External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G04410 107 / 1e-29
c-NAD-MDH1
cytosolic-NAD-dependent malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
AT5G43330 103 / 2e-28
c-NAD-MDH2
cytosolic-NAD-dependent malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1)
AT5G56720 85 / 3e-21
c-NAD-MDH3
cytosolic-NAD-dependent malate dehydrogenase 3, Lactate/malate dehydrogenase family protein (.1)
AT5G58330 42 / 7e-06
lactate/malate dehydrogenase family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011819
110 / 1e-30
AT5G43330 613 / 0.0
cytosolic-NAD-dependent malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1)
Lus10021870
109 / 1e-30
AT5G43330 606 / 0.0
cytosolic-NAD-dependent malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1)
Lus10041144
99 / 1e-25
AT5G43330 590 / 0.0
cytosolic-NAD-dependent malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1)
Lus10037935
40 / 4e-05
AT5G58330 709 / 0.0
lactate/malate dehydrogenase family protein (.1.2.3)
Lus10038668
40 / 4e-05
AT5G58330 707 / 0.0
lactate/malate dehydrogenase family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G071000
108 / 6e-30
AT5G43330 602 / 0.0
cytosolic-NAD-dependent malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1)
Potri.008G166800
106 / 2e-29
AT5G43330 605 / 0.0
cytosolic-NAD-dependent malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1)
Potri.002G141700
97 / 1e-25
AT1G04410 572 / 0.0
cytosolic-NAD-dependent malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Potri.002G141900
97 / 1e-25
AT1G04410 501 / 5e-180
cytosolic-NAD-dependent malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Potri.008G031700
42 / 1e-05
AT5G58330 710 / 0.0
lactate/malate dehydrogenase family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00056
Ldh_1_N
lactate/malate dehydrogenase, NAD binding domain
Representative CDS sequence
>Lus10021183 pacid=23171378 polypeptide=Lus10021183 locus=Lus10021183.g ID=Lus10021183.BGIv1.0 annot-version=v1.0
ATGGCGAAGGATCTAGCTCGCATTCTCGTCACCGAAGCTGCAGGACAAATTGGATATGCTCTTGCTCCCATGATTGCTAGGGGAGTGATGCTTGGTGCTG
ACCAGCCAGTCATCCTCCACATGCTTGATATCCCACCTGCAGCTGAGGCTTTGAATGGTGTCAAGATGGAGTTGGTTGATGCTGCGTTCTCTCTTCTTAA
AGGTTTCTCCAGTTCTTTACTAGCTCTCTTTGGGATTTCTGTTGGATGA
AA sequence
>Lus10021183 pacid=23171378 polypeptide=Lus10021183 locus=Lus10021183.g ID=Lus10021183.BGIv1.0 annot-version=v1.0
MAKDLARILVTEAAGQIGYALAPMIARGVMLGADQPVILHMLDIPPAAEALNGVKMELVDAAFSLLKGFSSSLLALFGISVG
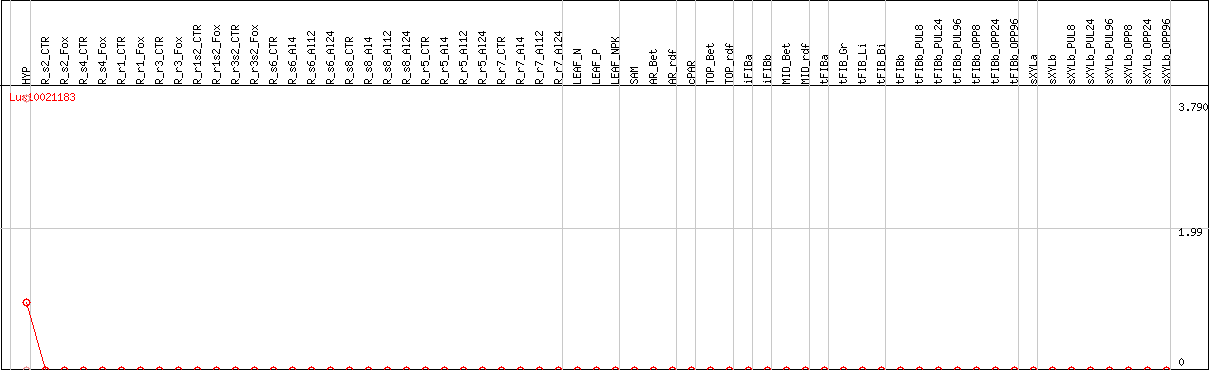
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021183 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.