External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G57270 238 / 9e-78
BG1
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
AT3G57260 201 / 4e-63
AtPR2, PR-2, PR2, BG2, BGL2
PATHOGENESIS-RELATED PROTEIN 2, "beta-1,3-glucanase 2", beta-1,3-glucanase 2 (.1)
AT4G16260 190 / 7e-59
Glycosyl hydrolase superfamily protein (.1)
AT3G57240 186 / 1e-57
BG3
"beta-1,3-glucanase 3", beta-1,3-glucanase 3 (.1)
AT5G56590 163 / 3e-47
O-Glycosyl hydrolases family 17 protein (.1)
AT2G26600 154 / 1e-45
Glycosyl hydrolase superfamily protein (.1.2)
AT1G32860 156 / 4e-45
Glycosyl hydrolase superfamily protein (.1)
AT5G20390 153 / 1e-44
Glycosyl hydrolase superfamily protein (.1)
AT2G01630 153 / 3e-44
O-Glycosyl hydrolases family 17 protein (.1.2)
AT3G15800 151 / 1e-43
Glycosyl hydrolase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014110
384 / 8e-136
AT3G57270 359 / 2e-124
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Lus10014108
249 / 1e-79
AT3G57270 366 / 3e-123
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Lus10019801
235 / 1e-76
AT3G57270 367 / 6e-127
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Lus10014109
235 / 2e-76
AT3G57270 361 / 1e-124
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Lus10027860
213 / 1e-67
AT3G57260 322 / 4e-109
PATHOGENESIS-RELATED PROTEIN 2, "beta-1,3-glucanase 2", beta-1,3-glucanase 2 (.1)
Lus10002807
204 / 2e-64
AT3G57240 339 / 7e-116
"beta-1,3-glucanase 3", beta-1,3-glucanase 3 (.1)
Lus10027859
189 / 6e-61
AT3G57240 185 / 4e-58
"beta-1,3-glucanase 3", beta-1,3-glucanase 3 (.1)
Lus10016883
183 / 2e-56
AT4G16260 391 / 1e-136
Glycosyl hydrolase superfamily protein (.1)
Lus10031037
182 / 1e-55
AT3G57240 312 / 3e-105
"beta-1,3-glucanase 3", beta-1,3-glucanase 3 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G057400
229 / 2e-74
AT3G57270 416 / 4e-146
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Potri.016G057600
226 / 7e-73
AT3G57270 414 / 2e-145
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Potri.006G046100
206 / 4e-65
AT3G57270 370 / 4e-128
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Potri.006G048100
204 / 2e-64
AT3G57270 382 / 6e-133
"beta-1,3-glucanase 1", beta-1,3-glucanase 1 (.1)
Potri.001G255100
197 / 6e-62
AT4G16260 369 / 9e-128
Glycosyl hydrolase superfamily protein (.1)
Potri.010G143166
191 / 1e-59
AT4G16260 432 / 5e-153
Glycosyl hydrolase superfamily protein (.1)
Potri.010G142800
191 / 5e-59
AT4G16260 433 / 2e-152
Glycosyl hydrolase superfamily protein (.1)
Potri.009G050300
177 / 2e-54
AT4G16260 285 / 5e-95
Glycosyl hydrolase superfamily protein (.1)
Potri.002G089200
166 / 9e-50
AT1G77780 309 / 6e-104
Glycosyl hydrolase superfamily protein (.1)
Potri.018G068600
165 / 6e-48
AT5G56590 685 / 0.0
O-Glycosyl hydrolases family 17 protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0058
Glyco_hydro_tim
PF00332
Glyco_hydro_17
Glycosyl hydrolases family 17
Representative CDS sequence
>Lus10019800 pacid=23164061 polypeptide=Lus10019800 locus=Lus10019800.g ID=Lus10019800.BGIv1.0 annot-version=v1.0
ATGCCTGGGTCCAAACCAACGTCTTCAACCACCTACCCGGCGTCAGATTCCGTTACATCGCTGTCGGAAACGAAGTCAAACCCGACGACCCGTTCGCCCA
ATTCCTCTATCCAGCGATGGTCAACATCCATAACTCCATTAGCCGATCTCCAGCGGCCAAACGGAATCAAAGTCTCCTCCGCATTCATCTACGGCGTCAT
CGGAGTCTCATACCCGCCGTCTCAGGGCGATTTCAACAAAGATTTGTGGTCGATCGTCGAACCCATTCTCAAATTCTTGGTCAAGAACAGATGCCCTCTG
CTTCTAAACGTGTACCCGTTTTTTGCGTACTTATTTAATCCGGGTCAGATTAGATTAGACTACGCTCTGTTCACGCCTCCAGCCGGGTTGGTACCCGACC
CGCCGCTGAGCTACGGTAATCTGTTCGATGCCATGGTGGACGCGGCGTACTCTGCTTTGGAGAAGGCTGGGGCAGGGGAACTGAAGATTGTGGTGGCGGA
GACGGGTTGGACGACCGACGGCGCCGGTAATGCGGCGACGGTGGAGAATGCGAGGCTTTATAATAACAATTTGGTGAAGCATGTGATGGGAGGGACGCCG
AAGCGGCCCGGAAAGGAGATTGAGACGTATATATTTGGGTTGTTTGACGAGGATTTGAAGTCGCCGGAGACTGAGAAGCATTGGGGAGTGTTTAGTCCCG
ATAAGAATCTTAAGTATCCGATGAATTTTAGAACATGA
AA sequence
>Lus10019800 pacid=23164061 polypeptide=Lus10019800 locus=Lus10019800.g ID=Lus10019800.BGIv1.0 annot-version=v1.0
MPGSKPTSSTTYPASDSVTSLSETKSNPTTRSPNSSIQRWSTSITPLADLQRPNGIKVSSAFIYGVIGVSYPPSQGDFNKDLWSIVEPILKFLVKNRCPL
LLNVYPFFAYLFNPGQIRLDYALFTPPAGLVPDPPLSYGNLFDAMVDAAYSALEKAGAGELKIVVAETGWTTDGAGNAATVENARLYNNNLVKHVMGGTP
KRPGKEIETYIFGLFDEDLKSPETEKHWGVFSPDKNLKYPMNFRT
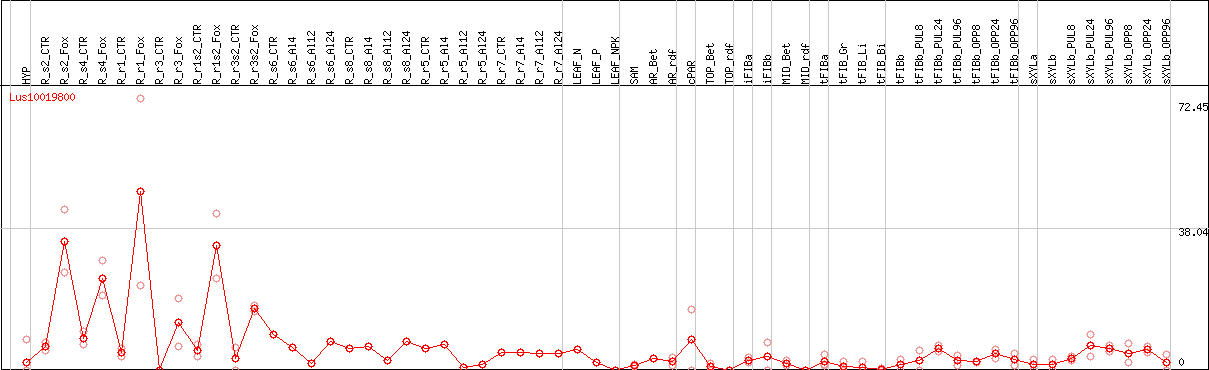
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.