External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G43570 51 / 2e-08
CHI
"chitinase, putative", chitinase, putative (.1)
AT2G43620 50 / 6e-08
Chitinase family protein (.1)
AT2G43580 49 / 8e-08
Chitinase family protein (.1)
AT2G43590 48 / 2e-07
Chitinase family protein (.1)
AT3G54420 48 / 2e-07
ATCHITIV, CHIV, ATEP3
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
AT2G43610 48 / 3e-07
Chitinase family protein (.1)
AT2G43600 42 / 4e-05
Chitinase family protein (.1)
AT3G12500 39 / 0.0005
PR-3, PR3, CHI-B, B-CHI, ATHCHIB
PATHOGENESIS-RELATED 3, basic chitinase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001773
66 / 4e-14
AT2G43570 51 / 6e-08
"chitinase, putative", chitinase, putative (.1)
Lus10020253
62 / 5e-13
AT2G43610 54 / 4e-09
Chitinase family protein (.1)
Lus10028377
49 / 2e-07
AT3G12500 393 / 1e-137
PATHOGENESIS-RELATED 3, basic chitinase (.1)
Lus10025948
48 / 2e-07
AT2G43610 50 / 3e-07
Chitinase family protein (.1)
Lus10000453
48 / 4e-07
AT3G54420 382 / 3e-135
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Lus10041830
45 / 3e-06
AT3G12500 402 / 4e-141
PATHOGENESIS-RELATED 3, basic chitinase (.1)
Lus10041828
42 / 6e-06
AT3G12500 59 / 8e-12
PATHOGENESIS-RELATED 3, basic chitinase (.1)
Lus10014253
42 / 2e-05
AT2G43570 46 / 1e-06
"chitinase, putative", chitinase, putative (.1)
Lus10006552
42 / 2e-05
AT3G04720 244 / 1e-82
HEVEIN-LIKE, pathogenesis-related 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G093700
50 / 6e-08
AT3G54420 413 / 3e-147
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.019G094000
49 / 7e-08
AT3G54420 340 / 2e-118
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.019G093800
49 / 8e-08
AT3G54420 369 / 8e-130
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.013G125100
49 / 9e-08
AT3G54420 336 / 1e-116
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.019G094100
49 / 1e-07
AT3G54420 350 / 3e-122
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.019G093900
49 / 1e-07
AT3G54420 360 / 3e-126
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.013G125000
46 / 9e-07
AT3G54420 421 / 1e-150
CHITINASE CLASS IV, homolog of carrot EP3-3 chitinase (.1)
Potri.009G141700
46 / 2e-06
AT3G12500 405 / 3e-142
PATHOGENESIS-RELATED 3, basic chitinase (.1)
Potri.009G142300
45 / 2e-06
AT3G12500 340 / 1e-116
PATHOGENESIS-RELATED 3, basic chitinase (.1)
Potri.009G141800
44 / 6e-06
AT3G12500 351 / 9e-121
PATHOGENESIS-RELATED 3, basic chitinase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00187
Chitin_bind_1
Chitin recognition protein
Representative CDS sequence
>Lus10020338 pacid=23144161 polypeptide=Lus10020338 locus=Lus10020338.g ID=Lus10020338.BGIv1.0 annot-version=v1.0
ATGGCACCCATTAACGTTCTTAATTATTTCCTGTCTCTCATACTAGTCCTAATAGCGTCGACTAAACTCCAACATTTTGCCGCCGCCGGCACGTCTTTCC
GGCTAGTCACGGCAGCCTCCGCGCCCTACAATTCGCCGCCGGCGCCTGTGATGGTAAACCTCACCCACGACGACGGCTGGTACGACCACCCGCACTGCGG
ATACCAGTCCCCGAAAGAAACCTGCGCGACGGGATTATGCTGCAGCCGGTTTGGCTACTGTGGGACAAAACCTGAGTATTGCGGCAAGGGCTGCCAAGAC
GGCCCCTGTCAAGGCGGCAGCGGCGGCGTGCCACACACTCCTTCGCCGTCTTCGGGAACCCCTTCGCCAGCTAATTATTAA
AA sequence
>Lus10020338 pacid=23144161 polypeptide=Lus10020338 locus=Lus10020338.g ID=Lus10020338.BGIv1.0 annot-version=v1.0
MAPINVLNYFLSLILVLIASTKLQHFAAAGTSFRLVTAASAPYNSPPAPVMVNLTHDDGWYDHPHCGYQSPKETCATGLCCSRFGYCGTKPEYCGKGCQD
GPCQGGSGGVPHTPSPSSGTPSPANY
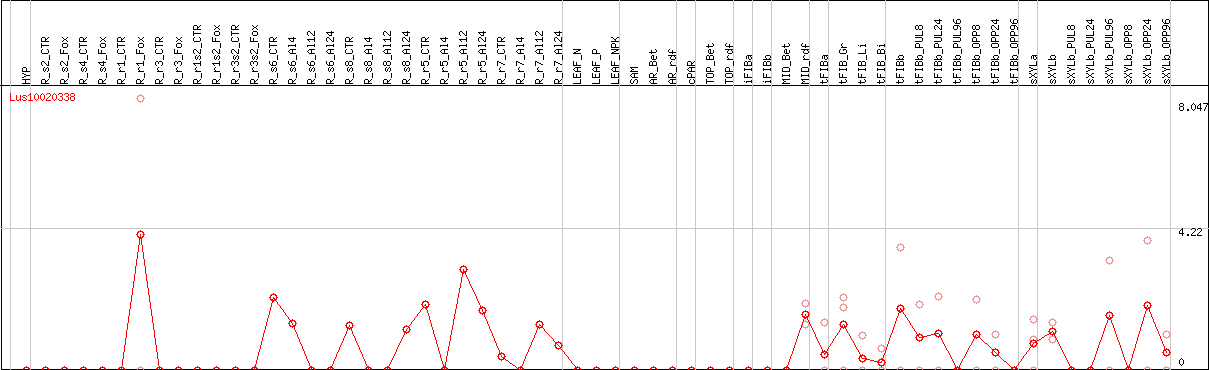
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020338 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.