Lus10020835 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10020835 pacid=23177938 polypeptide=Lus10020835 locus=Lus10020835.g ID=Lus10020835.BGIv1.0 annot-version=v1.0
ATGCTGACCAAAGCCCTCCTACTCGTTTTCCCACTCCTCCTCCTCCTCCTCTCATCGGCGGCGAGTCAAGACAGCCGGAGTCTCCTCACCGCCTTCAAAT
CCTCCTTACCCCACCCACCGCCGCCACTCCTCTCCAACTGGGTACCGAACCAAGACCCTTGTACCTTCAATGGAGTTAAATGTCATGACAACAACGCAGT
CTCCTCCATTGACCTAAGCTTCGTCCCTTTGAGCACCGATTTCAAATCCGTGGCGTCTTTTCTTCTCTCCATCGACTCCCTGGAAACGCTATCTCTCCAA
TCAGCTAATATATCCGGCTCCTTATCTTCCGGATCTTCTTCCAAGTGCAGCTCTGTTTTGACCGGATTAGATCTCTCCCTCAACTCCCTCTCTGGTCCTA
TCTCTGATATTGCTGCTCTCTCTTCTTGCTCGGGTTTAAAAGCTTTGAATCTCTCCAGGAACGCTCTTGATTTCTCTGTTAAGGGAAAATGGAGCGGCCT
CAAGCTTGGATTGTTGGAGTCGTTGGATCTTTCCTGGAACAAGATTTCAGGCTCCAACGCTATCTCCTTCGTCCTGCTCGGAGGCTGCAAAGACTTGAAG
CTTCTGACGATGAGAGGGAACAAAGTCGCCGGCGACGTCGACGTTTCCGGTTGCTCGAGTCTCGAGTATTTCGATATATCGTCGAATAACTTCTCCGGCT
CTGCTCTTCCGAACTTCGGCGGCTGCTCTTCTCTCCAACACCTCGACATCTCCCGGAACAGATTCTCCGGCGATCTCGTCCGGTCAATCGGTTCGTGTTC
AAAGCTCAGTTCCCTCAACGCCTCCGCTAATCTCTTCTCCGGCGAAATCCCGGCGGGACTTCCGACGGGGAATTTGCAGTTTCTATCGTTATCCGGCAAC
AATTTCCAAGGAAAAATCCCTCCGGAGATCACCGATGCTTGTCGAGGTCTCGTAATCCTCGACCTCTCTTACAACAATCTCTCCGGCGAGGTTCCGCCGG
CGTTGACTTCCTGCACTTCCCTGCAGACTCTCGACATTTCCGTGAACAACTTCACCGGCGAAATCCCTATTCCCGTCCTGCTAAAAATGGTAAGTTTGAA
GAGAATACTTCTTTCTTACAACAATTTCGTTGGTGAGCTGTCTGATTCTTTTTCGGCGATGATGAATTTGGAGTCGTTAGATGTGAGTTCCAACGATTTG
AAAGGTCAAATCCCCACCGGAATCTGTCAAAACCCTAACGGAAACTTGAAAGAGATCTTCTTCCAGAACAATCAGTTCACCGGCTCTATACCTGCTAGTT
TAAGCAACTGTACACAGTTAACTTCACTTCATTTGAGCTTCAATTACCTCACAGGAACAATTCCTCCAAGCTTGGGATCCTTAACCAAGCTCCAGGATTT
GAAGCTCTGGTTCAACCAGCTTCACGGAGAGATCCCTCTGGAGATAATGTCGATTCCGAATTTGCAGACATTGATTCTCGACTTCAACGAGCTTACCGGC
TCAATACCTTCAAGCATAAGCAACTCAACTAATCTGAACTGGATTTCATTATCCAACAACAAGCTGCACGGCGAAATCCCTTCTTCTATTGGCAAGCTCG
ATAGCCTCGCAATCTTGAAGCTCAGTAACAACTCGCTCCAAGGTCCGATTCCAGATGAGCTTGGGGACTGCAAGAGCTTGATTTGGCTCGATTTGAACAC
CAACGAGCTAACGGGTTCGATACCGTCCAATCTGTTCAAGCAGTCTGGGAAAATTGCTGTGAATTTCATCACGGGGAAGAGGTTTGTGTTCTTGAAGAAT
GAGAAGAGTGATTACTGCCATGGAACCGGGAATCTGCTCGAGTTTGCCGGGATAAGGCCCGAAAACCTCGATCGAGTCTCGGCCAGGCATCCGTGCAACT
TCACCAGGGTGTATGGAGGGCATACTCAGCCTACATTCAACGACAACGGTTCGATGATCTTTCTCGATCTCTCTTACAATCTGCTGACCGGTACGATTCC
TAAGGAGATTGGGACAATGTCATACTTATACATTATGAATTTCGGACACAACAATTTATCTGGAAACATACCTCAGGAGCTTGGCGACCTGAAAGGTATC
AACATTTTGGACCTTTCGAGCAATAGCCTCGGAGGGACTATTCCGCAGTCGATGACTGGACTTTCCTTGTTGACGGAGATCGATCTTTCGAACAACCATC
TCACCGGGACGATTCCTGAAATGGGACAGTTCGAGACGTTCCAAGATCGTAGCTTTTTAAACAACTCCGGCCTCTGCGGAATCCCTCTTCCTCCCTGTGG
CTCGCCCTCTCCGAACTCTCCGCAAAAGTCTCACCGGAGGAGACAAGCGTCTCTCATCTGGAGCGTTGCGATGGGGCTTTTGTTCTCGTTTTTATGCATC
TTCGTTGCGTTTTTCATTTCCTCGGAGGTCAAGAGAAGGAAGAGGAAGAAGGAGGAAGCTAATCTTGATGTGTTCATGGAGAATCATAATAATTCGCAGT
CCGGGAATGCTAATACGAGCTGGAAGCTGACTGGTGCGAGGGAAGCGTTGAGCATAAACCTTTCGACATTCGACAGGCCACTTCGGAGGCTGCCATTCGC
GGATCTTCACCAAGCTACAAACGGCTTCCACAACGATAGTCTTATCGGTTCCGGAGGGTTTGGAGATGTGTACAAGGCTCAACTGAAAGACGGCAGTATT
GTAGCAATCAAGAAGCTTATACAGATAAGCGGACAGGGCGATAGAGAATTCACCGCCGAAATGGAGACCATCGGTAAGATAAAGCACCGAAACCTCGCCC
CGTTACTCGGATACTGCAAAGTGGGGGAAGAGAGGCTGTTGGTGTATGAGTACATGAAGTATGGAAGCTTGGAAGATGTTCTCCACGACCAGACGAAATC
CGGGATAACACTTAACTGGAATGCGAGACGGAAGATCGCCATCGGTGCTGCGAGGGGGTTAGCATTCCTTCACCACAACTGCAGTCCACATATAATCCAC
CGAGACATGAAATCGAGCAATGTACTTCTCGACGAGAATTTGGAACCGCGAGTTTCCGATTTTGGGATGGCGAGGCTTATGAGTGCCATGGACACACATC
TAAGTGTGAGCACATTGGCAGGGACGCCCGGATACGTCCCTCCAGAATACTACCAAAGTTTTCGATGTTCGGTGAAAGGCGACGTGTACAGCTATGGAGT
TGTGTTGCTCGAACTGTTAACAGGGAGGAGGCCTACAGATTCGGCGGATTTCGGAGACAATAATTTAGTAGGGTGGGTGAAGCAGCATGCCAAGTTAAGA
ATAAGCAACGTGTTCGATCCACTGTTAATGAAAGAGGATCCGGGTTTGGAAATAGAGCTTTTGCAGCACTTGAAAGTGGCCTGTGCTTGTCTCGAGGATA
GGCAATGGAAGCGGCCGACGATGATTCAAGTGATGGCGATGTTTAAGGAGATCCAAGCCGGGTCGGGGCTGGATTCGCAGTCGACTATTGGTAAAGAAGA
TGATGATGGAGATTTTGTTGCGGTAGATATGATAGAGATGACTATAAAAGAAGTCCCTGAAGCAGGCTGCAAAGAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10020835 pacid=23177938 polypeptide=Lus10020835 locus=Lus10020835.g ID=Lus10020835.BGIv1.0 annot-version=v1.0
MLTKALLLVFPLLLLLLSSAASQDSRSLLTAFKSSLPHPPPPLLSNWVPNQDPCTFNGVKCHDNNAVSSIDLSFVPLSTDFKSVASFLLSIDSLETLSLQ
SANISGSLSSGSSSKCSSVLTGLDLSLNSLSGPISDIAALSSCSGLKALNLSRNALDFSVKGKWSGLKLGLLESLDLSWNKISGSNAISFVLLGGCKDLK
LLTMRGNKVAGDVDVSGCSSLEYFDISSNNFSGSALPNFGGCSSLQHLDISRNRFSGDLVRSIGSCSKLSSLNASANLFSGEIPAGLPTGNLQFLSLSGN
NFQGKIPPEITDACRGLVILDLSYNNLSGEVPPALTSCTSLQTLDISVNNFTGEIPIPVLLKMVSLKRILLSYNNFVGELSDSFSAMMNLESLDVSSNDL
KGQIPTGICQNPNGNLKEIFFQNNQFTGSIPASLSNCTQLTSLHLSFNYLTGTIPPSLGSLTKLQDLKLWFNQLHGEIPLEIMSIPNLQTLILDFNELTG
SIPSSISNSTNLNWISLSNNKLHGEIPSSIGKLDSLAILKLSNNSLQGPIPDELGDCKSLIWLDLNTNELTGSIPSNLFKQSGKIAVNFITGKRFVFLKN
EKSDYCHGTGNLLEFAGIRPENLDRVSARHPCNFTRVYGGHTQPTFNDNGSMIFLDLSYNLLTGTIPKEIGTMSYLYIMNFGHNNLSGNIPQELGDLKGI
NILDLSSNSLGGTIPQSMTGLSLLTEIDLSNNHLTGTIPEMGQFETFQDRSFLNNSGLCGIPLPPCGSPSPNSPQKSHRRRQASLIWSVAMGLLFSFLCI
FVAFFISSEVKRRKRKKEEANLDVFMENHNNSQSGNANTSWKLTGAREALSINLSTFDRPLRRLPFADLHQATNGFHNDSLIGSGGFGDVYKAQLKDGSI
VAIKKLIQISGQGDREFTAEMETIGKIKHRNLAPLLGYCKVGEERLLVYEYMKYGSLEDVLHDQTKSGITLNWNARRKIAIGAARGLAFLHHNCSPHIIH
RDMKSSNVLLDENLEPRVSDFGMARLMSAMDTHLSVSTLAGTPGYVPPEYYQSFRCSVKGDVYSYGVVLLELLTGRRPTDSADFGDNNLVGWVKQHAKLR
ISNVFDPLLMKEDPGLEIELLQHLKVACACLEDRQWKRPTMIQVMAMFKEIQAGSGLDSQSTIGKEDDDGDFVAVDMIEMTIKEVPEAGCKE
|
DESeq2's median of ratios [FLAX]
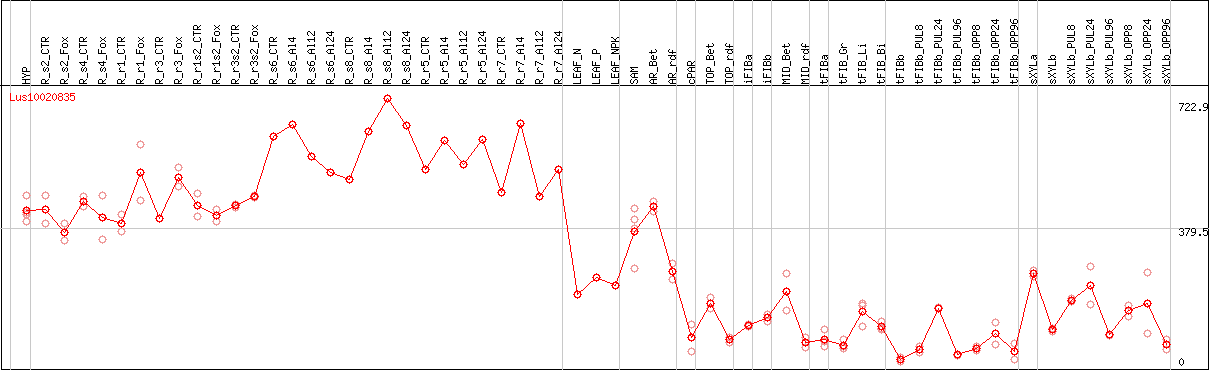
Coexpressed genes
Lus10020835 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
