Lus10020887 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10020887 pacid=23177966 polypeptide=Lus10020887 locus=Lus10020887.g ID=Lus10020887.BGIv1.0 annot-version=v1.0
ATGGCGTCCCAAATTGGCGGCCGCCATTCAGTTCTGATCTTTCTATTCCTTCTCTCCATCGTTTCCCAAACAGCACTCTCATGGAAGAAAGAAGAGTTCC
GTAATTGTAATCAGACCCCTTTCTGCAAGCGTGCTCGCTCTCGCAACCCCGGCGCTTGTAACTTGATCGCCGATCAAGTTTCCATCTCCGAAGGTCGGCT
CACTGCCAAGCTCATCCCGAGACCTATTCCCGATCAGGGGGATCAGATCTCCAAGCCTCTGCTTCTCTCGCTCTCGGTACACCAGAACGGGATCTTACGG
TTGAAAATCGATGAGGAATCTTCGCTTGACCCGCCAAAATCTCGATTCCAAGTCCCCGACGTGGTAGTCCCCGAGTTTGAGGAGAGTATGCTATGGCTGT
CTAGGGTTTCTACTGAGACTGTCGATGGCGATTCAGCTCTTTCCACTGTCGTCTATCTAGCGGACGGGTATGAGGCTGTTCTTCGACATGATCCGTTTGA
GATTTACGTCCGAGAGAAGTCAGGTAAGCGCCGTTTGGTTTCTATGAACTCTCATCTGCTGTTTGATTTCGAGCAATTGAGGTCGAAAGATGAAGGGGAA
GATTGGGAAGAGAGGTTCAAGAGTCATACAGATAGTAGACCTTACGGACCTCAGTCGATCAGTTTTGATGTGTCGTTTTATGATGCTGAAATTGTCTCCG
GTCTACCGGAGCATGCTACTAGCGTAGCTCTCAAGCCTACTAAAGGTCCTGGAGTTGATGACTCAGAGCCTTACAGATTGTTTAACTTGGATGTTTTCGA
GTACCTCCATGAGTCGCCTTTCGGTCTCTATGGTTCAATACCATTGATGATTGCTCATGGGAAGGGTGGGAAGAGTTCTGGATTCTTCTGGTTAAACGCT
GGTGAGATGCAAATTGATGTATTGGGTGATGGCTGGAATGCTGATTCTGGGCTTTCCCTGCCTTCTGATCAGAAGAGAATCGATACACTCTGGATGAGTG
AGGCAGGCATTGTAGATGCTTTCTTCTTTGTTGGACCAGGTCCTAAGGACGTTGTGAGCCAATACATTGCTGTCACGGGGAAACCAGCAATGCCACAGCA
TTTCGCCACTGCTTATCATCAGTGTAGGTGGAATTACAGGGATGAGGAGGATGTCGACAATGTGGATGCCAAGTTCGACGAGCACGATATTCCCTATGAC
GTTTTGTGGCTAGACATTGAGCACACTGATGGTAAGAGGTACATGACGTGGGATTCTGTTACGTTCCCGCATCCCGAGGAAATGCAAAAGAAGTTGGCAG
CCAAAGGAAGGCACATGGTGACCATCGTTGATCCTCATATTAAGAGGGATGACAACTATTACATACATAAGGAGGCTACAGAGAAAGGATATTATGTGAA
GGATTCTAGTGGGAAGGACTTTGATGGGTGGTGCTGGCCAGGTTCCTCATCTTACTTGGATGTGTTGAGCCCCGAGGTTAGAACGTGGTTTAGTAGCAAG
TTTGCATTTGACAGCTACAAGGGTTCAACTCCTTCACTGTATATTTGGAATGATATGAATGAGCCTTCCGTGTTCAATGGTCCAGAGATTTCAATGCCAA
GAGATGCTTTACATTTTGGGGGAGTGGAGCATCGGGAACTACACAACGCATATGGCTACTACTACCATATGGCAACATCAAATGGTCTTATCAAGCGTGG
AGATGGGACCGACAGGCCCTTTGTTCTGTCAAGAGCATTCTTTGCTGGAACTCAGAGGTTTGGATCAGTCTGGACAGGGGACAACACTGCAGACTGGGAT
CATTTGAGAGCGTCTGTACCAATGCTTTTGACCCTTGGTGCGGATGTCGGAGGGTTTTTCGGTAATCCGGAGCCGGAGTTATTAGTTCGTTGGTACCAAC
TTGGTGCTTTCTATCCTTTCTTCAGGGCCCATGCTCATCAGGACACGAAAAGACGGGAGCCTTGGTTATTTGGAGACCATAATACGAAGCTCATTAGACA
AGCTGTACATGTCCGTTACATGTTGCTTCCATATTTCTACACGTTGTTCCGAGAGGCCAACATGACCGGTACCCCCATTATGCGTCCTCTGTGGATGGAG
TTCCCTTCTGATGAGTCCACCTTTAACAATGATGAAGCTTTCATGCTCGGAAGCAGCATCTTAGTCCAAGGAATTTACACAGAGCGAGCGAAACACGCGT
CGGTTTATCTACCAGGCAAAGAACTCTGGTACGACCTTTTGAGCGGATCTCCGTACAAGGGTTCTCAAACTCACAAGCTAGTGGCTTTGGAGGACCGAAT
TCCTGCATTCCAAAAGGCCGGGACGATCATACCCAGGAAAGATCGTTATCGTCGCAGCTCAACACAGATGGCTAATGATCCCTATACTTTGGTGATAGCC
CTCAACAGTTCTCAAGCAGCTGAAGGCGAGCTCTACATCGACGACGGGAAGACCTTCGAGTTCCTTGAAGGAGCCTACATCCACCGCCGATTCGTGTTCT
CAAACGGCAAGCTCTCATCTTTCGACTTGACTCATAACAAGAAGCCGCGATTCACATCCCAGACTGTCATCGAAAGAATCATACTGCTGGGATACTCTCC
CGGACCTAAGAGTGCCCTGATCAAGCCAGGGAATGGTAAAGCCGAAGTCGGTCTAGGGCCACTCGTACTTGAAGAGGGGATTCGATCTTCGTCCTCGTCG
GTTGTAACCATCCGGAAGCCGAACCTCCCAGTGGACAAGGACTGGAGTGTAGAACTTGTGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10020887 pacid=23177966 polypeptide=Lus10020887 locus=Lus10020887.g ID=Lus10020887.BGIv1.0 annot-version=v1.0
MASQIGGRHSVLIFLFLLSIVSQTALSWKKEEFRNCNQTPFCKRARSRNPGACNLIADQVSISEGRLTAKLIPRPIPDQGDQISKPLLLSLSVHQNGILR
LKIDEESSLDPPKSRFQVPDVVVPEFEESMLWLSRVSTETVDGDSALSTVVYLADGYEAVLRHDPFEIYVREKSGKRRLVSMNSHLLFDFEQLRSKDEGE
DWEERFKSHTDSRPYGPQSISFDVSFYDAEIVSGLPEHATSVALKPTKGPGVDDSEPYRLFNLDVFEYLHESPFGLYGSIPLMIAHGKGGKSSGFFWLNA
GEMQIDVLGDGWNADSGLSLPSDQKRIDTLWMSEAGIVDAFFFVGPGPKDVVSQYIAVTGKPAMPQHFATAYHQCRWNYRDEEDVDNVDAKFDEHDIPYD
VLWLDIEHTDGKRYMTWDSVTFPHPEEMQKKLAAKGRHMVTIVDPHIKRDDNYYIHKEATEKGYYVKDSSGKDFDGWCWPGSSSYLDVLSPEVRTWFSSK
FAFDSYKGSTPSLYIWNDMNEPSVFNGPEISMPRDALHFGGVEHRELHNAYGYYYHMATSNGLIKRGDGTDRPFVLSRAFFAGTQRFGSVWTGDNTADWD
HLRASVPMLLTLGADVGGFFGNPEPELLVRWYQLGAFYPFFRAHAHQDTKRREPWLFGDHNTKLIRQAVHVRYMLLPYFYTLFREANMTGTPIMRPLWME
FPSDESTFNNDEAFMLGSSILVQGIYTERAKHASVYLPGKELWYDLLSGSPYKGSQTHKLVALEDRIPAFQKAGTIIPRKDRYRRSSTQMANDPYTLVIA
LNSSQAAEGELYIDDGKTFEFLEGAYIHRRFVFSNGKLSSFDLTHNKKPRFTSQTVIERIILLGYSPGPKSALIKPGNGKAEVGLGPLVLEEGIRSSSSS
VVTIRKPNLPVDKDWSVELV
|
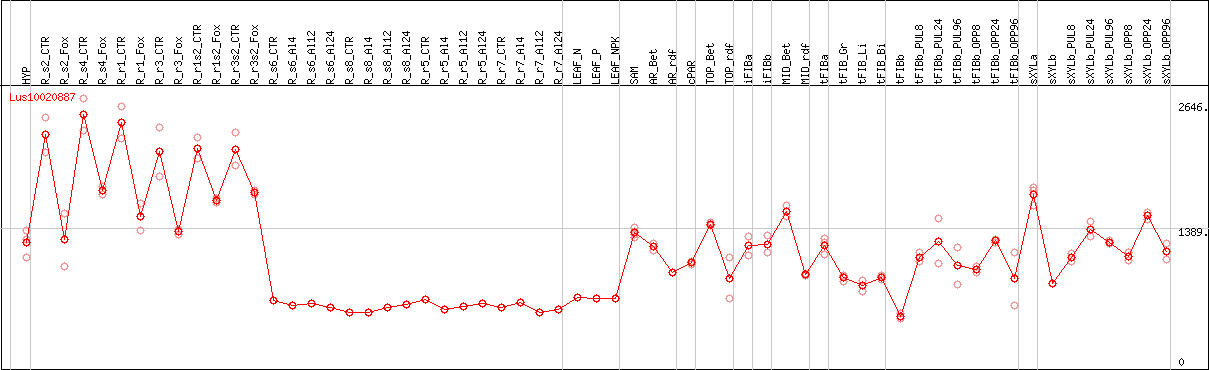
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020887 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
