External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G03170 246 / 3e-82
ATFLA11, FLA11, IRX13
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
AT5G60490 201 / 2e-64
FLA12
FASCICLIN-like arabinogalactan-protein 12 (.1)
AT1G03870 153 / 8e-46
FLA9
FASCICLIN-like arabinoogalactan 9 (.1)
AT5G44130 152 / 1e-45
FLA13
FASCICLIN-like arabinogalactan protein 13 precursor (.1)
AT2G20520 146 / 4e-43
FLA6
FASCICLIN-like arabinogalactan 6 (.1)
AT2G04780 85 / 1e-19
FLA7
FASCICLIN-like arabinoogalactan 7 (.1.2)
AT2G45470 79 / 8e-17
AGP8, FLA8
FASCICLIN-like arabinogalactan protein 8 (.1)
AT3G60900 78 / 2e-16
FLA10
FASCICLIN-like arabinogalactan-protein 10 (.1)
AT3G46550 74 / 6e-15
FLA4, SOS5
salt overly sensitive 5, fasciclin-like arabinogalactan-protein 4, Fasciclin-like arabinogalactan family protein (.1)
AT4G12730 66 / 3e-12
FLA2
FASCICLIN-like arabinogalactan 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026499
360 / 2e-127
AT5G03170 291 / 5e-100
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10036111
181 / 2e-56
AT5G03170 209 / 1e-67
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10002978
176 / 7e-55
AT5G03170 209 / 1e-67
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10036114
169 / 8e-52
AT5G03170 191 / 3e-60
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10017696
160 / 1e-48
AT2G20520 244 / 2e-81
FASCICLIN-like arabinogalactan 6 (.1)
Lus10036112
159 / 6e-48
AT5G03170 197 / 1e-62
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10002985
157 / 4e-47
AT5G60490 180 / 4e-56
FASCICLIN-like arabinogalactan-protein 12 (.1)
Lus10033651
156 / 4e-47
AT2G20520 234 / 2e-77
FASCICLIN-like arabinogalactan 6 (.1)
Lus10036113
154 / 7e-46
AT5G03170 173 / 6e-53
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G129200
277 / 1e-94
AT5G03170 199 / 9e-64
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Potri.016G088700
277 / 2e-94
AT5G03170 203 / 2e-65
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Potri.015G129400
192 / 2e-61
AT5G60490 206 / 2e-66
FASCICLIN-like arabinogalactan-protein 12 (.1)
Potri.012G127900
190 / 3e-60
AT5G60490 207 / 6e-67
FASCICLIN-like arabinogalactan-protein 12 (.1)
Potri.013G151366
179 / 2e-55
AT5G60490 167 / 7e-51
FASCICLIN-like arabinogalactan-protein 12 (.1)
Potri.013G151300
179 / 2e-55
AT5G03170 167 / 4e-51
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Potri.013G151432
177 / 4e-55
AT5G60490 167 / 6e-51
FASCICLIN-like arabinogalactan-protein 12 (.1)
Potri.013G014200
176 / 2e-54
AT5G60490 167 / 6e-51
FASCICLIN-like arabinogalactan-protein 12 (.1)
Potri.013G151500
176 / 2e-54
AT5G60490 167 / 4e-51
FASCICLIN-like arabinogalactan-protein 12 (.1)
Potri.019G121200
176 / 2e-54
AT5G03170 166 / 2e-50
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02469
Fasciclin
Fasciclin domain
Representative CDS sequence
>Lus10019929 pacid=23163998 polypeptide=Lus10019929 locus=Lus10019929.g ID=Lus10019929.BGIv1.0 annot-version=v1.0
ATGAGGAGCCGACTTCAATCTCCATTCATGTTCTTGTTCTTCTTTTTCCTCTACTGCTCCATTTCATATGCGCAATCCCCTGCGCCAGGCCCCTCTGGCC
CTGCCAACATCACAGCCATCTTAGAGAAGGCCGGCCAGTTCACGACCTTGATCAAGCTGATGAAGACCACCCAAGAAGCCGACCAAATCAACACTCAGTT
GAACAGCAGCTCGAGCTCGGGCCTCACCATCTTCGCTCCTACAGACAACGCGTTCACTAGCCTCAAAGCCGGAACTCTAAACTCCCTCTCGGATCAGCAG
AAAGTGCAGCTGATCCAGTTCCACATCCTCCCAACTTTCATCTCCATGTCCCAGTTCCAGACCGTGAGCAACCCTCTGAGGACCCAGGCTGGCGACAGCG
CCAACGGGAAGTTCCCTCTCAACGTGACCACTTCGGGGACTCAAGTGAACGTCACGACCGGAGTCGATACTGCTAATGTGGACAACACATTGTACACTGA
CGGCCAGCTCGTTGTTTACCAAGTGAGCAAGGTGCTCTTGCCCTTGGATATCTTTGGTTCGTCGGCTGCACCAGCTCCGGCTCCGACCAAGTCCGGGAAG
GCTCTCACTTCCAAGGCTCCCACAGCTGGGAATGCAGACGATTCCCCTGGCGATGCTTCTGGCGGTGAAGGAAGAGTTGTATCCCTTGGTTTGATGGCTT
TTGCTGCAGCTGTCCTTGTGTTATGA
AA sequence
>Lus10019929 pacid=23163998 polypeptide=Lus10019929 locus=Lus10019929.g ID=Lus10019929.BGIv1.0 annot-version=v1.0
MRSRLQSPFMFLFFFFLYCSISYAQSPAPGPSGPANITAILEKAGQFTTLIKLMKTTQEADQINTQLNSSSSSGLTIFAPTDNAFTSLKAGTLNSLSDQQ
KVQLIQFHILPTFISMSQFQTVSNPLRTQAGDSANGKFPLNVTTSGTQVNVTTGVDTANVDNTLYTDGQLVVYQVSKVLLPLDIFGSSAAPAPAPTKSGK
ALTSKAPTAGNADDSPGDASGGEGRVVSLGLMAFAAAVLVL
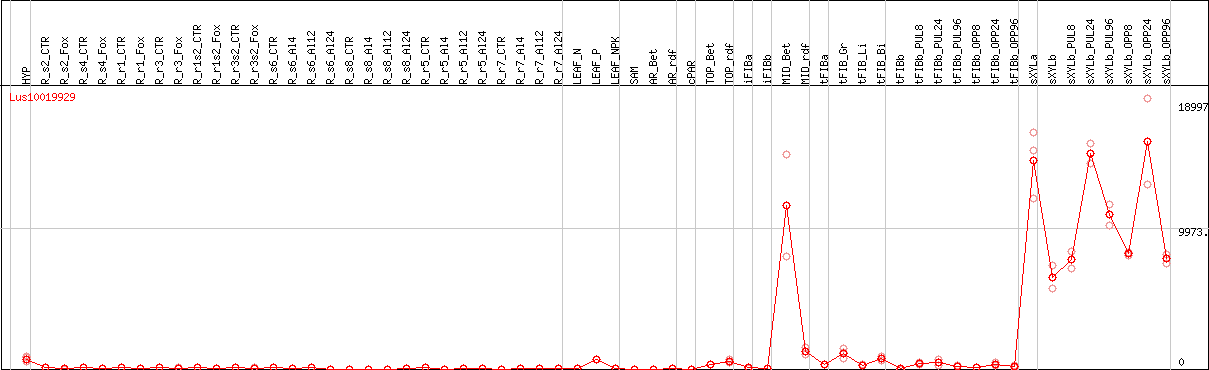
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019929 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.