External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G34780 258 / 8e-85
ATAPRL4
APR-like 4 (.1.2)
AT4G08930 224 / 9e-72
ATAPRL6
APR-like 6 (.1)
AT3G03860 119 / 2e-31
ATAPRL5
APR-like 5 (.1)
AT5G18120 112 / 7e-29
ATAPRL7
APR-like 7 (.1)
AT3G54960 51 / 7e-07
ATPDI1, ATPDIL1-3
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 1, PDI-like 1-3 (.1.2)
AT2G47470 50 / 1e-06
ATPDI11, ATPDIL2-1, UNE5, MEE30
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
AT2G32920 47 / 2e-05
ATPDI9, ATPDIL2-3
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 9, PDI-like 2-3 (.1)
AT1G04980 41 / 0.0007
ATPDI10, ATPDIL2-2
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 10, PDI-like 2-2 (.1)
AT5G60640 41 / 0.0009
ATPDI2, ATPDIL1-4
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 2, PDI-like 1-4 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033449
584 / 0
AT1G34780 260 / 8e-86
APR-like 4 (.1.2)
Lus10036486
375 / 9e-131
AT1G34780 272 / 2e-90
APR-like 4 (.1.2)
Lus10010354
370 / 3e-129
AT1G34780 268 / 2e-89
APR-like 4 (.1.2)
Lus10005703
134 / 1e-36
AT3G03860 310 / 7e-106
APR-like 5 (.1)
Lus10020283
121 / 4e-32
AT3G03860 296 / 3e-100
APR-like 5 (.1)
Lus10010274
49 / 3e-06
AT2G47470 549 / 0.0
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Lus10036337
48 / 5e-06
AT2G47470 548 / 0.0
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Lus10020691
46 / 2e-05
AT2G47470 473 / 3e-168
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Lus10015160
45 / 7e-05
AT1G04980 621 / 0.0
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 10, PDI-like 2-2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G164200
306 / 9e-104
AT1G34780 364 / 1e-126
APR-like 4 (.1.2)
Potri.002G097601
306 / 2e-103
AT1G34780 357 / 9e-124
APR-like 4 (.1.2)
Potri.013G057100
119 / 2e-31
AT5G18120 293 / 3e-99
APR-like 7 (.1)
Potri.019G035700
115 / 1e-29
AT5G18120 277 / 4e-93
APR-like 7 (.1)
Potri.014G122800
47 / 6e-06
AT2G47470 551 / 0.0
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Potri.002G198300
47 / 8e-06
AT2G47470 530 / 0.0
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Potri.008G040100
44 / 7e-05
AT3G54960 288 / 7e-92
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 1, PDI-like 1-3 (.1.2)
Potri.002G082100
42 / 0.0006
AT1G21750 689 / 0.0
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 5, PDI-like 1-1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF00085
Thioredoxin
Thioredoxin
Representative CDS sequence
>Lus10020928 pacid=23177987 polypeptide=Lus10020928 locus=Lus10020928.g ID=Lus10020928.BGIv1.0 annot-version=v1.0
ATGGGGTCTACGATTAGGGTTTCGAAGAGGAGGACATGGGAGTTGGTTGTTGTGGTGGTGGTATTGGCACTCACTCTTTCTGGCACCTCCGCTGCTTCTA
AGGATCTGATGTGCCCTAGGCTCTCTATCACCGATTCCATCTTTGGGGTCGGAGAATTGAAACCCGGTTTTCACTCTGCTAATTCTCTGTCTTCCGCCTG
CGTCAAAGAGGGAGGCGAAGCTGCATTATGGAAAGCACTGGATTTGGTTTACAGTGGTAGTTTGGAGAATGTAGCTGTGCTATTTTATGCAGCATGGTGC
CCTTTTTCGACGGCATTAAAACTGGAATTCCCTATCTTGGCTTCCCTGTATCCATCTATCCCCCATATAGCTGTTGAAGAATCAGCCATCCGGCCCAGCA
CGCTTTCCAAGTATGGAGTTCATGGCTTTCCCACTCTTCTGCTTCTGAATTCTACAATGCGTGTACGCTACCATGGTTCCCGGACATTTAGCTCTCTGGT
TACCTTCTACAATGATGCAACTGGTGTGTTACCTGATAATGCGTCATTTTCGAAGATATGGTACCCTTTGACCCTTGGAAAGCATGACATTATTGACCCA
GAAAGCTGCCCGTTTTCCTGGGCTAGGTCTCCAGAGAAATTGCTGATGCAAGAAACGTATCTGGCTCTTGCCTCAGCCTTTCTGGTTTTTAGATTCCTCT
ACTTAATGTTACCGAGTCTGGTTGTATTTGCGCAACTAGTTTGGAGGAGGCAGGTGAGTCGTGCAAGGATCAGGAGCATCCTGGGTCAGCCGCGAGTCTT
TATGACCGATGTTGTCCAGGCATTCAAGTGCCTTTTGAAACCTTGCAAGAAAGGTAATTTACAGGAAGGGGCAATGAATGCCGGAGCATGGGCTTCGAAG
TCGCTCGTCACAGTAGCTAGAGCGGGAGAAGCAAGCACGAGTGGCACGGCACAGAAATCGTGA
AA sequence
>Lus10020928 pacid=23177987 polypeptide=Lus10020928 locus=Lus10020928.g ID=Lus10020928.BGIv1.0 annot-version=v1.0
MGSTIRVSKRRTWELVVVVVVLALTLSGTSAASKDLMCPRLSITDSIFGVGELKPGFHSANSLSSACVKEGGEAALWKALDLVYSGSLENVAVLFYAAWC
PFSTALKLEFPILASLYPSIPHIAVEESAIRPSTLSKYGVHGFPTLLLLNSTMRVRYHGSRTFSSLVTFYNDATGVLPDNASFSKIWYPLTLGKHDIIDP
ESCPFSWARSPEKLLMQETYLALASAFLVFRFLYLMLPSLVVFAQLVWRRQVSRARIRSILGQPRVFMTDVVQAFKCLLKPCKKGNLQEGAMNAGAWASK
SLVTVARAGEASTSGTAQKS
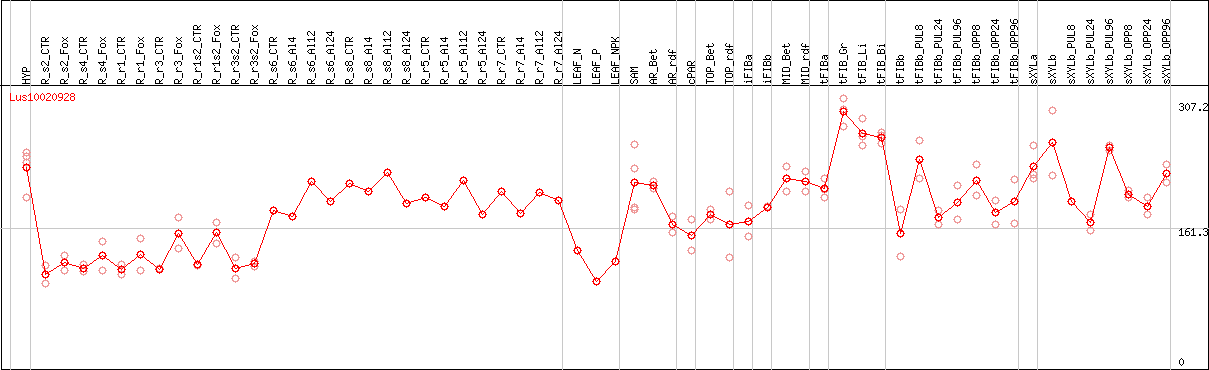
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10020928 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.