External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G79460 129 / 2e-34
ATKS1, ATKS, GA2
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
AT1G70080 49 / 1e-06
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
AT4G16740 43 / 7e-05
ATTPS03
terpene synthase 03 (.1.2)
AT2G24210 43 / 8e-05
AtTPS10
terpene synthase 10 (.1)
AT3G25810 43 / 0.0001
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
AT4G16730 43 / 0.0001
AtTPS02
terpene synthase 02 (.1)
AT1G48820 43 / 0.0001
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
AT5G48110 42 / 0.0002
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
AT4G15870 42 / 0.0002
ATTS1
terpene synthase 1 (.1)
AT3G29110 41 / 0.0004
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029033
296 / 1e-95
AT1G79460 662 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10014673
218 / 2e-66
AT1G79460 650 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10012880
205 / 2e-61
AT1G79460 677 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10006923
182 / 9e-54
AT1G79460 538 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10004906
157 / 3e-44
AT1G79460 531 / 1e-178
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10004443
144 / 2e-39
AT1G79460 564 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10016598
140 / 6e-38
AT1G79460 776 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10034229
132 / 2e-35
AT1G79460 583 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10034232
125 / 1e-32
AT1G79460 569 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G082400
157 / 4e-44
AT1G79460 816 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Potri.008G082700
156 / 6e-44
AT1G79460 818 / 0.0
GA REQUIRING 2, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE 1, ARABIDOPSIS THALIANA ENT-KAURENE SYNTHASE, Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Potri.011G142800
63 / 2e-11
AT5G23960 405 / 2e-135
terpene synthase 21 (.1.2)
Potri.015G085500
56 / 4e-09
AT5G23960 456 / 1e-155
terpene synthase 21 (.1.2)
Potri.015G032100
56 / 6e-09
AT5G23960 444 / 8e-151
terpene synthase 21 (.1.2)
Potri.017G041700
56 / 6e-09
AT3G25810 432 / 3e-145
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Potri.007G119700
55 / 9e-09
AT4G16740 438 / 8e-147
terpene synthase 03 (.1.2)
Potri.007G118600
48 / 2e-06
AT4G16740 462 / 3e-157
terpene synthase 03 (.1.2)
Potri.004G037900
45 / 3e-05
AT1G61120 782 / 0.0
TERPENE SYNTHASE 4, GERANYLLINALOOL SYNTHASE, terpene synthase 04 (.1)
Potri.019G022338
43 / 0.0001
AT3G25830 483 / 8e-165
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0613
Terp_synthase
PF03936
Terpene_synth_C
Terpene synthase family, metal binding domain
Representative CDS sequence
>Lus10020865 pacid=23178004 polypeptide=Lus10020865 locus=Lus10020865.g ID=Lus10020865.BGIv1.0 annot-version=v1.0
ATGAGACTGGCCAAGGGGAGGATGGGATATTGTATATTCACCGTGGCAGCGACTTGCTTCGAGCCTAAGCACTCCCACGCCCGCATCTCCTGGGCAAAAC
ACAGTCTTCTTACTGCTCTCATCGATGAAATTTATGACATGTATGGAACTCCAGAGGAACAGCTTAACCTTGTTCATCTTCAAGAGAGATGGGATATTGC
TAGCCCTAAGGTAGATTTCTGTTCGGAGCGAGTAGAAAGCTTATATTGGGCACTTCACAACGCCATATGCGAGACTGCCGCAAAGGTATCATTTGCTTTA
GGACCAATTGTTCTGCCAACCCTTTACTTGCTAGGACCTAAGCTCTCCGATGAAGTTGCCAAGGGGCCAGAAATCCGAAACATGTTCAACGTTGTGACAG
AAGACTGTGATGAAGAAATCAAAGAGCTAGCAGAAGATCTGAGGAAACAACTGTCGAGATTAGAGTTGGATGAACAGAGTAGTAGCAGTATAGTTCCAAA
GGAATGCAGGGTATTGTTTCGCAAAATGAACGATGCGCTGCACTGTTCTTACAAGAAAGAAGACGGGTTCGCGTCCCTTCAATTGGGCCGAGACCTTGAG
TTTTTGATTGAGAAACCCATCCCACTTAGCGATAGAGTTTTGAAGGATGAGTATGGATATGTTAAATGA
AA sequence
>Lus10020865 pacid=23178004 polypeptide=Lus10020865 locus=Lus10020865.g ID=Lus10020865.BGIv1.0 annot-version=v1.0
MRLAKGRMGYCIFTVAATCFEPKHSHARISWAKHSLLTALIDEIYDMYGTPEEQLNLVHLQERWDIASPKVDFCSERVESLYWALHNAICETAAKVSFAL
GPIVLPTLYLLGPKLSDEVAKGPEIRNMFNVVTEDCDEEIKELAEDLRKQLSRLELDEQSSSSIVPKECRVLFRKMNDALHCSYKKEDGFASLQLGRDLE
FLIEKPIPLSDRVLKDEYGYVK
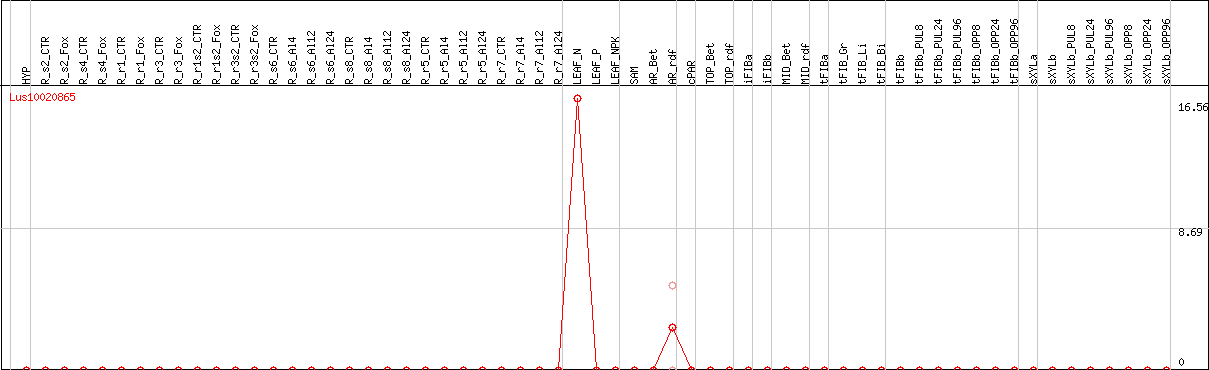
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020865 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.