External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G12700 239 / 9e-73
RPF1
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
AT1G12775 230 / 5e-70
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63150 218 / 9e-66
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G12300 218 / 2e-65
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G22470 216 / 4e-65
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62930 214 / 3e-64
RPF3
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63230 206 / 4e-64
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63080 212 / 2e-63
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62680 210 / 3e-63
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G64100 211 / 1e-62
pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014245
461 / 2e-160
AT1G12700 427 / 5e-141
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10014244
425 / 2e-146
AT1G12700 397 / 7e-130
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10003433
402 / 7e-137
AT1G12700 396 / 2e-129
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10003427
374 / 3e-129
AT1G12700 291 / 8e-92
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10008593
369 / 3e-124
AT1G12700 400 / 7e-131
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10009201
339 / 4e-116
AT1G12700 274 / 3e-86
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10021074
318 / 5e-107
AT1G62930 273 / 5e-86
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10022861
318 / 2e-104
AT1G12700 370 / 7e-121
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10014247
315 / 3e-103
AT1G62930 340 / 3e-109
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G046200
290 / 2e-93
AT1G12700 501 / 5e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.013G032600
288 / 2e-92
AT1G12700 493 / 2e-166
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.006G257300
286 / 7e-92
AT1G62930 481 / 3e-163
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G038400
284 / 5e-91
AT1G12700 478 / 7e-161
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.019G021200
283 / 8e-91
AT1G12700 491 / 2e-166
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.006G271400
281 / 4e-90
AT1G12700 486 / 3e-164
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G050180
280 / 2e-89
AT1G12700 488 / 2e-164
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.006G271200
279 / 2e-89
AT1G62930 475 / 3e-161
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G045000
280 / 3e-89
AT1G12700 502 / 4e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.013G034200
278 / 1e-88
AT1G12700 511 / 6e-174
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10020134 pacid=23142030 polypeptide=Lus10020134 locus=Lus10020134.g ID=Lus10020134.BGIv1.0 annot-version=v1.0
ATGAATGCATGTGAAGCTGCAGCTTCAGTGTCTAGGAAAATGCAATTCGTTGGGTTGTTACATGATTCATTTGGATCCAATACACTGATTTATTGTTTGT
TGGTTGTGCATTCTCAGTACTGGGCAAAAAATTTTAAATTTGGTCATCAACCTTGTGCGGTAACTTTTAATACCTTAATTGTTGGATTGTGCAAAGATGG
AAAAGTTGATCATGCTGTGAAGCTTTTCGATAAGATTGTGTCTTTAAGATATCAACCGGATGTTTTCACTTATAGTTCCATTATTGATGGCCTGTGTAAG
ATGGGTAAAACTAGTGCTGCTCATAGATTGCTGGAGAAAATGGAAGATGTAGGTTGTCAACCAGATGTGGTTACTTACAGTACAATTATAGATAGCCTAT
GTAAGGATGGTTTGGTATCCAAAGCCGTAGATGTGTTTATCAAGATGAAGGATGAAAAGTGTTCACCAAATGTTGTCACTTACAGCTGTTTGATTCATGG
TTTATGTGTTTCAAGTCGAAAGAATGAAGCTATATTGTCGTTCGGCAAAATGTTAGAAAGGGATGTCGTTCCAAATGTAGTCACCGATAACATTCTGATT
GATTCGTTTTGCAAAGATGGAATGGTCCGTGAGGCTAAAGCTATAGTCAAAGTGATGATACAAAAAGGACAGCATCCGGATATCGTCACTTACAACTCAT
TGCTAGATGGTATGATGATTAATGGATATTGTAAGAGTGAAAGGTTCAGTGAAGCCAAACAGTTAGTGAATAATATGCTTGAGAAGGATTTAGTTCTAGA
CACTGTTACGTATACTACTCTTGTGGATGGGTTTTGCCGAGCGCGGAGGTTCCAAGATGCAGAAAAAGTTCTTAAGGAAATGTGCCATCAGGGACAGCTT
CCGAATGTCGTGACATTCAACAGTTTGCTGAATGGTTGGTGCCAAATAGGTCATCTTGACAAGGCATTAGCTCTATTCCGAGAAATCAAAATTATGGACT
TTGTAGGGATGGTCTGGTAG
AA sequence
>Lus10020134 pacid=23142030 polypeptide=Lus10020134 locus=Lus10020134.g ID=Lus10020134.BGIv1.0 annot-version=v1.0
MNACEAAASVSRKMQFVGLLHDSFGSNTLIYCLLVVHSQYWAKNFKFGHQPCAVTFNTLIVGLCKDGKVDHAVKLFDKIVSLRYQPDVFTYSSIIDGLCK
MGKTSAAHRLLEKMEDVGCQPDVVTYSTIIDSLCKDGLVSKAVDVFIKMKDEKCSPNVVTYSCLIHGLCVSSRKNEAILSFGKMLERDVVPNVVTDNILI
DSFCKDGMVREAKAIVKVMIQKGQHPDIVTYNSLLDGMMINGYCKSERFSEAKQLVNNMLEKDLVLDTVTYTTLVDGFCRARRFQDAEKVLKEMCHQGQL
PNVVTFNSLLNGWCQIGHLDKALALFREIKIMDFVGMVW
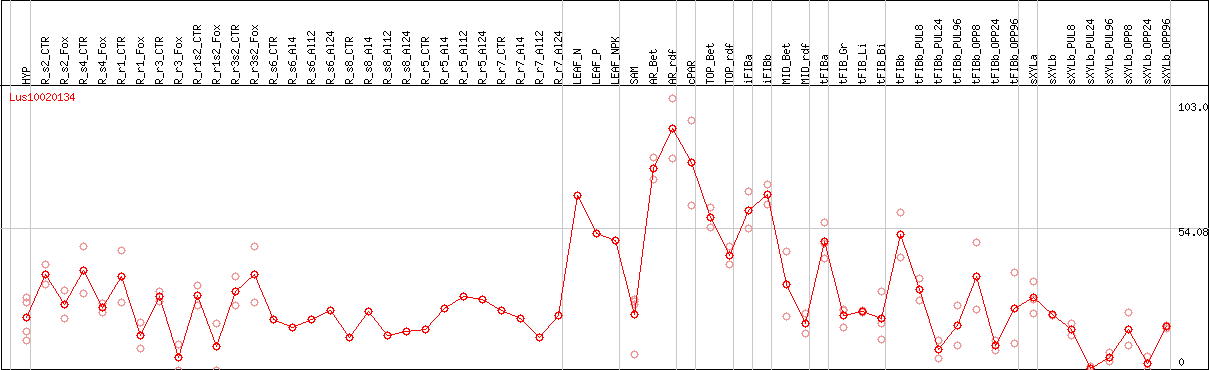
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020134 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.