External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39770 474 / 5e-169
TPPH
trehalose-6-phosphate phosphatase H, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT2G22190 464 / 4e-165
TPPE
trehalose-6-phosphate phosphatase E, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT5G10100 463 / 2e-164
TPPI
trehalose-6-phosphate phosphatase I, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
AT5G65140 461 / 2e-163
TPPJ
trehalose-6-phosphate phosphatase J, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
AT1G35910 428 / 1e-150
TPPD
trehalose-6-phosphate phosphatase D, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT1G78090 408 / 1e-142
ATTPPB
Arabidopsis thaliana trehalose-6-phosphate phosphatase B, trehalose-6-phosphate phosphatase (.1)
AT5G51460 377 / 3e-130
ATTPPA
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
AT1G22210 373 / 1e-129
TPPC
trehalose-6-phosphate phosphatase C, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT4G22590 347 / 1e-118
TPPG
trehalose-6-phosphate phosphatase G, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT4G12430 343 / 3e-117
TPPF
trehalose-6-phosphate phosphatase F, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000687
544 / 0
AT5G65140 451 / 3e-159
trehalose-6-phosphate phosphatase J, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Lus10027208
391 / 6e-136
AT5G51460 515 / 0.0
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
Lus10031146
381 / 5e-132
AT5G51460 517 / 0.0
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
Lus10031723
372 / 2e-128
AT5G51460 511 / 0.0
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
Lus10024607
371 / 7e-128
AT4G22590 534 / 0.0
trehalose-6-phosphate phosphatase G, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10032244
369 / 6e-127
AT4G22590 531 / 0.0
trehalose-6-phosphate phosphatase G, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10038921
370 / 8e-127
AT5G51460 488 / 2e-172
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
Lus10019010
315 / 2e-106
AT1G78090 338 / 6e-115
Arabidopsis thaliana trehalose-6-phosphate phosphatase B, trehalose-6-phosphate phosphatase (.1)
Lus10039339
313 / 1e-105
AT1G78090 339 / 2e-115
Arabidopsis thaliana trehalose-6-phosphate phosphatase B, trehalose-6-phosphate phosphatase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G077200
520 / 0
AT5G65140 545 / 0.0
trehalose-6-phosphate phosphatase J, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.007G090900
520 / 0
AT5G65140 548 / 0.0
trehalose-6-phosphate phosphatase J, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.002G094500
469 / 8e-167
AT5G65140 489 / 3e-174
trehalose-6-phosphate phosphatase J, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.012G126100
374 / 5e-129
AT5G51460 526 / 0.0
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
Potri.015G126900
372 / 2e-128
AT5G51460 531 / 0.0
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.3)
Potri.001G120500
365 / 1e-125
AT4G22590 501 / 2e-178
trehalose-6-phosphate phosphatase G, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.003G112400
361 / 4e-124
AT4G12430 485 / 3e-172
trehalose-6-phosphate phosphatase F, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.012G001000
346 / 2e-118
AT1G35910 336 / 4e-114
trehalose-6-phosphate phosphatase D, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.015G020300
344 / 1e-117
AT1G78090 333 / 4e-113
Arabidopsis thaliana trehalose-6-phosphate phosphatase B, trehalose-6-phosphate phosphatase (.1)
Potri.005G166700
316 / 2e-107
AT5G65140 362 / 8e-126
trehalose-6-phosphate phosphatase J, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0137
HAD
PF02358
Trehalose_PPase
Trehalose-phosphatase
Representative CDS sequence
>Lus10019666 pacid=23139231 polypeptide=Lus10019666 locus=Lus10019666.g ID=Lus10019666.BGIv1.0 annot-version=v1.0
ATGACGAAGAAGAAGCTTCTGCTGGAGAATCTGGAAATCAACGGTGGTGGCGGAGGAGCGAGGATCAATTCGTGGGTAGATTCGATGAGGGCTTCGTCCC
CGACCCGTCTGAGATCGAATTCGAAATCGGAAGAGCAGCAGCAGGGGTTTCAGTGGCAGGAGGAGCAGTTTCTTCATCTTCCGTCGGCTCTGGATTCTTT
CGATGAGATTATCGAAGCTTCGCAGGGGAAGCAAATCGTCATGTTTCTGGACTACGACGGCACTCTTTCTCCCATTGTCGACGATCCTGATCGAGCTTTC
ATGTCCAAGAAGATGAGAACTACGGTTAGAAGACTCGCTAGATGCTTTCCTACTGCCATAGTTAGTGGAAGATGCAGAGACAAGGTGTACAACTTTGTAA
GGTTAGCGGAGCTTTACTATGCCGGAAGCCACGGAATGGACATCAAGGGGCCGAAGAAAGGGTCAAAGTACACGAAAGGAGTCGAGCCAGTGATCTGTCA
GCCGGCAAGCGAGTTCCTCCCCATGATCGACGAAGTATTCAAAGAGCTGATCGAGAAAACCAAATCGACCCCTGGAGCGAAAGTTGAGAACAACAAATTT
TGCGTTTCGGTCCATTTCCGGTGTGTCGAGGAGAAGATGTGGGGCGAGGTGGCGGAAGCCGTTAAGTCGGTTTTGAAAAAGTACCCCAAACTTAGACTCT
CGCACGGACGAAAGGTGTTGGAAATCCGACCAACCATCAAGTGGGACAAAGGAAAGGCTCTTGAATTTTTGTTGGAGTCTTTGGGATTTGCCAACTGTAC
CGATGTTTTTCCTATTTATATCGGAGATGATCGAACAGACGAGGATGCATTTAAAGTATTAAAAAAAAGAGGGCAAGGATTTGGGATTCTCGTGTCCAAA
GTTCCAAAAGAAACCAATGCTTCTTATGCCCTACAAGAACCCACTGAGGTTATGGACTTCTTGCAAAGATTGGTCGATTGGAAAAGGCGGGATTTCCAAA
GGCAATCCAGGATGTAG
AA sequence
>Lus10019666 pacid=23139231 polypeptide=Lus10019666 locus=Lus10019666.g ID=Lus10019666.BGIv1.0 annot-version=v1.0
MTKKKLLLENLEINGGGGGARINSWVDSMRASSPTRLRSNSKSEEQQQGFQWQEEQFLHLPSALDSFDEIIEASQGKQIVMFLDYDGTLSPIVDDPDRAF
MSKKMRTTVRRLARCFPTAIVSGRCRDKVYNFVRLAELYYAGSHGMDIKGPKKGSKYTKGVEPVICQPASEFLPMIDEVFKELIEKTKSTPGAKVENNKF
CVSVHFRCVEEKMWGEVAEAVKSVLKKYPKLRLSHGRKVLEIRPTIKWDKGKALEFLLESLGFANCTDVFPIYIGDDRTDEDAFKVLKKRGQGFGILVSK
VPKETNASYALQEPTEVMDFLQRLVDWKRRDFQRQSRM
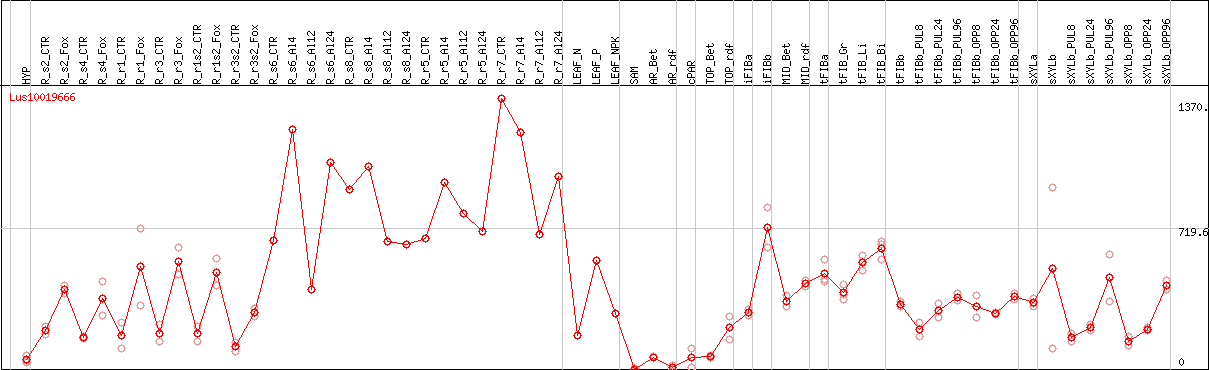
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019666 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.