External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G37020 184 / 9e-55
ARF
ARF8, ATARF8
auxin response factor 8 (.1.2)
AT1G30330 152 / 3e-43
ARF
ARF6
auxin response factor 6 (.1.2)
AT1G19220 115 / 2e-30
ARF
IAA22, ARF11, ARF19
indole-3-acetic acid inducible 22, AUXIN RESPONSE FACTOR11, auxin response factor 19 (.1)
AT5G20730 114 / 4e-30
ARF
MSG1, IAA21, BIP, ARF7, IAA25, IAA23, TIR5, NPH4
MASSUGU 1, indole-3-acetic acid inducible 25, indole-3-acetic acid inducible 23, indole-3-acetic acid inducible 21, BIPOSTO, AUXIN RESPONSE FACTOR 7, Transcriptional factor B3 family protein / auxin-responsive factor AUX/IAA-related (.1.2.3)
AT1G19850 108 / 7e-28
ARF
IAA24, ARF5, MP
MONOPTEROS, indole-3-acetic acid inducible 24, AUXIN RESPONSE FACTOR 5, Transcriptional factor B3 family protein / auxin-responsive factor AUX/IAA-related (.1)
AT5G60450 84 / 2e-19
ARF
ARF4
auxin response factor 4 (.1)
AT1G59750 75 / 3e-16
ARF
ARF1
auxin response factor 1 (.1.2.3.4)
AT5G62000 74 / 5e-16
ARF
ORE14, HSS, ARF1-BP, ARF2
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
AT4G23980 69 / 4e-14
ARF
ARF9
auxin response factor 9 (.1.2)
AT3G17600 66 / 4e-14
AUX_IAA
IAA31
indole-3-acetic acid inducible 31 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10007386
303 / 1e-99
AT5G37020 1021 / 0.0
auxin response factor 8 (.1.2)
Lus10010969
248 / 1e-78
AT5G37020 1013 / 0.0
auxin response factor 8 (.1.2)
Lus10031354
244 / 3e-77
AT5G37020 1013 / 0.0
auxin response factor 8 (.1.2)
Lus10020804
173 / 1e-50
AT5G37020 913 / 0.0
auxin response factor 8 (.1.2)
Lus10017640
149 / 4e-42
AT5G37020 762 / 0.0
auxin response factor 8 (.1.2)
Lus10033597
149 / 5e-42
AT1G19220 842 / 0.0
indole-3-acetic acid inducible 22, AUXIN RESPONSE FACTOR11, auxin response factor 19 (.1)
Lus10017641
145 / 5e-41
AT1G30330 810 / 0.0
auxin response factor 6 (.1.2)
Lus10022943
118 / 3e-31
AT1G19220 546 / 4e-177
indole-3-acetic acid inducible 22, AUXIN RESPONSE FACTOR11, auxin response factor 19 (.1)
Lus10027600
117 / 3e-31
AT1G19220 670 / 0.0
indole-3-acetic acid inducible 22, AUXIN RESPONSE FACTOR11, auxin response factor 19 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0072
Ubiquitin
PF02309
AUX_IAA
AUX/IAA family
Representative CDS sequence
>Lus10020805 pacid=23140829 polypeptide=Lus10020805 locus=Lus10020805.g ID=Lus10020805.BGIv1.0 annot-version=v1.0
ATGCCTTTAAGCGTGGCTGGGTTTCAGAATTCTCTCTACAGTTCTTCTGAGTTCTTACCCGGTACAGGAGGGCAAGTAGAGGCGCTAAACGCAGGTCCAA
TATTTGTGAAGGTTTACAAATCTGGATCGGTTGGGCGGTCCCTTGACATCTCCCGGTTCAGCAGCTATCATCAGCTGCGGGAGGAACTGGCACAGATGTT
CGGGATCGAGGGGAAGTTGGAAAACCCTCATAGATCAGGCTGGCAGCTTGTATTTGTCGACAAGGAGGATGATGTTCTTCTCCTTGGAGACGGTCCTTGG
GACGCATTTGTAAACAACGTGTGGTATATCAAGATACTTTCTCCGGAGGATGTAGTGAAGATGGGGGATCAAGGGGTTGAATCGTTCAGCCCGAATACAC
GCCAAAGAACGATCAGCAGCAGGGAACCGGACTTTAGCTCGGGTTCACTTAGATACTGA
AA sequence
>Lus10020805 pacid=23140829 polypeptide=Lus10020805 locus=Lus10020805.g ID=Lus10020805.BGIv1.0 annot-version=v1.0
MPLSVAGFQNSLYSSSEFLPGTGGQVEALNAGPIFVKVYKSGSVGRSLDISRFSSYHQLREELAQMFGIEGKLENPHRSGWQLVFVDKEDDVLLLGDGPW
DAFVNNVWYIKILSPEDVVKMGDQGVESFSPNTRQRTISSREPDFSSGSLRY
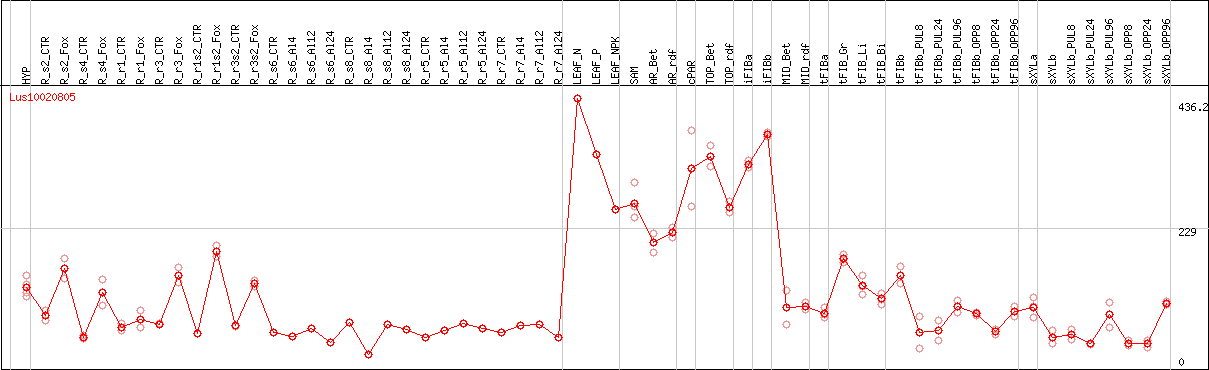
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020805 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.