External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G31590 56 / 9e-11
ATCSLC5, ATCSLC05
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
AT2G24630 54 / 3e-10
ATCSLC8, ATCSLC08
CELLULOSE-SYNTHASE LIKE C8, Glycosyl transferase family 2 protein (.1)
AT4G07960 53 / 8e-10
ATCSLC12
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
AT3G07330 52 / 1e-09
ATCSLC6, ATCSLC06
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
AT3G28180 47 / 1e-07
ATCSLC4, CSLC4, ATCSLC04
CELLULOSE-SYNTHASE LIKE C4, Cellulose-synthase-like C4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026923
61 / 2e-12
AT4G31590 1078 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Lus10038217
55 / 2e-10
AT3G07330 821 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Lus10018651
52 / 1e-09
AT4G07960 978 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Lus10007715
52 / 1e-09
AT4G07960 1010 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Lus10025886
52 / 2e-09
AT3G07330 994 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Lus10039440
49 / 3e-08
AT3G28180 1035 / 0.0
CELLULOSE-SYNTHASE LIKE C4, Cellulose-synthase-like C4 (.1)
Lus10039475
48 / 4e-08
AT3G28180 1035 / 0.0
CELLULOSE-SYNTHASE LIKE C4, Cellulose-synthase-like C4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G270900
56 / 5e-11
AT4G31590 1085 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Potri.018G009300
56 / 6e-11
AT4G31590 1077 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Potri.002G248400
54 / 5e-10
AT3G07330 957 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Potri.002G114200
51 / 4e-09
AT4G07960 1031 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Potri.005G146900
51 / 4e-09
AT4G07960 1050 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
PFAM info
Representative CDS sequence
>Lus10020121 pacid=23142059 polypeptide=Lus10020121 locus=Lus10020121.g ID=Lus10020121.BGIv1.0 annot-version=v1.0
ATGGAAGAAGCAGCTGCAGCAGTGCCGAAACCGAAAAAGATAGTGAACAAGATTTACAGGAAGGAGCTAGCTCTTGCATTCCTCTTGCTCACAGCTTCTG
CAAGGAGTCTCTTATCTGCTCAAGGAGTCCATTTCTACTTCCTTCTGTTCCAAGGTGTCACATTTCTGTTGGTGGGTTTGGACCTCATTGGACAGCAAAT
GACTTGA
AA sequence
>Lus10020121 pacid=23142059 polypeptide=Lus10020121 locus=Lus10020121.g ID=Lus10020121.BGIv1.0 annot-version=v1.0
MEEAAAAVPKPKKIVNKIYRKELALAFLLLTASARSLLSAQGVHFYFLLFQGVTFLLVGLDLIGQQMT
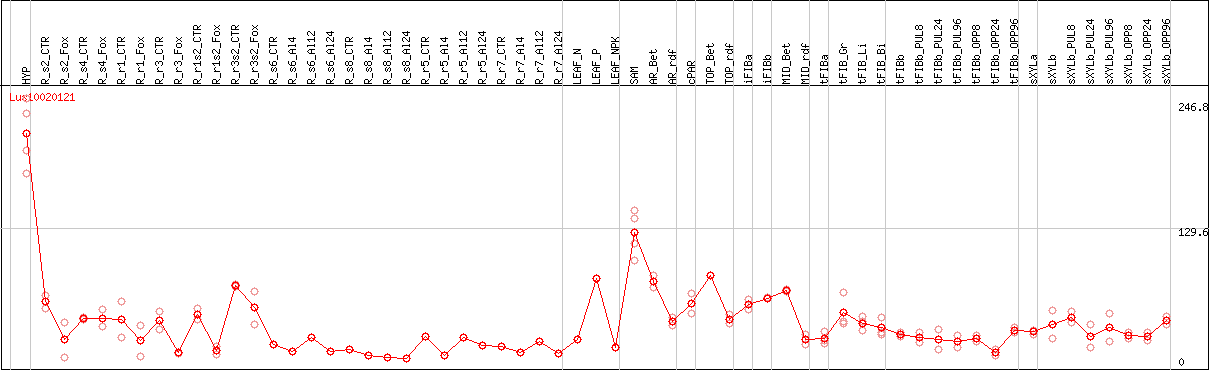
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020121 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.