External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G27610 422 / 1e-142
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G22070 325 / 3e-106
pentatricopeptide (PPR) repeat-containing protein (.1)
AT1G20230 311 / 4e-101
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G09950 315 / 2e-100
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 301 / 2e-100
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT4G02750 309 / 7e-100
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G16835 303 / 4e-99
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G68930 305 / 1e-98
pentatricopeptide (PPR) repeat-containing protein (.1)
AT4G33990 306 / 3e-98
EMB2758
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G49170 302 / 1e-96
EMB2261
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004892
558 / 0
AT2G27610 998 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010998
325 / 1e-106
AT4G16835 778 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10016387
309 / 3e-103
AT4G33990 717 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040577
302 / 1e-101
AT1G25360 577 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10016425
308 / 3e-99
AT2G22070 1002 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10029436
304 / 6e-99
AT3G08820 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10000117
295 / 6e-99
AT4G21065 404 / 5e-139
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10014956
293 / 3e-98
AT5G13230 484 / 1e-165
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019735
305 / 4e-98
AT4G33990 1015 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G184800
432 / 1e-146
AT2G27610 1061 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.015G018700
320 / 3e-103
AT3G49170 1040 / 0.0
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.016G075000
313 / 2e-102
AT2G41080 755 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.016G077700
310 / 5e-102
AT4G30700 489 / 1e-164
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G085500
314 / 8e-102
AT2G22070 1088 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.004G047800
317 / 2e-101
AT4G13650 654 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G027800
313 / 2e-101
AT1G25360 1047 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.006G105700
307 / 3e-100
AT3G08820 810 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.018G040100
308 / 6e-100
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G155100
309 / 9e-100
AT3G61170 933 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10020588 pacid=23172270 polypeptide=Lus10020588 locus=Lus10020588.g ID=Lus10020588.BGIv1.0 annot-version=v1.0
ATGCGGGCTCGGAACGTGGAAATGGATGGTGTGACGTTCATTAGTGTGATATCGGCGTGCACGCACGCTGGATTGATTGAGGAAGGTCGAAAGTATTTCG
ACTTGATGGGGAAGGATTATCGAATCGAGCCGACAATGGAGCATTATTCCTGTTTGGTGGATCTTTATGGTCGAGCTGGGAGGCTCGAAAAGGCAATGGA
TGTCATAAAGATGATGCCTTTTCCACCAGGTGCGACGTTGTGGAGGACGGTTCTCGGTGCAGCCCGTGTGCACCGGAATCTCGAGATAGGCGAGCTCGCT
GCAGAGAATCTTATATCGATCCAGCCAGAGGATTCGGCAGCCTATGTGCTTTTGTCTAATATGCACGCGGTGGCCGGGAACTGGAAGGAGAGAACTAAAG
TAAGGAAGTTGATGGATGTTAGGAATGTGAAGAAGGAAGCCGGGTACAGCTGGATCGAGGTGAAGAACACGACCCACTCATTTTTAGCGGCCGATGTTTC
GCACCCCATGTCGAATTCGATATATGCAAAGCTTAAGGAGCTCGGTTCGAAGTTAAGAGATGTTGGGTATAGACCGGATACTGAAAATGTGCTCCTTGAT
GTGGATGATGAACATATGGAATCGGTTCTTTCGCAGCATAGCGAGAGGCTGGCAATTGCGTTCGGGTTAATTTCGACGCCTCGTTGGACTCCGCTGCGGA
TTATGAAGAATTTGAGGGTTTGTGGGGACTGCCATGCTGTGATTAAGTTGATCTCGGAAATAGAAGGGAGGTACATTGTTGTTAGGGATACTAATCGGTT
CCATCACTTCAAGGAGGGTTCTTGCTCTTGTGGAGATTATTGGTGA
AA sequence
>Lus10020588 pacid=23172270 polypeptide=Lus10020588 locus=Lus10020588.g ID=Lus10020588.BGIv1.0 annot-version=v1.0
MRARNVEMDGVTFISVISACTHAGLIEEGRKYFDLMGKDYRIEPTMEHYSCLVDLYGRAGRLEKAMDVIKMMPFPPGATLWRTVLGAARVHRNLEIGELA
AENLISIQPEDSAAYVLLSNMHAVAGNWKERTKVRKLMDVRNVKKEAGYSWIEVKNTTHSFLAADVSHPMSNSIYAKLKELGSKLRDVGYRPDTENVLLD
VDDEHMESVLSQHSERLAIAFGLISTPRWTPLRIMKNLRVCGDCHAVIKLISEIEGRYIVVRDTNRFHHFKEGSCSCGDYW
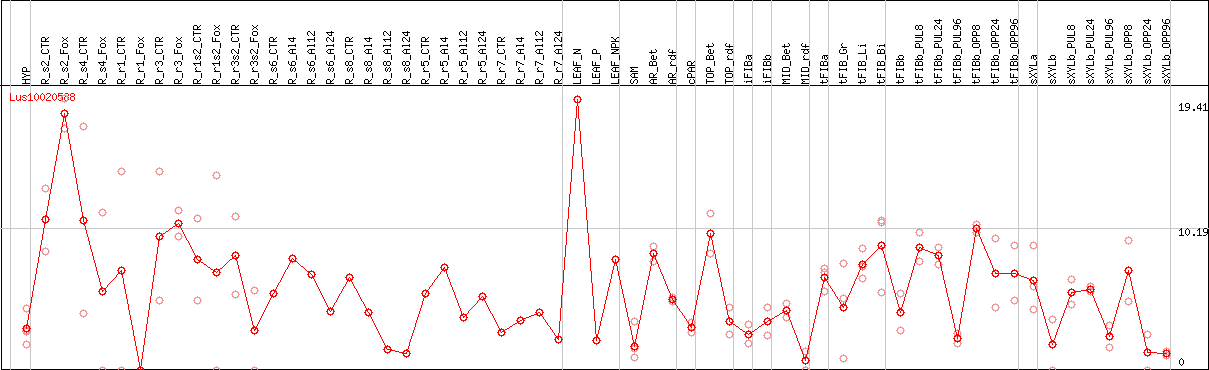
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020588 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.