External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39700 211 / 3e-71
Heavy metal transport/detoxification superfamily protein (.1)
AT1G22990 184 / 1e-60
HIPP22
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
AT1G71050 178 / 3e-58
HIPP20
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
AT4G08570 171 / 2e-55
Heavy metal transport/detoxification superfamily protein (.1)
AT4G38580 167 / 5e-54
HIPP26, ATFP6
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
AT5G66110 152 / 5e-48
HIPP27
heavy metal associated isoprenylated plant protein 27, Heavy metal transport/detoxification superfamily protein (.1)
AT5G17450 149 / 1e-46
HIPP21
heavy metal associated isoprenylated plant protein 21, Heavy metal transport/detoxification superfamily protein (.1.2)
AT4G35060 141 / 1e-43
HIPP25
heavy metal associated isoprenylated plant protein 25, Heavy metal transport/detoxification superfamily protein (.1)
AT2G18196 107 / 6e-30
Heavy metal transport/detoxification superfamily protein (.1)
AT1G06330 101 / 5e-28
Heavy metal transport/detoxification superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016436
320 / 2e-114
AT4G39700 209 / 1e-70
Heavy metal transport/detoxification superfamily protein (.1)
Lus10022508
245 / 1e-84
AT4G39700 201 / 2e-67
Heavy metal transport/detoxification superfamily protein (.1)
Lus10016812
225 / 1e-76
AT4G39700 181 / 3e-59
Heavy metal transport/detoxification superfamily protein (.1)
Lus10039419
190 / 1e-62
AT4G08570 215 / 5e-73
Heavy metal transport/detoxification superfamily protein (.1)
Lus10042946
185 / 7e-61
AT1G71050 196 / 3e-65
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Lus10016708
183 / 5e-60
AT1G71050 192 / 1e-63
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Lus10036004
182 / 1e-59
AT1G22990 187 / 4e-62
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Lus10019789
173 / 3e-56
AT4G38580 254 / 4e-88
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
Lus10014120
172 / 5e-56
AT4G38580 253 / 5e-88
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G087300
244 / 2e-84
AT4G39700 215 / 9e-73
Heavy metal transport/detoxification superfamily protein (.1)
Potri.005G079800
232 / 1e-79
AT4G39700 189 / 2e-62
Heavy metal transport/detoxification superfamily protein (.1)
Potri.005G167000
194 / 1e-64
AT4G08570 229 / 1e-78
Heavy metal transport/detoxification superfamily protein (.1)
Potri.002G092200
192 / 8e-64
AT4G08570 193 / 3e-64
Heavy metal transport/detoxification superfamily protein (.1)
Potri.010G114600
192 / 1e-63
AT1G22990 194 / 9e-65
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Potri.005G003700
181 / 2e-59
AT4G08570 207 / 5e-70
Heavy metal transport/detoxification superfamily protein (.1)
Potri.017G123400
170 / 4e-55
AT5G17450 220 / 4e-75
heavy metal associated isoprenylated plant protein 21, Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.009G135300
167 / 1e-53
AT4G38580 234 / 3e-80
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
Potri.004G175400
165 / 6e-53
AT4G38580 254 / 2e-88
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
Potri.005G110400
151 / 2e-47
AT5G66110 189 / 2e-62
heavy metal associated isoprenylated plant protein 27, Heavy metal transport/detoxification superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00403
HMA
Heavy-metal-associated domain
Representative CDS sequence
>Lus10019676 pacid=23139208 polypeptide=Lus10019676 locus=Lus10019676.g ID=Lus10019676.BGIv1.0 annot-version=v1.0
ATGGGGATTGCAGGAACTTTGGAGTACTTGTCTGAGTTGATGGCAAGCTCTGGCCATAAACACAAGAAGAGGAAGCAATTGCAGACTGTGGAGCTCAAGG
TCAGGATGGATTGTGATGGTTGTGAGCTCAAGGTCAAGAATTCACTTTCCTCTCTAAGTGGGGTGAAGAGTGTGGAGATAAACAGGAAGCAACAGAAGGT
GACAGTAACCGGGTTCGTCGAGCCACACAAAGTGCTGAAGAAGGCGAAATCGACAGGGAAAAGAGCGGAGATTTGGCCATACGTTCCATATAACTTGGTG
GCTCAGCCATACATTGCTGGAACTTATGACAAGAAAGCACCTCCTGGCTATGTCAGGAAAGCTGAGCAAACCGGTGTTTCTAGCGGAACCTTAGTTATAA
ACAGAGGAGGAGACCCTTACACTTCCATGTTCAGTGATGACAACCCTAATGCTTGCTCCGTCATGTAA
AA sequence
>Lus10019676 pacid=23139208 polypeptide=Lus10019676 locus=Lus10019676.g ID=Lus10019676.BGIv1.0 annot-version=v1.0
MGIAGTLEYLSELMASSGHKHKKRKQLQTVELKVRMDCDGCELKVKNSLSSLSGVKSVEINRKQQKVTVTGFVEPHKVLKKAKSTGKRAEIWPYVPYNLV
AQPYIAGTYDKKAPPGYVRKAEQTGVSSGTLVINRGGDPYTSMFSDDNPNACSVM
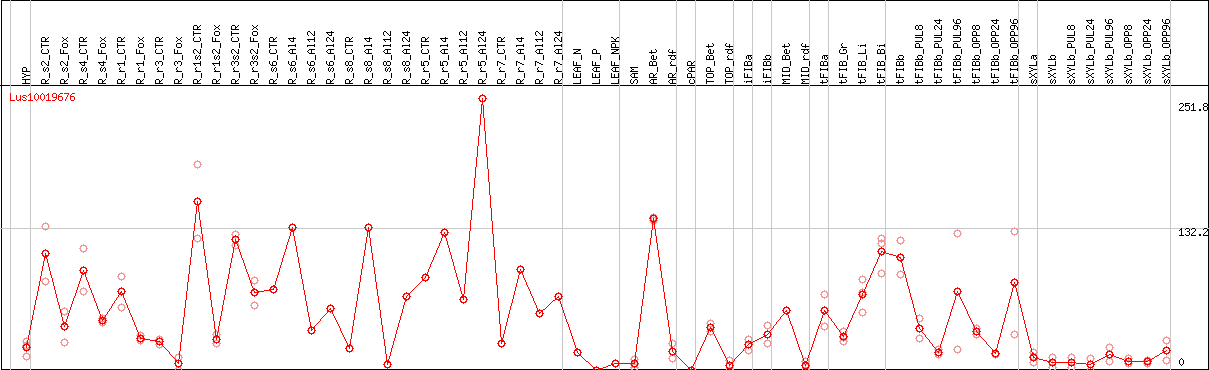
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019676 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.