Lus10019868 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Lus10019868 pacid=23164018 polypeptide=Lus10019868 locus=Lus10019868.g ID=Lus10019868.BGIv1.0 annot-version=v1.0
ATGTTCCCGTGGAACATCGCCAAGTCGGCGGAGGCCGTCTTATCGCAGTGGGCGCTAAAGAGGCTGTGTAAGTTTGTGCTGAAGAAGAAGCTAGGGCAAT
TCATTTTGGGGGAAATTGATCTTCAGCAGCTTGATGTTCAGCTCGGTCAAGGTGTGATCCAGCTCAGCGACCTTGCTCTTGATGTTGATTATCTTAATGA
CAAGTTTGGTGCAGAAGCATCTGTTGTTATAAAGGAGGGATCAATCGGCTCTCTTTCGGTTAAAATGCCTTGGAAGGGTAACGGTTTTGAGGTTGAGGTG
GATGAGCTTGAGCTTGTTTTTGGACCCTGTTTAAAAAATGAAGCACCGATTGTTGGTGATAAATTTCGTGCCAGTCAAGATAAAAGAACGAGCATGCATG
ATGAGATGGGAAAACTTGGCAATGTTTTGGACCCTGCAGCAAGATCGGGTTCTGTTGATATCCATGAAGGCGTGAAGACAATTGCAAAAATGGTTAAATG
GTTCTTAGCAAGCTTCCATGTGCGTGTTAAGAAATTACTGGTTGCTTTTGATCGATACTTGGAAAGTGAAAAGGAAGTGGGCTGCCACAAACTCGTGGTT
TTACGCATAGCTGAAGTGGAATGTGGAACATGTGTTTCTGATGATGTTGGTTCGATCACTTATGGAACCACTGATGATTTGCTCGGAATTAGTCGATTGA
CAAACTTTCTTAAATTTCAAGGTGCAGTCCTTGAACTCCTAGAAATGAATGATGCCAAAAGAGAATCTGTGTTATCTGAATCATTATCAGGTTCGACTTT
CGATGAGGTTCCTGCAAATGTAACAATGCCTATATTGTTGGGGAGTAAGGATGGATTTTCCGGGGAATTGAAGTTGAGTATACCCTGGAATAATGGTTCT
CTAGATATTCGCCAAGTGGATGCAGATATTTGTATAGATCCCGTGGAATTGAGAGTTCAGCCTGGTACTATTAAATATTTCCTTCTTGTGTGGGAAGCTT
ATAAGATCTTGAACAGGGATTCTGCTCTCAATTCACAGTCTAAGCCTGTGGACTCTGTTTACTTCAATTCAGCTTCTCAGTTGTATCCATCAACCCATAA
TTCTGCTTCAACACCAGCTGCGAATTTGTTTTGTGCTCCTGGTACTTCTCCACGGGTACAACAATCCTTCAGTGATTCGGTTCTTCCTGGATCACATCTT
ATACCTGACTGGATGCCGATATCTGTTAAAGATGGTGTGCAGGAGCTTGACCTTGGAGCAAGCGTGGATCAGTTCTTTGAGTGTTTTGACGGAATTAGAA
ATTCACAGTCAGCTTTGGGTAGTAGTGGGATGTGGGGCTGGACATGCTCTGTATTTGGTGCATTAACTGCTGCTTCCAGCCTTGCTTCTGGTTCTTTGCT
CGTTACATCTGAGCAGCAGCATATGCAGACGAATCTCAAAGTCCGTTGGACTGGAATTTCTGTTGTTTTATTGTTCCAAGATGAATACGAGCAGTGTTGC
TGCAAATCAACCGGGGACCAACGTAGTCTGAAACCTCATCATGTTACTGCTGAGTGCAAAGACATTTCAGTTGTTGTGCAGGTAAGTCCACAACAAATGA
TAATTGATGGTCATGTGAAATGTATTGAGGTTTCTGAAAACTCACAGAGTGGAAATGACGGTATGTTCTTCCCTCGGTCTTCTACTAATGGCCAGGATGA
GACTATTCAGATTCAGCATCTGCAAGCTGCTCTACTTAGTGCTCTCCCATTGGCCAATAATCTAAATTCAGATGATTGGAGTACTCCGCATGCTTCTGAT
ATTCATGCTGGAAAGACAAAAGTTAAAATGCTCAGTACCTCAGGTGTAACTCATTGCCAATTTGTTGTAAGTTCAGATTCATCTCACCGAGATGCGAAAG
TACCAGCATCTTTTTCATTGCATCTCCCACACTTTGTGTTCTGGGTGAACCTTATTTCGATAAAGATGCTAATGGATCTTGTTGAAGATGTGTCAAGTGC
TGTTAGTACCAGTAGCGGGAGCGATGAATTTAAAACTTCCAGTGAGAAGTGTGGCTCTACAGTTCATGAATCCAAAGGAGATGTGGAAGCATCATCTTCA
ACCGAAAACTGGCAATGTAACATTTCAATTCCCAGTGCAAGGATAATACTCTGTTTTCCATTAGAGATGGACAAACTAATGCAAAACAGCTGGTGCTGGG
ATCAGTTTGTTGCAATTGATTTCTCTTCGCCATCTGTCCGGGAGAAAGGGAAAATACAGGGTCACGGCCCTGTAGCTGATTCTACATCATGGAGAAGGTA
TATATCAAAGCCTGCATGGTCTGTGTATGTATCCGTTGGCGATCTAAGAGTCTATTTGATCAGCTCCAGAAAAAATGACGACGCAGTTATGGCTAGGATG
ACATTCTGCGTTCAGAAGGTTTTATCTGTCAGCAATAGAAGTGGGTGTCTTTCTACAGTTAGCATAGTGTCTCAGGATACTGTTATGACTGGTCCTTGGG
TTGCCGAGAGAGCAAAGTTCATGGCTACTTCAGAAGAGTCCAGGATGAGAGGAAACAACAACGCTAAAGGTCATGAATTTGCAACAGTGACAGCCGTGAA
AGACACAGATGATATGAGTTCTCGAACTCGCAGGGACATCTTGTTGACCTCTGCTTTTGTTTTACATGTTCATCTACCTCCTTGTATGATTGACATGGAC
AGTTCTCAGTATTGTAAGCTAGTTCTTCTGATTGGGCATATGATAAAAGGGTTGTCATGTACACCCTTGGAATCAGTTAGTCTAGATGAAGAACCATCTG
TTTCGCAGATATCAGTCCTAGTGGAGTGTGATTCGCTAGAAGTTTCAATTAGACCAGACACAATGGAGGAAAGACACAGTTCCTTGCAAAGTGAACTTCC
TGGATCATGGCTATGTTTCAAACTCAAAATTCGGAAACTTGAATTGATGTCTGTCTCGAGAATTGGAGGCATCGAAGATGCTACTTTCTTGTGGATAACC
CACAGAGAAGGTAAACTATGGGGATCAGTCACAAGCAACCCAAGCAAAATGTTTCTTCTCATATCATGCAGCAACTCTTCCAGGAAACGTGGGGATGGAG
GCGGTTCAAATGCATTGTCTTCTCGGTGGGCTGGTTCTGATATTGTCCACCTATGTGTTCCAGAGGCTATTCGTTCAATTACATCTATTACAATCAGATG
TGCGACAGTTGTGGCTGTTGGTGGTCGGTTGGACTGGTTTGATGCAATATTTTCTTTCTTCAATTTGTCCTCTCCTGAAGTTGTAAACACAGATCACACT
AATCTGCAGGAAGAAGGTCTCCAACCTACACACCAGACCTCTTTTCTTCTGAAGTTGGTTGATATTGGCTTAAGTTATGAACCACACTTGCAGAGTTCAA
TATCAACGGATCACAGTTCTTTCTCCAGTTCCACAAATACAGGAGAGAGTGATGTGGGGCCATATTTTTCCTGTCTTATAGCTGCTTCTTCCTTGACAGT
ATCCACTACTGCAATGGAAGATTCTGTGGACAATGACTACAAAATAAGAGTACAAGATCTAGGATTCCTTCTTTCTAAGGCACCTCATAATCCTGGTGGT
ATTTACAGTGTGGATTACCTTCGTGATATGAGATATGTTAAAGTTGCTCAAGAAGCCCTTATAGAAGCCGTTTTGAGAACTAACAGTGAGAATGGTCTTC
AATGGGAGTTAGAGTGTTCAAAGCTCCATGTTTGGATGAACACTTGTCATGATACTACCTCAGGACTTACTGGATTGGCTGCTCAACTGCAGCAGCTTTT
TGCTCCTGACTTGGAGGAAGCAGTTGTGCACTTGCAAGCTCGGTGGAACAATTTCTGTGAGGCACAAGAGAAGAGCAAGTTTAACGAGGAAGGTAAGGCT
ATTGTTTATGATTCCACCACAGCAACCTCTGAAGTGTTTGGCCTAAGTGCTTTCACTGTGAACGAACATAAGGCTGTAGTTGGGTTGATGGATGAGATAC
GCGGAGATGCATTTTCCTTGGATGCAAAGCAGAAGCAGGCCTGTCAAAATTCGGCTGGACCCGGTCTTCGTTTGCCACCTGCTGATAGCATCTTTGGACA
GCCTTATGGAATAGGTTATGAAGCTCAAGATAATTTCAAGATAATTTCTTGTGAACAGTCGTCATCAGTTGGCTTGGATAGAACTGAATCATCATTTTTA
GAGAATGGTTCCTTGCCGGAATTTATAGAAAGCTATTGCATATCTGACTTACGCCCTTTGTCAGAACTATCTATAGGCAGCATAGATCCCCCTGCTCGGG
TAAACAGTGGATGGTACGAAGAGTCGCCTGTAAGCATTGTAGAGGACCACATTTCAGATACTGGTGGGAGTGGTTACCTGAACCAGTTTCTTGATGACAA
GTTTCTGGAGGGCTTCTCTTTAACATCAGATCATTCTGAAAAGGCTACAGGGCGTGTACTTCTTAAGGACATAAATGTGAGCTGGAAAATGTATGCTGGT
TTTGACTGGGATCCACACGGCATGAACACTGAGCCATCTAAATGTACTGGTGGAAGGGACACAACTACTTGTCTAGAGGTTTCATTATGTGGGATGGAAT
TACGTTGTGAGTTGTTCCCACTGACGGGAGTATGTGCATCTAAGATTTCAATTTCCATTCAGGACTTCTATCTTTACGACAAGAGCTATACTGCACCTTG
GAAACTGATGCTTGGATATTACCATTCGAAAGATCATCCTAGGACAACAAATGCCAAAGCGCTCAAGCTGGACTTAGAAGCTAAATTTGATGTATGCCCT
ATTCTTATACGGGTCGATTACACACCTAGTCGTGTTGATTTAACAGCATTAGGAGGTGGGAAGTATGTGGAACTTGTAAACCTTGTTCCTTGGAAGGGGG
TTGAACTCGAGCTTAAACACGTACATGCTTCTGGTATCTATGGCTGGGAAAGCGCATGTGAAACAATGTTCGGGGAGTGGCTGGAAGATATATCACAAAA
TCAGGTACGAAAGATCTTACGAGGCCTCCCCACTATACGATCCTTTGTTGCTGTTGGTTCTGGGGCTGCAAAGCTCGTATCACTGCCAGTTCAGAGCTAT
AAGAAAGATCAGCGAGTCCTCAAGGGAATGCAAAGAGGTACCGTTGCATTTCTTAGAAGTATATCACTTGAAGCTGTCGGGCTTGGAGTGCATTTGGCAG
CCGGAGCACATGATATTTTACTTCAAGCCGAGTATATCCTCACCAGTGTTCCTTCCCCAGTATCATCATCGTCCTCCACACAGGGCAAAAAGAAGTCAAA
TGTAAGAAATAATCAACCAAAAGATGCCCAAGAAGGAATCCAAAAGGCTTGTGCGAGCATCAGTGATGGCCTGGGGAAAACAGCTTCTGCACTTGTTCGA
ACACCGCTGAAAAAATACCAATACGGGTCCAGTGCTGGATCCGCATTGGCAACTGCAGTTCGGGGCACTCCTGCTGCTGCTATTGCTCCAGTCTCTGCCT
GTGCCAGTGCTGCACATTACGCTCTACTTGGACTTAGAAATAGCCTTGATCCCGAGCACAAAAAAGAGTCGATGGAGAAATATCTAGGTCCGACTCAAAC
ACACAATAGCGATTGA
|
|||||||||||||||
|
AA sequence
|
>Lus10019868 pacid=23164018 polypeptide=Lus10019868 locus=Lus10019868.g ID=Lus10019868.BGIv1.0 annot-version=v1.0
MFPWNIAKSAEAVLSQWALKRLCKFVLKKKLGQFILGEIDLQQLDVQLGQGVIQLSDLALDVDYLNDKFGAEASVVIKEGSIGSLSVKMPWKGNGFEVEV
DELELVFGPCLKNEAPIVGDKFRASQDKRTSMHDEMGKLGNVLDPAARSGSVDIHEGVKTIAKMVKWFLASFHVRVKKLLVAFDRYLESEKEVGCHKLVV
LRIAEVECGTCVSDDVGSITYGTTDDLLGISRLTNFLKFQGAVLELLEMNDAKRESVLSESLSGSTFDEVPANVTMPILLGSKDGFSGELKLSIPWNNGS
LDIRQVDADICIDPVELRVQPGTIKYFLLVWEAYKILNRDSALNSQSKPVDSVYFNSASQLYPSTHNSASTPAANLFCAPGTSPRVQQSFSDSVLPGSHL
IPDWMPISVKDGVQELDLGASVDQFFECFDGIRNSQSALGSSGMWGWTCSVFGALTAASSLASGSLLVTSEQQHMQTNLKVRWTGISVVLLFQDEYEQCC
CKSTGDQRSLKPHHVTAECKDISVVVQVSPQQMIIDGHVKCIEVSENSQSGNDGMFFPRSSTNGQDETIQIQHLQAALLSALPLANNLNSDDWSTPHASD
IHAGKTKVKMLSTSGVTHCQFVVSSDSSHRDAKVPASFSLHLPHFVFWVNLISIKMLMDLVEDVSSAVSTSSGSDEFKTSSEKCGSTVHESKGDVEASSS
TENWQCNISIPSARIILCFPLEMDKLMQNSWCWDQFVAIDFSSPSVREKGKIQGHGPVADSTSWRRYISKPAWSVYVSVGDLRVYLISSRKNDDAVMARM
TFCVQKVLSVSNRSGCLSTVSIVSQDTVMTGPWVAERAKFMATSEESRMRGNNNAKGHEFATVTAVKDTDDMSSRTRRDILLTSAFVLHVHLPPCMIDMD
SSQYCKLVLLIGHMIKGLSCTPLESVSLDEEPSVSQISVLVECDSLEVSIRPDTMEERHSSLQSELPGSWLCFKLKIRKLELMSVSRIGGIEDATFLWIT
HREGKLWGSVTSNPSKMFLLISCSNSSRKRGDGGGSNALSSRWAGSDIVHLCVPEAIRSITSITIRCATVVAVGGRLDWFDAIFSFFNLSSPEVVNTDHT
NLQEEGLQPTHQTSFLLKLVDIGLSYEPHLQSSISTDHSSFSSSTNTGESDVGPYFSCLIAASSLTVSTTAMEDSVDNDYKIRVQDLGFLLSKAPHNPGG
IYSVDYLRDMRYVKVAQEALIEAVLRTNSENGLQWELECSKLHVWMNTCHDTTSGLTGLAAQLQQLFAPDLEEAVVHLQARWNNFCEAQEKSKFNEEGKA
IVYDSTTATSEVFGLSAFTVNEHKAVVGLMDEIRGDAFSLDAKQKQACQNSAGPGLRLPPADSIFGQPYGIGYEAQDNFKIISCEQSSSVGLDRTESSFL
ENGSLPEFIESYCISDLRPLSELSIGSIDPPARVNSGWYEESPVSIVEDHISDTGGSGYLNQFLDDKFLEGFSLTSDHSEKATGRVLLKDINVSWKMYAG
FDWDPHGMNTEPSKCTGGRDTTTCLEVSLCGMELRCELFPLTGVCASKISISIQDFYLYDKSYTAPWKLMLGYYHSKDHPRTTNAKALKLDLEAKFDVCP
ILIRVDYTPSRVDLTALGGGKYVELVNLVPWKGVELELKHVHASGIYGWESACETMFGEWLEDISQNQVRKILRGLPTIRSFVAVGSGAAKLVSLPVQSY
KKDQRVLKGMQRGTVAFLRSISLEAVGLGVHLAAGAHDILLQAEYILTSVPSPVSSSSSTQGKKKSNVRNNQPKDAQEGIQKACASISDGLGKTASALVR
TPLKKYQYGSSAGSALATAVRGTPAAAIAPVSACASAAHYALLGLRNSLDPEHKKESMEKYLGPTQTHNSD
|
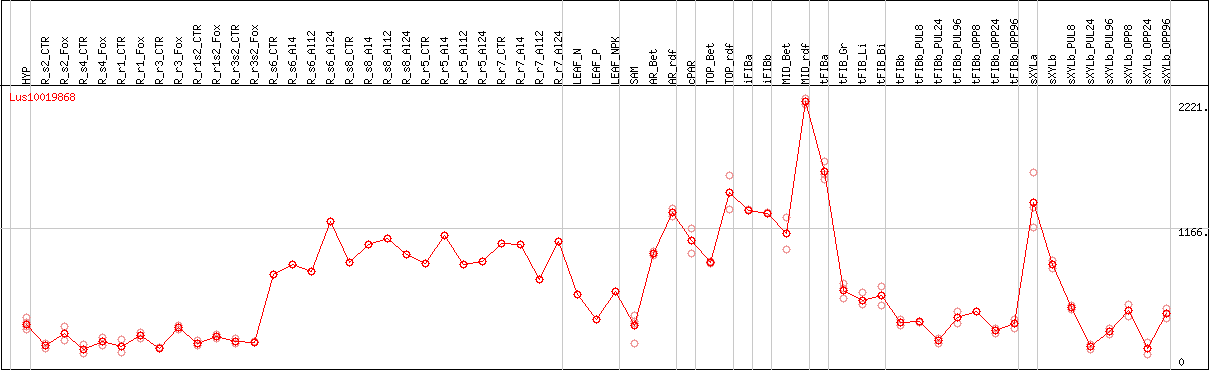
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019868 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
