Lus10020350 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10020350 pacid=23144084 polypeptide=Lus10020350 locus=Lus10020350.g ID=Lus10020350.BGIv1.0 annot-version=v1.0
ATGCATCCGAAGGCTCAAACCAGAGTGGAATCTGGAACGTGTAACGTGTGCTCCACTCCTTGTTCATCTTGTATGCACCGGAAGCGTGCTTTTATGGGAT
CAAAGGGTGATGAGGGTGATGAATTTTCCGACGAAACTTGCCACGTAGCTGAAACAAGTCAGAATTCCATTAACAAGGATGATTCCTCACCTGCCAAAGG
CGGCATGCTAGCCACGTTACAACATGCAACAAGTGCACCTAGTAATGCGCCAAATGTTGATTCATGTCAGGATTCAGTGTCTGAAAGTGCAGAAATTAAA
GGTAATGTGAAGATTTCGGATAGAGTGGACGTCTCAATGAAGTCTGAGGCTCTCCCAGAGTCCTCACACGTCGGATCTGTCTCGTTGGTGGACCGGATTT
CTCCTAAGCCAGAGCTTATTGCATATAAGCGAGCTGCGTCTGTTAAGCTTGCTGCTTCTAAAGCTGTAGAAGGCCATGGTGATAATATATCATGTGTGAC
GGGTGCTGATGTGAGCAGTAACGGAGATTCTGCCAGTAAGGTGCCTGTCCAGAATAACAATCTCTCGTCCAACGCAGATGCTGTAAACTTGAACAAGGAT
CCAGTCGTTCAAGTTTATAAAGAAAGTTCTAAACCCATAAAGGAACATGCAAGCCCAATGTTGTCACAAGAGGCAGCACCAGATATTCTACTCCGGGATA
ACTCGCAAAATGGTGCAGACAACAACCTGCAAAGTAGAAGTGTAACTCCGGGCGATGGTCCAGAGGTTTCAAGTAAACTTGGTCAAGAGATCAAAGAGGA
AACTGGTAATGATAGTCGATCACGTCCAGACAAAGGTATTAGTTCTTCTGTCAAAGGCGAGGAAGATGAGAAGGTTAACGAGTCAGTTGAGCTACCTCAT
ACTCGTGAGTTGCCTGAACAATCTTCATCTGCCAATGATGGTGATGAGTCGGAAATTGTGGAACAAGACGTGAAAGTATGTGATATCTGTGGAGACTCAG
GTCAGGAAGAGAAGCTTGCATTTTGTAGTAGGTGCAGCGATGGTGCTGAACACACATATTGCATGAAGGAAATGCTCCTAGAAGTTCCGGATGGTGATTG
GTTGTGTGAAGAATGCAAGATAGCTGAAGATTCCGAAAATCAGAAGCAAGGTGCAGAGGAAAAAGATGCAAGCAAATTAACTTACCAGAGATTAAAGAAG
AGACCTGCTGAGGCTGTAGAGGTGGCCGCTATTCCCAAAAGGCAGGCTGTTGACATGGATTCAGGTTCGTCGAATCCATCGAGCCCTATCAAAGTGGCTG
CCCTGCAGCGTAACAGCTCCTCCAGGAGTTGGGATAAGGAGAAAGTAAAGCCAAGCCATCCTGTTAATGACATCACAGAAAGTACATATCTGAAGTCCAA
TTCCTTCAACGCAATAAGTTCCAAGCCGAAAGTTAAGCTTGTTGATGAAGTGCCACACAAGCAAAAGAGCTTTAAGGAGAGCTCCTTATCTGACAGTAAG
GATGGGCCTGGCAGAACAATGGGCAAATCTATGTCATTTCATTACGCAGGATCTGGAAAAGCTGAGTCAAAAGTGAAATTGTTACCGTCAAAACTTGATA
GTAAAGGAGTAAAGCAAGCACGAAACACCTTTGAGAGAAGAAGTTTGCCTAAGTTAGACCGTCCTGGTGGTATGGTGGCACCGAGTTCGAAAGTCACAAC
ACCGAAACTTGAACAAAAGCTCACTCGTAACGAAGGTGTCCAATCGTCATCGTCTGGCAACAACAGAGACTTGAATGCCGCTGCTTCTGAGGGAAGGTTG
GGCAGCTTATCCAGATCAAATGCTGGCATTACCCGTAAGGGCGACGGTCCTGGTACTTCTGTTCGATCCTCATCTCTTCATGGAGCAGGCAACGTTTCCA
TTGAAAGAACTACAAATCAAGCTATCCCCAAGGATGAACCCTCACCTAGTACTTCTTCGTCTGCAGAAAGAACACTGGATACCGGTGATGAAAGCTCATA
TGATGGCCTTCGTCCACAGAAATCATTGAGCCTGAATGAGAAAAGCAGGGCGAGTTCTGTTAGTCGCACAAGACCTGCTGGTGCAGTGGGCCTAAAAGAA
GCAGGTACTAGTCGGAAAAGGAAGGATATTGGTCTTTCCGGGGAAAACCCCACATCTGTTAGTCCTCGTGGTCCTTCTAGTGAAGCACCAGGTACTAGAA
GTGTTGGTGATGAAATAAATAGAGACAGTAAACTAAAAGCTGCAATTGAAGTTGCTATGCTGAAGAGACCGGGAATTTATAGAAAGAAGATTAATCGTGA
CCGATCTGACGGGCTATCCTCATCAAATGTGGATGTGAACTCTCCTAAGGATCCACTGGCTAAGTTTTCAGGTTTAAATAGGCTGAAAGTAGCCACTTTT
GACGAACGAGCTCATGAAGAGAAACCAGTCCTTGGCCCTTGGTCAACGGACTCTCCAAAACAATCTGACATGGAAAGTCTTAACCTATCTAATACACACT
CTGTTAGTACTGCATTCCCGTTGAAAGGGGACTTAGACACAGTAAACACTTCTGCTGGAAAATCTGGTCCTATCCCACTTACAACCATTGAGAAAGCGTC
CCCAATTCCAGAGCATGACTTTATCTGGCAGGGAACCTTTGAGGTTAACCGGGGTGGAAAAGTCGCAAATATCCATGGAGGCATTCAAGCACACTTGTCA
ACTTGTGCATCATCCAAAGTTCTTGAAGTGGTAAACAAGTTTCCTGAGAATTTTCATTTAGAGGAAGTATCTCGATCAAGTACGTGGCCGAGACAGTTCC
AAGAAATTGGCGCTACTGGTGATAATATTGCTCTTTACTTCTTTGCCAAGGACCACGAAAGTTACAACAAGCATTACAAGGGCTTATTGGACAATGCGAT
GAAAAGAGATTTGGCCTTCAAAGGATACTTTGATGGTGTTGAACTCTTAATATTCCCATCCACTCAGCTCCCAGAGGATTCACAACGTTGGAACATGATG
TTTTTTCTTTGGGGCGTGTTCAAGGCGAGGAAATCCGGTTCTGATGACTGTCTAAGAAAGCTAGCTGTCCCCAGTTTGAATTTGGTTGCTCAGGGTGAAG
AAATGGCCGTGGCTAATGTATCATCGCCGGCTAATAGGTTCCCACCTCAATATGCAGGGAGAGAAATTTCCGCATTAGATGATCACCGCAACACTGCTGT
TTCATCAAGGGACTTGAAAAACACGCATGGAACAGAAAATGGTGCTGTGGATAAGGGAAAAGTAGACTCTGCGATCAAGGTAGAGCAGCCAGAATGTATA
CTCGACTGTGGTGGAACGACAAGCGTGGCATCGTCAAATCAAGTTCGGAGCAGCAGAGCCATGGCTAGCCCGGTCGACATTGGCCGTCCAGAGTCCAGAA
CGGATACAGAAGAGAAACCATGTGCTCCGGTTGTCGAGTCCAAGAATGGCTTCAATGACAAGGAAGCACAGAAGCAAGTAGACACTTCAAATCTCCGAGC
TGATGCATCTAAACTCGAGAACCTTCAGATCGAGACTCAAAACGTAGGTGTGAGGGGCTGCATTTCTGATCAAGAGCCAGCAGTTACCAGGAAAGACCGT
AGCAATACGGGCGGCGGAGTTAACATCATGAGAGACTTGAACGAAGTTGTTGATATGGAGATCGAAACTTCCGAGGATAGAGATGACATGGCCTCCGACA
AAAGGAAAATGCCGCTCTTGGATCTCTCCAAGCTGCCACCCGAGAGTCCAAGTAGCATCATAGATCATGAAATGCCATGGAGTAACGCAACAGCAAAAAC
TGGGTCTGTAGATGGACAAGGTGCCAACAAGAAGCGGAGAGTGAGCTACGAAGGTTCTGAGTCTTCCCCACTACCTATTGGTTCTTCGGTAGAGAAGAGA
TACAATGAGATGTCGGAGAAGAAAGCCGTTCCTGAAAACTCGAAAGCTCCGGAGTGGTTTTCCTTCCCGGCTTTGGATTCACACAGTGCGTCAATGAGGC
CGAACACCATCCCCTGGGCACGGGGCTCCTCGTCGGACCAACCATCATCGCGTCACGAAATCCCTAACCTTAATCTCGCTTTGGAGGAAGATGAGACAGA
GTCCGACACTGTACCACCCCCTCCATCAAGGAAGGGAATGCTGCCTTTCATAATTGCAACAGTAGACAATCATGCTGCTAGCCAGAACAACTCGTTGCTG
CCGCAGCCACCGCCGGCAAAGGAAGTGAAGGAGAGAGAGCAAGAGGAGAGCGTTTCGGCGGCCCTTTCTCTATCTCTTTCGTTTCCCTTCCCGGATAAGG
AACAACCGACCAAACCACCACCGGAGTATGCTCCCTTTTGGAAGCTCCACAGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10020350 pacid=23144084 polypeptide=Lus10020350 locus=Lus10020350.g ID=Lus10020350.BGIv1.0 annot-version=v1.0
MHPKAQTRVESGTCNVCSTPCSSCMHRKRAFMGSKGDEGDEFSDETCHVAETSQNSINKDDSSPAKGGMLATLQHATSAPSNAPNVDSCQDSVSESAEIK
GNVKISDRVDVSMKSEALPESSHVGSVSLVDRISPKPELIAYKRAASVKLAASKAVEGHGDNISCVTGADVSSNGDSASKVPVQNNNLSSNADAVNLNKD
PVVQVYKESSKPIKEHASPMLSQEAAPDILLRDNSQNGADNNLQSRSVTPGDGPEVSSKLGQEIKEETGNDSRSRPDKGISSSVKGEEDEKVNESVELPH
TRELPEQSSSANDGDESEIVEQDVKVCDICGDSGQEEKLAFCSRCSDGAEHTYCMKEMLLEVPDGDWLCEECKIAEDSENQKQGAEEKDASKLTYQRLKK
RPAEAVEVAAIPKRQAVDMDSGSSNPSSPIKVAALQRNSSSRSWDKEKVKPSHPVNDITESTYLKSNSFNAISSKPKVKLVDEVPHKQKSFKESSLSDSK
DGPGRTMGKSMSFHYAGSGKAESKVKLLPSKLDSKGVKQARNTFERRSLPKLDRPGGMVAPSSKVTTPKLEQKLTRNEGVQSSSSGNNRDLNAAASEGRL
GSLSRSNAGITRKGDGPGTSVRSSSLHGAGNVSIERTTNQAIPKDEPSPSTSSSAERTLDTGDESSYDGLRPQKSLSLNEKSRASSVSRTRPAGAVGLKE
AGTSRKRKDIGLSGENPTSVSPRGPSSEAPGTRSVGDEINRDSKLKAAIEVAMLKRPGIYRKKINRDRSDGLSSSNVDVNSPKDPLAKFSGLNRLKVATF
DERAHEEKPVLGPWSTDSPKQSDMESLNLSNTHSVSTAFPLKGDLDTVNTSAGKSGPIPLTTIEKASPIPEHDFIWQGTFEVNRGGKVANIHGGIQAHLS
TCASSKVLEVVNKFPENFHLEEVSRSSTWPRQFQEIGATGDNIALYFFAKDHESYNKHYKGLLDNAMKRDLAFKGYFDGVELLIFPSTQLPEDSQRWNMM
FFLWGVFKARKSGSDDCLRKLAVPSLNLVAQGEEMAVANVSSPANRFPPQYAGREISALDDHRNTAVSSRDLKNTHGTENGAVDKGKVDSAIKVEQPECI
LDCGGTTSVASSNQVRSSRAMASPVDIGRPESRTDTEEKPCAPVVESKNGFNDKEAQKQVDTSNLRADASKLENLQIETQNVGVRGCISDQEPAVTRKDR
SNTGGGVNIMRDLNEVVDMEIETSEDRDDMASDKRKMPLLDLSKLPPESPSSIIDHEMPWSNATAKTGSVDGQGANKKRRVSYEGSESSPLPIGSSVEKR
YNEMSEKKAVPENSKAPEWFSFPALDSHSASMRPNTIPWARGSSSDQPSSRHEIPNLNLALEEDETESDTVPPPPSRKGMLPFIIATVDNHAASQNNSLL
PQPPPAKEVKEREQEESVSAALSLSLSFPFPDKEQPTKPPPEYAPFWKLHR
|
DESeq2's median of ratios [FLAX]
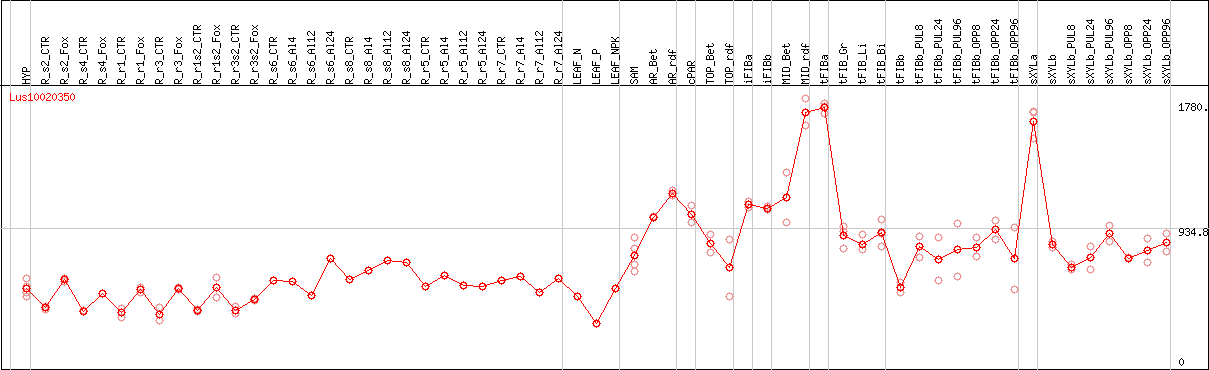
Coexpressed genes
Lus10020350 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
