External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G54890 318 / 4e-109
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
AT4G31010 228 / 3e-73
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1), RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.2)
AT2G20020 207 / 2e-62
CAF1, ATCAF1
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
AT1G23400 194 / 8e-59
CAF2, ATCAF2
ARABIDOPSIS THALIANA HOMOLOG OF MAIZE CAF2, RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
AT3G01370 54 / 2e-08
ATCFM2
Arabidopsis thaliana CRM family member 2, CRM family member 2 (.1)
AT3G23070 51 / 3e-07
ATCFM3A
CRM family member 3A (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040080
364 / 9e-127
AT5G54890 484 / 2e-172
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Lus10036545
221 / 1e-70
AT4G31010 446 / 8e-157
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1), RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.2)
Lus10041383
216 / 1e-68
AT4G31010 526 / 0.0
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1), RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.2)
Lus10012974
207 / 8e-63
AT2G20020 544 / 0.0
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Lus10011119
204 / 8e-62
AT2G20020 598 / 0.0
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Lus10043237
204 / 9e-62
AT2G20020 605 / 0.0
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Lus10029134
196 / 3e-59
AT1G23400 712 / 0.0
ARABIDOPSIS THALIANA HOMOLOG OF MAIZE CAF2, RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Lus10013029
188 / 2e-55
AT1G23400 658 / 0.0
ARABIDOPSIS THALIANA HOMOLOG OF MAIZE CAF2, RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Lus10034952
150 / 4e-43
AT2G20020 274 / 2e-85
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G423800
336 / 4e-116
AT5G54890 514 / 0.0
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Potri.014G074400
231 / 2e-74
AT4G31010 501 / 1e-177
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1), RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.2)
Potri.006G163100
208 / 8e-63
AT2G20020 644 / 0.0
RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Potri.010G042500
198 / 3e-60
AT1G23400 655 / 0.0
ARABIDOPSIS THALIANA HOMOLOG OF MAIZE CAF2, RNA-binding CRS1 / YhbY (CRM) domain-containing protein (.1)
Potri.010G077100
57 / 3e-09
AT3G23070 927 / 0.0
CRM family member 3A (.1)
Potri.012G056100
54 / 3e-08
AT3G18390 935 / 0.0
embryo defective 1865, CRS1 / YhbY (CRM) domain-containing protein (.1)
Potri.001G350801
53 / 7e-08
AT3G01370 972 / 0.0
Arabidopsis thaliana CRM family member 2, CRM family member 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01985
CRS1_YhbY
CRS1 / YhbY (CRM) domain
Representative CDS sequence
>Lus10019623 pacid=23142558 polypeptide=Lus10019623 locus=Lus10019623.g ID=Lus10019623.BGIv1.0 annot-version=v1.0
ATGATACGTTGGTCACCCACTCATGGATTGCGCAATCCCCCAAGGTATAATCCGGCTCAGGCCAGACATGTTGAGTATGCCTACAGATGCTGCTCACACG
GGAAAGGAGGAGTGACTCATAATATGTTGGATGATATCCACAACCATTGGAAACGAGCTGAAGCCGTCAGAATCAAGTGCTTGGAGGTTCCAACTCTAGA
TATGGACAATGTCTGCTTCCACCTTGAGGACAAATCTGGTCGGAAGATTATATACCGGCACATCAACGTCCTTCTTTTGTATCGGGGGAGAAACTATGAT
CCCAAGAATCGGCCTCGTATTCCCTTGATGCTATGGAAGCCTTATCCACCTATTTATCCAAGGCTTGCCAAGAATGTTGCTGATGGCTTAACATTTGAAG
AAACCAAGGAAATGAGGAACAGAGAACTCAATTCTCTTCCGCTCATGAAGCTTCCCAGAAATGGTGTTTATGTGAATGTAGTGGATAGGGTGAGTGAGGC
TTTCACTACAAAAGAGGTTGTGAGACTTGATTGCACGCATGTCGGTACCAGTGACTGCAAAAGGATTGGTGTGAAGCTAAGGGATTTAGTGCCTTGTGTT
CCTATTTTGGTCAAGGACGAGCAGGTTATACTTTGGAGGGGGAGGGTTGACAAGGGTCAGCAGCATGATTCTTCAGCTATTCCAGCTGCAGACCCTTCCT
CAGATGATTAA
AA sequence
>Lus10019623 pacid=23142558 polypeptide=Lus10019623 locus=Lus10019623.g ID=Lus10019623.BGIv1.0 annot-version=v1.0
MIRWSPTHGLRNPPRYNPAQARHVEYAYRCCSHGKGGVTHNMLDDIHNHWKRAEAVRIKCLEVPTLDMDNVCFHLEDKSGRKIIYRHINVLLLYRGRNYD
PKNRPRIPLMLWKPYPPIYPRLAKNVADGLTFEETKEMRNRELNSLPLMKLPRNGVYVNVVDRVSEAFTTKEVVRLDCTHVGTSDCKRIGVKLRDLVPCV
PILVKDEQVILWRGRVDKGQQHDSSAIPAADPSSDD
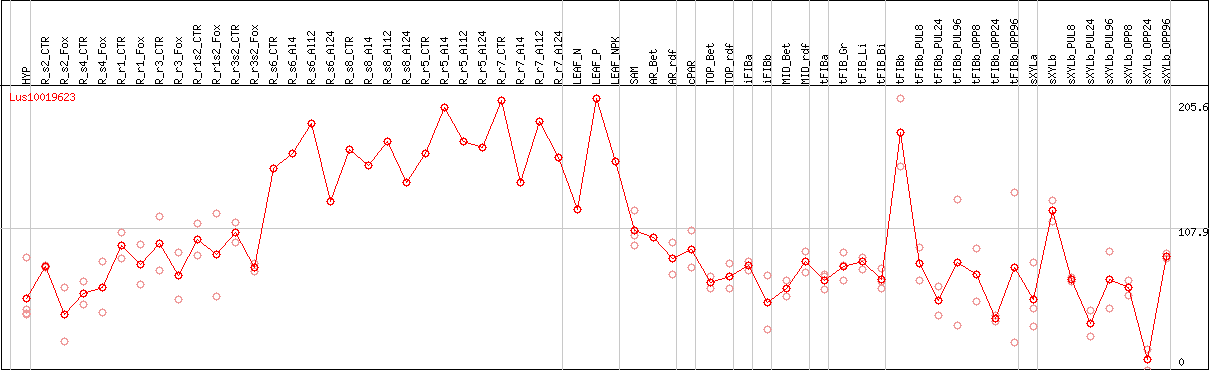
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019623 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.