External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G55190 406 / 3e-146
RAN3, ATRAN3
RAN GTPase 3 (.1)
AT5G20010 400 / 6e-144
ATRAN1, RAN-1
ARABIDOPSIS THALIANA RAS-RELATED NUCLEAR PROTEIN, RAS-related nuclear protein-1 (.1)
AT5G20020 390 / 1e-139
RAN2
RAS-related GTP-binding nuclear protein 2 (.1)
AT5G55080 293 / 2e-101
ATRAN4
ras-related nuclear protein 4 (.1)
AT4G39890 102 / 1e-26
AtRABH1c
RAB GTPase homolog H1C (.1)
AT1G07410 99 / 3e-25
ATRAB-A2B, AtRABA2b
ARABIDOPSIS RAB GTPASE HOMOLOG A2B, RAB GTPase homolog A2B (.1)
AT1G22740 98 / 3e-25
RAB75, RAB7, AtRABG3b
RAB GTPase homolog G3B (.1)
AT2G44610 97 / 8e-25
RAB6, AtRABH1b, AtRab6A
Ras-related small GTP-binding family protein (.1)
AT4G19640 97 / 9e-25
ATRAB-F2B, ARA7, Ara-7, AtRABF2b, AtRab5B
ARABIDOPSIS RAB GTPASE HOMOLOG F2B, Ras-related small GTP-binding family protein (.1)
AT5G45750 97 / 1e-24
AtRABA1c
RAB GTPase homolog A1C (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G250300
407 / 2e-146
AT5G55190 424 / 4e-153
RAN GTPase 3 (.1)
Potri.018G030900
404 / 2e-145
AT5G55190 424 / 3e-153
RAN GTPase 3 (.1)
Potri.018G030700
404 / 2e-145
AT5G55190 424 / 3e-153
RAN GTPase 3 (.1)
Potri.006G250400
391 / 3e-140
AT5G55190 411 / 3e-148
RAN GTPase 3 (.1)
Potri.001G152800
100 / 9e-26
AT1G02130 382 / 2e-137
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.003G086700
100 / 9e-26
AT2G44610 397 / 4e-143
Ras-related small GTP-binding family protein (.1)
Potri.002G135500
99 / 2e-25
AT2G44610 385 / 3e-138
Ras-related small GTP-binding family protein (.1)
Potri.001G080400
98 / 4e-25
AT1G02130 387 / 2e-139
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.018G079300
98 / 4e-25
AT5G45130 273 / 2e-94
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
Potri.003G081800
97 / 6e-25
AT1G02130 391 / 9e-141
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00071
Ras
Ras family
Representative CDS sequence
>Lus10019441 pacid=23181571 polypeptide=Lus10019441 locus=Lus10019441.g ID=Lus10019441.BGIv1.0 annot-version=v1.0
ATGGCTCTCCCGAATCAGCAAGTAGTTGACTACCCGAGCTTCAAGCTCGTTATTGTTGGCGATGGAGGCACCGGGAAAACCACCTTCGTCAAGAGACATT
TGACTGGAGAGTTTGAGAAGAAATATGAACCAACCATTGGCGTGGAAGTCCATCCATTGGATTTCACTACCAACTGTGGAAAGATCCGATTCTATTGCTG
GGACACAGCTGGACAAGAGAAATTTGGCGGACTCAGGGATGGATACTACATCCATGGTCAATGTGCGATTATAATGTTTGATGTCACCGCAAGATTGACC
TACAAGAATGTTCCTACATGGCATCGTGATCTCTGCAGGGTTTGTGAGAATATACCCATTGTTCTCTGTGGAAACAAGGTTGATGTAAAGAACAGGCAGG
TTAAGGCCGATGTGAAGAACAGGCAGGTTAAGGCCAAGCAGGTTACTTTCCACAGGAAGAAGAATCTCCAGTACTATGAGATATCTGCCAAGAGCAACTA
CAACTTTGAGAAGCCTTTCCTCTACCTTGCTCGCAAGCTTGCTGGGGACCCCAATCTTCACTTTGTGGAGGCGGTTGCCCTCCGTCCTCCAGAGGTTCAA
ATTGACATGGCTACACAGCAACAACATGAACAAGAGCTTGCTCAAGCTGCTAACCAGCCTCTTCCTGATGACGATGATGATGCATTCGAGTAA
AA sequence
>Lus10019441 pacid=23181571 polypeptide=Lus10019441 locus=Lus10019441.g ID=Lus10019441.BGIv1.0 annot-version=v1.0
MALPNQQVVDYPSFKLVIVGDGGTGKTTFVKRHLTGEFEKKYEPTIGVEVHPLDFTTNCGKIRFYCWDTAGQEKFGGLRDGYYIHGQCAIIMFDVTARLT
YKNVPTWHRDLCRVCENIPIVLCGNKVDVKNRQVKADVKNRQVKAKQVTFHRKKNLQYYEISAKSNYNFEKPFLYLARKLAGDPNLHFVEAVALRPPEVQ
IDMATQQQHEQELAQAANQPLPDDDDDAFE
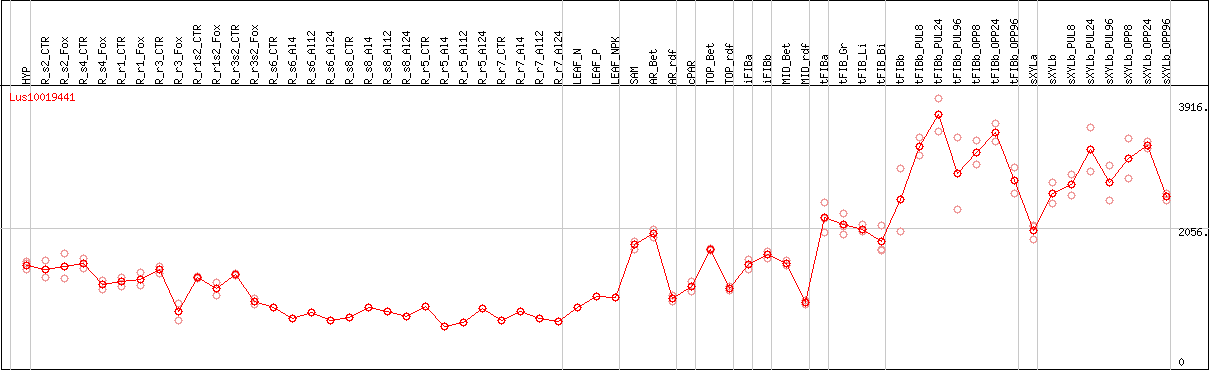
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019441 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.