGACC
ATGTTGATGGAAACAAATGTAATCTGGAGTCGAATCATAACCTGTCTTGGAAACCCTTTGAAAAAAGTATGTTTGAGAAAGGTGTGGAAATTTTTGGCTG
GAATAGCTGTCTAATTACTAGGAATCTATTAAATGGTTTAAAGACGTGCACGGAAGTTTATCATTACATGACACGAGGAGACAACAAACTAGGCTGCCAA
GCAGGTGATGCTGCAAATTCTCTTGGCGAAGGCTATTCCAAGTTTGAACATCAAGGCAATAATGAAGTTAGGAGAAGATCAAGGTTTTTACGCCGAAGAG
GTAAAGTTCGCCGTTTGAAGTATTCATGGAAGTCTACAGCTTACCATTCTATCCGAAAACGGATAAGTGACAAAAAAGATCAACCGTGCCGCCAGTACAA
TCCTTGTGGCTGCCAAGCAGCTTGTGGAAAGCAGTGCGCTTGCTTGCTGAATGGCACTTGCTGTGAGAAGTATTGTGGTTGTGGCGATGGCTCACTTGGT
GTTCCGAATCAAAGCGGAGACAATTACGAGTGCCGGAATATGAAACTTCTTCTGAAACAACAGCAAAGGGTTTTACTTGGAAGATCCAATATATCTGGCT
GGGGAGCATTTCTGAAGAATAGCGTCGGTAAGCATGATTACCTCGGCGAATACACTGGAGAGTTGATATCACATAGGGAAGCAGACAAGCGTGGCAAGAT
TTACGACCGCGAAAACTCGTCATTCCTCTTCAATTTGAATGATCAGGCATATGGTCTTTCATCCCAACTTTTATTCGTCCTTGATGCTTACCGGAAGGGC
GACAAGTTGAAGTTTGCCAACCATTCCCCTGATCCAAACTGCTACGCCAAGGTGATCATGGTTGCAGGAGATCACAGAGTGGGGATATTCGCCAAAGAAA
GAATTGGTGCTGGCGAGGAGCTTTTCTACGATTACCGTTACGAACCAGACAGAGCTCCTGCCTGGGCAAGGAAGCCCGAAGCATCCGGGTCGAAAAGGGA
AGAGACGGGTCATACAAGTGGCCGAGCCAAGAAGCTGGCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10019336 pacid=23141317 polypeptide=Lus10019336 locus=Lus10019336.g ID=Lus10019336.BGIv1.0 annot-version=v1.0
MASKASPSASASRSGSLKDTSMEKAEESGPTSEEIGSIIESLKTLVADERFVSIKERMKENRNKLASVTSHLQKLSMERRNSWINIGEASQDLLKKRQDE
AVGMHSGFGISNGDRDSDSYQEDSHGSTAVLLGTSIPPGKNSVRPIKLPGVKKLPPYTTWIFLDRNQRMTEDQSVVGRRRIYYDSGEAVICSDSEEEAVD
EEEEKRFFLSSEDYIIRMTLDKTGLSNGALQSLAECLSRSKDEVKARYESLTKEEKPSVDSMDVGSETQTVNSILDKDLEAALDSFDNLFCRRCLVFDCR
LHGCSQDLVFPVEKPLPWNHPDIENVPCGSHCYKSVSKSGKLVTLNSPEQLSDTKENSLQPSGTAGIRMSVRNKSTSSTRKKVKPSQSESASSNVKNMSE
SSDSEIGPQHDTSSTGHLSPSKNNIIGKGGISHRNSKRVAERVLSCIKKRQKKLIGSDSDSVASGGLLGGDATSPHKEDEDATSSSKKARKSSNTGRSRK
TEFSAHQVQGEVSSQMCTDPPGSSSDDTLRKDDHVDGNKCNLESNHNLSWKPFEKSMFEKGVEIFGWNSCLITRNLLNGLKTCTEVYHYMTRGDNKLGCQ
AGDAANSLGEGYSKFEHQGNNEVRRRSRFLRRRGKVRRLKYSWKSTAYHSIRKRISDKKDQPCRQYNPCGCQAACGKQCACLLNGTCCEKYCGCGDGSLG
VPNQSGDNYECRNMKLLLKQQQRVLLGRSNISGWGAFLKNSVGKHDYLGEYTGELISHREADKRGKIYDRENSSFLFNLNDQAYGLSSQLLFVLDAYRKG
DKLKFANHSPDPNCYAKVIMVAGDHRVGIFAKERIGAGEELFYDYRYEPDRAPAWARKPEASGSKREETGHTSGRAKKLA
|
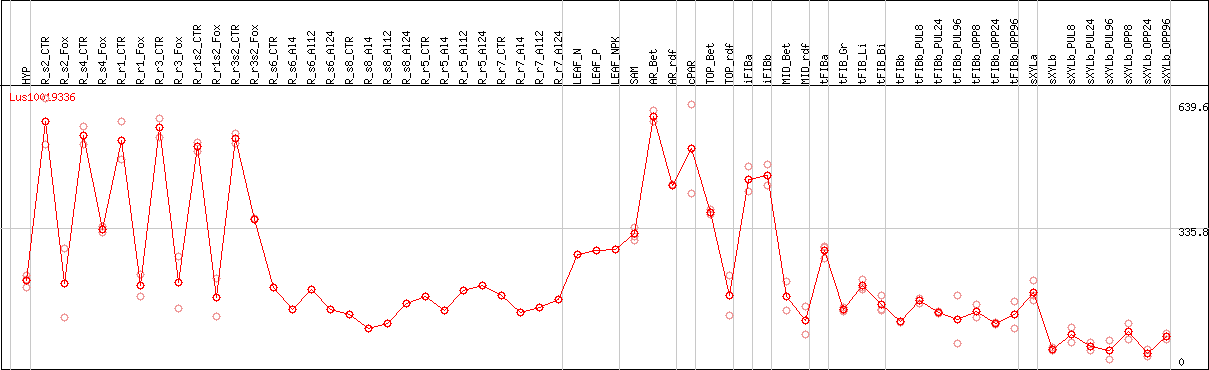
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019336 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
