External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G04350 100 / 5e-25
EMB2369
EMBRYO DEFECTIVE 2369, tRNA synthetase class I (I, L, M and V) family protein (.1)
AT5G49030 46 / 4e-06
OVA2
ovule abortion 2, tRNA synthetase class I (I, L, M and V) family protein (.1), tRNA synthetase class I (I, L, M and V) family protein (.2), tRNA synthetase class I (I, L, M and V) family protein (.3)
AT4G10320 46 / 6e-06
tRNA synthetase class I (I, L, M and V) family protein (.1)
AT2G37550 45 / 6e-06
ASP1, AGD7
yeast pde1 suppressor 1, ARF-GAP domain 7 (.1.2)
AT3G53710 39 / 0.0008
AGD6
ARF-GAP domain 6 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015552
106 / 5e-27
AT4G04350 1412 / 0.0
EMBRYO DEFECTIVE 2369, tRNA synthetase class I (I, L, M and V) family protein (.1)
Lus10038317
66 / 6e-14
AT2G37550 166 / 5e-50
yeast pde1 suppressor 1, ARF-GAP domain 7 (.1.2)
Lus10023766
49 / 7e-07
AT2G37550 485 / 4e-170
yeast pde1 suppressor 1, ARF-GAP domain 7 (.1.2)
Lus10003978
45 / 1e-05
AT2G37550 516 / 0.0
yeast pde1 suppressor 1, ARF-GAP domain 7 (.1.2)
Lus10023765
43 / 5e-05
AT2G37550 507 / 4e-178
yeast pde1 suppressor 1, ARF-GAP domain 7 (.1.2)
Lus10024041
42 / 0.0001
AT4G10320 1819 / 0.0
tRNA synthetase class I (I, L, M and V) family protein (.1)
Lus10041706
42 / 0.0001
AT4G10320 1820 / 0.0
tRNA synthetase class I (I, L, M and V) family protein (.1)
Lus10033331
40 / 0.0005
AT1G14610 1580 / 0.0
VALYL TRNA SYNTHETASE, TWIN 2, valyl-tRNA synthetase / valine--tRNA ligase (VALRS) (.1)
Lus10016658
40 / 0.0006
AT4G13780 1280 / 0.0
methionine--tRNA ligase, putative / methionyl-tRNA synthetase, putative / MetRS, putative (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G009600
102 / 2e-25
AT4G04350 1443 / 0.0
EMBRYO DEFECTIVE 2369, tRNA synthetase class I (I, L, M and V) family protein (.1)
Potri.007G126501
47 / 2e-06
AT4G10320 1167 / 0.0
tRNA synthetase class I (I, L, M and V) family protein (.1)
Potri.017G033200
47 / 3e-06
AT4G10320 1940 / 0.0
tRNA synthetase class I (I, L, M and V) family protein (.1)
Potri.010G014500
44 / 3e-05
AT5G49030 1612 / 0.0
ovule abortion 2, tRNA synthetase class I (I, L, M and V) family protein (.1), tRNA synthetase class I (I, L, M and V) family protein (.2), tRNA synthetase class I (I, L, M and V) family protein (.3)
Potri.016G095100
43 / 5e-05
AT3G53710 462 / 9e-161
ARF-GAP domain 6 (.1.2)
Potri.006G084000
40 / 0.0005
AT3G53710 498 / 8e-175
ARF-GAP domain 6 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0039
HUP
PF00133
tRNA-synt_1
tRNA synthetases class I (I, L, M and V)
Representative CDS sequence
>Lus10020034 pacid=23156214 polypeptide=Lus10020034 locus=Lus10020034.g ID=Lus10020034.BGIv1.0 annot-version=v1.0
ATGACGAAACAACTCAATACCACCACCAGTACAGTACCTCCTCATTTTTGCTTCCACTTGAGAAGTAATGATGGAGTTTGGATTGGCAACCCAAATCCCG
ACCCTCTTCCTTCCTTCTCTTCCCTTTCCTCCTCCTCCTCCTCTCGCCGGAGGTCTCGACAGTCGTCTCTTGGGGCTGGTTTGCACGTTGGGCATCCACT
TGGCTATACTGCCACTGATATTCTTGCTAGGTATAAGCGAATGCAGGGTTATAATGTTTTGCACCCTATGGGATGGGATGCCTTTGGTTTGCCCGCTGAG
CAATACGCCATTGATGACCTCCATGCGGTTGGGAAGACAGAGGTGCTTGCAAGAGCAGCTGAAGCACATGAAGATGCCATAGGTGGCAGAGGCCCAATTA
GGATTCTTGTTGGAGCAGTCGACGCCGATCTTGTTTGCAGGTTGCCCTTGGAGATCCCCGTAAGCGATGTTTCTGGCGACGTCGAGCGGGAGGATCATGC
TATGGCTTTGCTTTGGCCATAG
AA sequence
>Lus10020034 pacid=23156214 polypeptide=Lus10020034 locus=Lus10020034.g ID=Lus10020034.BGIv1.0 annot-version=v1.0
MTKQLNTTTSTVPPHFCFHLRSNDGVWIGNPNPDPLPSFSSLSSSSSSRRRSRQSSLGAGLHVGHPLGYTATDILARYKRMQGYNVLHPMGWDAFGLPAE
QYAIDDLHAVGKTEVLARAAEAHEDAIGGRGPIRILVGAVDADLVCRLPLEIPVSDVSGDVEREDHAMALLWP
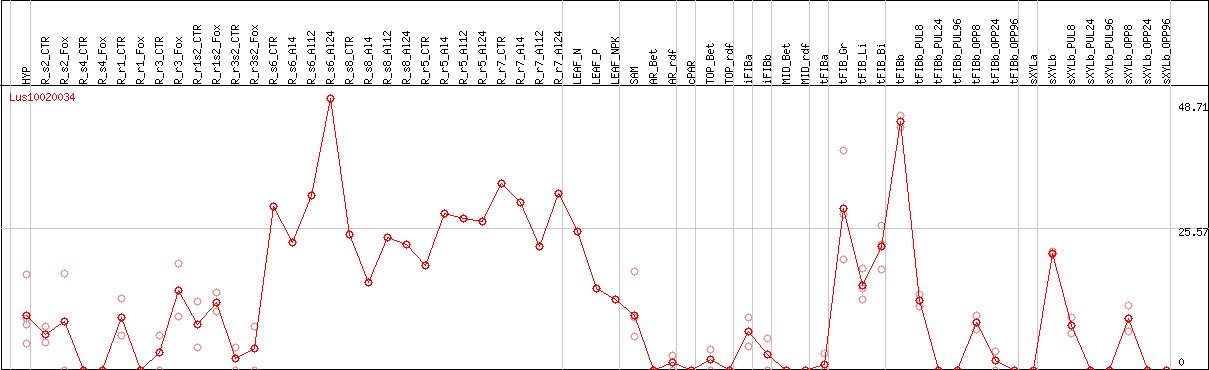
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020034 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.