Lus10020813 [FLAX]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Poplar homologues |
|
||||||||||
| PFAM info | |||||||||||
|
Representative CDS sequence |
>Lus10020813 pacid=23140801 polypeptide=Lus10020813 locus=Lus10020813.g ID=Lus10020813.BGIv1.0 annot-version=v1.0
ATGGAGGCCGACGACGAAGAGAGGATGCAACTTTGCCGTAAAATCGCCAAAATCCTCGACGAAACAAAGACATCAAACGCCACTCACATCCGAAAGCTGA
AAGACCTCTCTGCCGTCCTGTCAAAGTCCCCGTCTCCTCGCCAGTTCGCGGCTGCATTCTGCAGGACTCTGACTCCTCTCTTCTGCATCCAGCGCCGGAC
TTCCTCCGCCGAGCGCGTTGTCAGGTTTGTCTCCACCTTCGCCGGCGCCCAGGCGTCTGGTAGAGACTCCGGCTGCGATGAATTTCTCGAGGAGTTGCTC
AGGTTCTTACTAACTGCTTCTGCCGCCGCCAATAAGACCGCCAGGTTTAGGGCCTGTCAGATGATATCTGAGATAATTATGCGGCTGCCTGATGATGCCG
AAGTAAGTGATGATGTATGGGATGAGGTTATTGAATGTATGATGTTGCGTGTTCGAGACAAGGCTTCTGTTATTTGTACATTTGCTGTTAGAGCTCTTTC
CAGATTCGTAAACGATGCTGAGAATAGTGACATCCTTAATATGCTTCTGGAAGTGCTTGAACTGGAGCAAAGTGCAGAAGTTCGTAAAACAATTCTGTTA
GCTTTGCCTCCTACAAATGCAACCTCACTTGTGATTGTTGATTGCACCTTAGATGTGAGTGAATCAGTTCGCAAAGCTGCATACTCTGTTCTTGCTAACA
AATTTCCACTGCATAGTCTCAGCATTAAGCTGAGGACACTGATTCTTCAAAGAGGACTTGCTGATCGTTCCATCATTGTTTCAAAGGAGTGCCTAAAGCT
GCTGAGAGATGAGTGGCTTTCTAAGTGCTGCAATAATGATCCTACAGAACTTCTAAAGTATTTGGATGTTGAGACTTATGAGTTGGTTGGAGAAGCTGTG
ATGGATGCTTTGCTGAAAGAAGGTTCAATTTTATTTGATAATCAGAGCATCCAAGAGTACATTGTATATACCGCTGGTGAAAATGAAGATTCACCTCCAA
ATATTCAGTTGATGGAACCAGAGTCGGCTCTTTATTGGAAAACTGTGTGCAGGTTCTTACAGAGCAAGGCACAAGCAAAAGGCTCTGATGCTGCTGTAAC
AACGGGTACTGAAGCTGCAGTTTATGCCGCTGAGGCTTCAGACAGTAATGACCTTCTCGACAGAATACTTCCAGCTACAGTTTTTGATTATGTAGCTCTG
GTCAAAGCTCATATAGATGCTGGAGCAAACTTCCGTTTTGCATCTCGACAGCTGCTGCTACTTGGTACCATGGTGGATTTTTCTGATTCCACATGCAGAA
AGGTAGCTAGTGCTTTCCTTCAAAACCTGTTGCAGAGGCCCTTAGATTTTGAAACTGATGAGACTGGAAATGAAGTCGTCATTGGAGATGGCTTAAACCT
GGGTGGTGAAAAAGATTGGGCTGACGCTGTAACCAGCTTGGCAAAACAAGTTCATTCAGGAATTGGAGAGATTGAAAGTGCTTTTCTTGTGGTTCTAGAA
GAACTTGTCAGGCCTTGCCGGGAGAGGACAGCTGATTTTAAGCAGTGGATGCATTGCCTTTCTGTCACAGGTCTTCTTTTGCAAAATACAAAGTCGTTTC
ACCTGCTTCAAGGACAAGCTATCCAACCATCTGAATTGCTGCAGTCTTTGTTACTCCCTGCGGCAAAACATGTTCACTTAGATGTGCAGAGAGTTTCGAT
TCGATGCCTTGGTCTTTTTGGCTTGTTGGAGAAGAAGCCAAGTGAAGATGTGCTGAAGCAGTTAAGGGTTTCCTTTATAAGGGGTCCTTCTCCTATCAGT
GCAATGGCATGCAAAGCTGTAATGGACCTTGCTCTCTGGCATGGTCCTCATAACGTCGATAGTGCCCTGGGAAGCGGTGATGCATCTCCATCGGAGGGGA
TCAAAATGGAATTCAATCCTGTGAACTTCCTTGATGCTGATGATAATACCGATGTCGGGTTTCTAGACCTTTTGTATGCCGGTCTTGACAGAAATGATTG
GGGGGAAGCAGTAGAAAGTGATGAAAGCGAGATTGTGCGAGCTGTTCTAGGTGAAGGATTGGCGAAGATTCTTCTCCTCAGTGACAATTATACAACTATA
CCGTCTTCTTTACACCCTCTACTCTTTGCGAAGCTTATTGAACTATATTTTAGTGACCAAACTAATGACTTGCAGAGGCTAAAACAGTGTTTATCTGTTT
TCTTTGAGCACTACCCTACCATTTCAGCTAATCATAAGGCAATTATGCACGCAATGTGGCCAGCTGTTTTTGGAAATTCTGGAGGAAGTTCTGTTGTGGT
ATCTAACTTGCGAAAACGTGCAATCCAAGCATCACGGTTTATGCTACAGATGATGCAGGCTCCATTGTATGTCTTAAACAATGAGACTGAGAACGGAGAG
AGTGGTACTGGATCAGCAGAAAATACGAATCAAACGCCAGAGCCTACACTAGAATGTAGAGAGGAAGGACTCGCAGTGCGCATAGCCACAGAGTTGGCCA
GCTTCACAGGGAAGAGAACAGCTGCAGAGAAGGCATACATATCTGCACTCTGCAGAATACTCGTCCTGATCCAGTTTAGGATATCAGGGCAACACACAAT
AAAGCTAATGAGACGACTTCTGAATCAACTGATTGGAAATGTACCACTAGATAAAGATGTCATGAAAGAACTGAAGCTGATGGCTGAACGTCTAAGTTCA
ATGGATAGTCAACCAGAGGAGGAACTACCCCAAGATCAGGCCAATCTTATATTCGGTATACTGGAGGTGGAATTCAACTTGGACGCCAACATTCCTGCTG
CTAGTGCTGCTCCTAGTTCTACAAACCAGCAGACGCCAGATTGTCGAGCGAGCCGACCAGTCCGGCCGAGGAGGAGAGCAAAGGACGACGAAGAAACCTC
GTCGGAGGAAGAAACTTCATCGGTACCAGTCCCGGTAGCAGCTGCTGGCAGTAGCAGGTCGCAAAGGGCGAGCAAGACTGCTGCATTGTCCAAGCTCCAG
GGAGTAACAGGTCGCAAGAAAGCTGCTGCGGCTAGGATTGAGGAGGAAGAAGAAGAAGACGAAGAAGATGATGAGTCAGACGTGACGTCAGAAGATGCTG
ATGATGACAGTGATGACGGCTTGATCTGA
|
||||||||||
|
AA sequence
|
>Lus10020813 pacid=23140801 polypeptide=Lus10020813 locus=Lus10020813.g ID=Lus10020813.BGIv1.0 annot-version=v1.0
MEADDEERMQLCRKIAKILDETKTSNATHIRKLKDLSAVLSKSPSPRQFAAAFCRTLTPLFCIQRRTSSAERVVRFVSTFAGAQASGRDSGCDEFLEELL
RFLLTASAAANKTARFRACQMISEIIMRLPDDAEVSDDVWDEVIECMMLRVRDKASVICTFAVRALSRFVNDAENSDILNMLLEVLELEQSAEVRKTILL
ALPPTNATSLVIVDCTLDVSESVRKAAYSVLANKFPLHSLSIKLRTLILQRGLADRSIIVSKECLKLLRDEWLSKCCNNDPTELLKYLDVETYELVGEAV
MDALLKEGSILFDNQSIQEYIVYTAGENEDSPPNIQLMEPESALYWKTVCRFLQSKAQAKGSDAAVTTGTEAAVYAAEASDSNDLLDRILPATVFDYVAL
VKAHIDAGANFRFASRQLLLLGTMVDFSDSTCRKVASAFLQNLLQRPLDFETDETGNEVVIGDGLNLGGEKDWADAVTSLAKQVHSGIGEIESAFLVVLE
ELVRPCRERTADFKQWMHCLSVTGLLLQNTKSFHLLQGQAIQPSELLQSLLLPAAKHVHLDVQRVSIRCLGLFGLLEKKPSEDVLKQLRVSFIRGPSPIS
AMACKAVMDLALWHGPHNVDSALGSGDASPSEGIKMEFNPVNFLDADDNTDVGFLDLLYAGLDRNDWGEAVESDESEIVRAVLGEGLAKILLLSDNYTTI
PSSLHPLLFAKLIELYFSDQTNDLQRLKQCLSVFFEHYPTISANHKAIMHAMWPAVFGNSGGSSVVVSNLRKRAIQASRFMLQMMQAPLYVLNNETENGE
SGTGSAENTNQTPEPTLECREEGLAVRIATELASFTGKRTAAEKAYISALCRILVLIQFRISGQHTIKLMRRLLNQLIGNVPLDKDVMKELKLMAERLSS
MDSQPEEELPQDQANLIFGILEVEFNLDANIPAASAAPSSTNQQTPDCRASRPVRPRRRAKDDEETSSEEETSSVPVPVAAAGSSRSQRASKTAALSKLQ
GVTGRKKAAAARIEEEEEEDEEDDESDVTSEDADDDSDDGLI
|
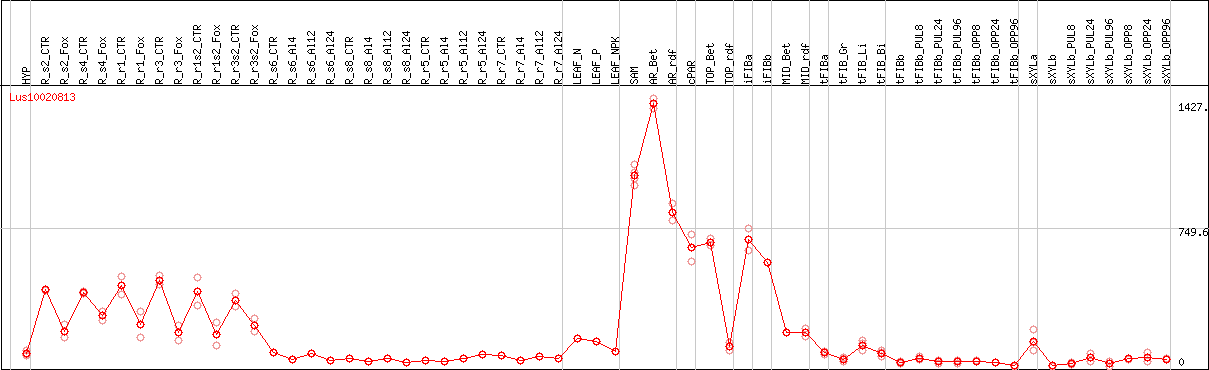
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020813 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
