External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G22380 283 / 8e-97
NAC
ANAC090
NAC domain containing protein 90 (.1)
AT3G44350 258 / 8e-87
NAC
ANAC061
NAC domain containing protein 61 (.1.2)
AT1G61110 144 / 2e-41
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT4G17980 142 / 3e-41
NAC
ANAC071
NAC domain containing protein 71 (.1)
AT2G17040 142 / 4e-41
NAC
ANAC036
NAC domain containing protein 36 (.1)
AT2G02450 143 / 1e-40
NAC
LOV1, ANAC034, ANAC035
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
AT5G46590 141 / 2e-40
NAC
ANAC096
NAC domain containing protein 96 (.1)
AT4G27410 139 / 6e-40
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT3G17730 135 / 5e-39
NAC
ANAC057
NAC domain containing protein 57 (.1)
AT1G32510 135 / 2e-38
NAC
ANAC011
NAC domain containing protein 11 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004846
498 / 0
AT5G22380 275 / 1e-93
NAC domain containing protein 90 (.1)
Lus10029410
179 / 2e-55
AT5G22380 171 / 1e-52
NAC domain containing protein 90 (.1)
Lus10023208
146 / 6e-42
AT2G02450 321 / 3e-108
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10008420
145 / 2e-41
AT2G02450 320 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10008419
147 / 3e-41
AT2G02450 327 / 7e-109
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10003367
147 / 1e-40
AT2G02450 333 / 3e-110
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10008897
142 / 1e-40
AT2G02450 315 / 5e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10026617
140 / 2e-40
AT1G69490 298 / 1e-101
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10030446
140 / 3e-40
AT1G69490 302 / 2e-103
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10020643 pacid=23172384 polypeptide=Lus10020643 locus=Lus10020643.g ID=Lus10020643.BGIv1.0 annot-version=v1.0
ATGGCTGGTCAGTTCCCACCTGGCTTTCGCTTCTTCCCAACTGAGGAAGAACTGGTCTCTTTCTACCTCCATAAACAGCTCGAAGGGACGATCCAGGAGG
GCTTGCACCAGGTTATACCTGTCATCGATATTTACGAGATCGACCCATGGCACCTTCCAAGCGTAGCAGGAGAGAGGTGTCGAGGAGATACAGAGCAGTG
GTTCTACTTCACGCCAAGACAAGAGCGAGAAGCTCGAGGAGGGAGGCCGAATCGAACAACCGGATCAGGGTACTGGAAGGCCACCGGTTCCCCGAGCTAC
GTCTATTCCTCTAATAATAGAGTGATTGGGATGAAGAAGACCATGGTGTTCTACAAGGGGAAAGCTCCAGCTGGAAGGAAAACCAAATGGAAGATGAATG
AGTACAGGGCTATAGATGGAGGACTTTCGGAGTCATCTAGTCCGCCCATTCCTCAGTTGAGGCGTGAGTTCAGCCTGTGCCGAATCTACGTGGTATCAGG
GACCTTTGGAGCATTTGACAGACGCCCACTGGAAGCTGCAACGAGTACTAGTATTGGCGCTCAGGTTCCGAGTGATGTGCCATCATCAAGTGCTCAAGTA
CCTGCTGCAGTGGTGGAGAGAACTAACTCATCTGAAACTTCCAACTCGGGAGGAGACCATGGCTTCTCCCAAGGTGCTGGAATCGCGGACATGGAGATTG
CTGATTGTCTGGATGAACAATTCATCGACTGGGAACAACTGGACTGGCCATGA
AA sequence
>Lus10020643 pacid=23172384 polypeptide=Lus10020643 locus=Lus10020643.g ID=Lus10020643.BGIv1.0 annot-version=v1.0
MAGQFPPGFRFFPTEEELVSFYLHKQLEGTIQEGLHQVIPVIDIYEIDPWHLPSVAGERCRGDTEQWFYFTPRQEREARGGRPNRTTGSGYWKATGSPSY
VYSSNNRVIGMKKTMVFYKGKAPAGRKTKWKMNEYRAIDGGLSESSSPPIPQLRREFSLCRIYVVSGTFGAFDRRPLEAATSTSIGAQVPSDVPSSSAQV
PAAVVERTNSSETSNSGGDHGFSQGAGIADMEIADCLDEQFIDWEQLDWP
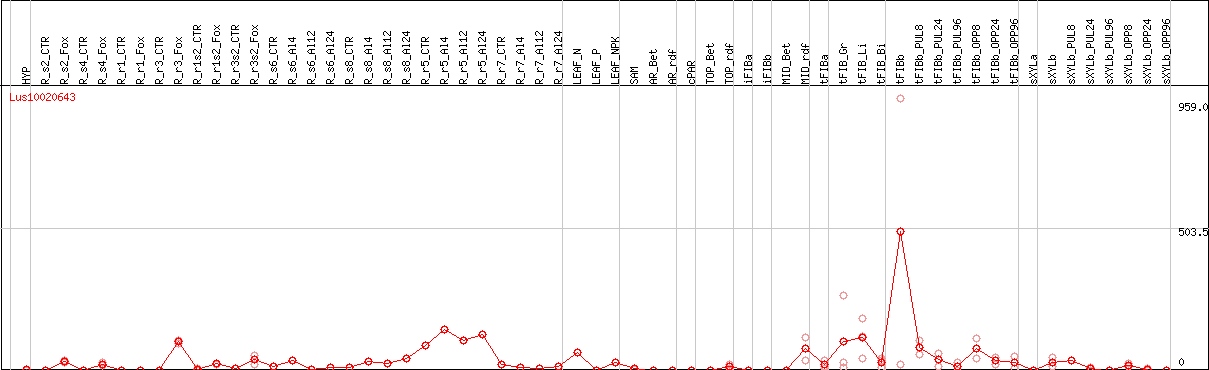
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020643 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.