External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G15570 184 / 6e-60
TRX-M3, GAT1, ATHM3, ATM3
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
AT3G15360 110 / 1e-30
ATHM4, ATM4, TRX-M4
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
AT1G03680 102 / 6e-28
ATHM1, ATM1, TRX-M1
ARABIDOPSIS THIOREDOXIN M-TYPE 1, thioredoxin M-type 1 (.1)
AT4G03520 100 / 1e-26
ATHM2
Thioredoxin superfamily protein (.1.2)
AT1G76760 76 / 2e-17
ATY1, TRX-Y1
thioredoxin Y1 (.1)
AT1G43560 71 / 9e-16
ATY2
thioredoxin Y2 (.1)
AT1G50320 69 / 1e-14
ATHX, ATX
thioredoxin X (.1)
AT1G52990 68 / 6e-14
thioredoxin family protein (.1)
AT4G12170 65 / 9e-14
Thioredoxin superfamily protein (.1)
AT3G06730 61 / 8e-12
TRXz, TRXP ,TRX z
thioredoxin putative plastidic, Thioredoxin z (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014069
342 / 3e-122
AT2G15570 182 / 3e-59
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
Lus10014798
121 / 3e-35
AT3G15360 181 / 2e-58
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10040887
121 / 4e-35
AT3G15360 182 / 4e-59
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10029755
115 / 1e-32
AT3G15360 162 / 3e-51
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10029752
114 / 2e-32
AT3G15360 165 / 3e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10042784
114 / 3e-32
AT3G15360 165 / 4e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10018875
69 / 5e-15
AT1G76760 176 / 6e-57
thioredoxin Y1 (.1)
Lus10028569
69 / 8e-15
AT1G76760 174 / 3e-56
thioredoxin Y1 (.1)
Lus10039499
61 / 1e-11
AT3G06730 238 / 9e-81
thioredoxin putative plastidic, Thioredoxin z (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF00085
Thioredoxin
Thioredoxin
Representative CDS sequence
>Lus10019847 pacid=23163960 polypeptide=Lus10019847 locus=Lus10019847.g ID=Lus10019847.BGIv1.0 annot-version=v1.0
ATGGCTACTTCTGCTTCCACGTGCTTCCGCGCTCTTCCTCTCCCCGCTTCACTCCAGCTCCGTCCAGCAGCGCAACTCAGTTCTAGCTCAGCAGTTTCTT
TCCCGTTCGACTTCTCCGCCGACGTGAGAGGTGGATTAGCCCTCCGTTCAACCAACTCGCGTCCGTCTTCTCCCTTCAAGGTCGTCTGCATGCGTGGATC
CAAAGCAGCTGTTGTCACCAAGGACTCATGGGAGAAGTCGATCCTGAAAAGCAATACCCCCGTGTTAGTCGAGTTCCATGCAAGCTGGTGTGGGCCCTGT
AAGATGGTTCACCGCGTTATTGATGAGCTTGCTGTAGAATACGAAGGGAAAATCCAATGCTTTGTCCTCAACGCCGACAATGATATAAAAATCGCTGAGG
ACTACCAGATCAAGGCTGTACCAGTGGTCCTGCTTTTCAAGAACGGTGAAAAGCGAGAGACTGTTGTCGGAACCATGCCCAAGGAATTCTACGCTGCTGC
CATCGAGAGGGTTCTGAAGGCATGA
AA sequence
>Lus10019847 pacid=23163960 polypeptide=Lus10019847 locus=Lus10019847.g ID=Lus10019847.BGIv1.0 annot-version=v1.0
MATSASTCFRALPLPASLQLRPAAQLSSSSAVSFPFDFSADVRGGLALRSTNSRPSSPFKVVCMRGSKAAVVTKDSWEKSILKSNTPVLVEFHASWCGPC
KMVHRVIDELAVEYEGKIQCFVLNADNDIKIAEDYQIKAVPVVLLFKNGEKRETVVGTMPKEFYAAAIERVLKA
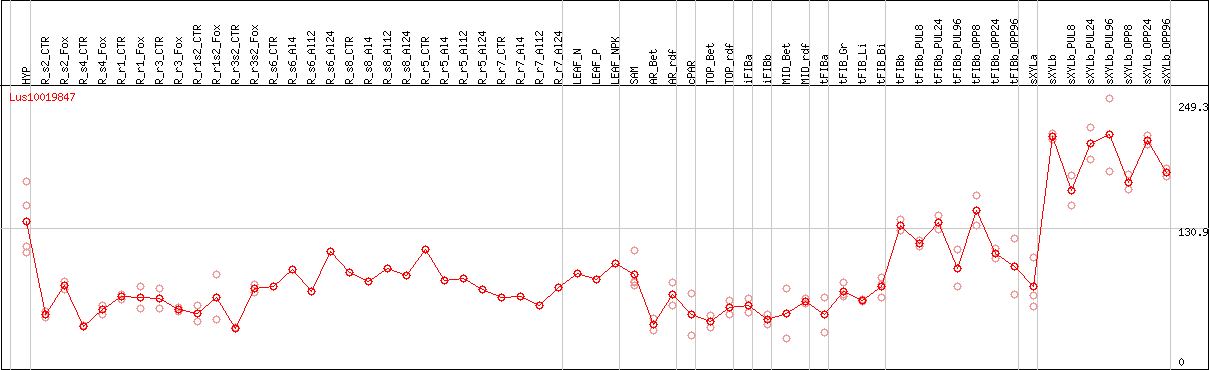
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019847 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.