External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06200 146 / 3e-44
CASP4
Casparian strip membrane domain protein 4, Uncharacterised protein family (UPF0497) (.1)
AT3G11550 145 / 1e-43
CASP2
Casparian strip membrane domain protein 2, Uncharacterised protein family (UPF0497) (.1)
AT2G36100 133 / 5e-39
CASP1
Casparian strip membrane domain protein 1, Uncharacterised protein family (UPF0497) (.1)
AT2G27370 116 / 3e-32
CASP3
Casparian strip membrane domain protein 3, Uncharacterised protein family (UPF0497) (.1)
AT5G15290 99 / 9e-26
CASP5
Casparian strip membrane domain protein 5, Uncharacterised protein family (UPF0497) (.1)
AT1G14160 89 / 8e-22
Uncharacterised protein family (UPF0497) (.1)
AT3G06390 46 / 4e-06
Uncharacterised protein family (UPF0497) (.1)
AT1G03700 42 / 7e-05
Uncharacterised protein family (UPF0497) (.1)
AT4G03540 42 / 9e-05
Uncharacterised protein family (UPF0497) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017008
230 / 1e-76
AT2G27370 228 / 5e-76
Casparian strip membrane domain protein 3, Uncharacterised protein family (UPF0497) (.1)
Lus10021333
158 / 1e-48
AT3G11550 246 / 1e-83
Casparian strip membrane domain protein 2, Uncharacterised protein family (UPF0497) (.1)
Lus10017009
155 / 2e-43
AT3G11540 1356 / 0.0
SPINDLY, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10004205
141 / 5e-42
AT3G11550 232 / 8e-78
Casparian strip membrane domain protein 2, Uncharacterised protein family (UPF0497) (.1)
Lus10029409
140 / 1e-41
AT3G11550 234 / 9e-79
Casparian strip membrane domain protein 2, Uncharacterised protein family (UPF0497) (.1)
Lus10015375
105 / 2e-28
AT5G15290 219 / 1e-73
Casparian strip membrane domain protein 5, Uncharacterised protein family (UPF0497) (.1)
Lus10033007
101 / 9e-27
AT5G15290 216 / 3e-72
Casparian strip membrane domain protein 5, Uncharacterised protein family (UPF0497) (.1)
Lus10037160
86 / 2e-20
AT1G14160 214 / 9e-71
Uncharacterised protein family (UPF0497) (.1)
Lus10036770
48 / 4e-07
AT1G14160 119 / 2e-34
Uncharacterised protein family (UPF0497) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G208800
160 / 9e-50
AT5G06200 201 / 7e-66
Casparian strip membrane domain protein 4, Uncharacterised protein family (UPF0497) (.1)
Potri.016G075400
159 / 5e-49
AT5G06200 179 / 2e-57
Casparian strip membrane domain protein 4, Uncharacterised protein family (UPF0497) (.1)
Potri.016G075300
134 / 2e-39
AT5G06200 162 / 2e-50
Casparian strip membrane domain protein 4, Uncharacterised protein family (UPF0497) (.1)
Potri.009G160000
109 / 3e-29
AT2G27370 146 / 2e-43
Casparian strip membrane domain protein 3, Uncharacterised protein family (UPF0497) (.1)
Potri.012G032300
98 / 2e-25
AT5G15290 207 / 1e-68
Casparian strip membrane domain protein 5, Uncharacterised protein family (UPF0497) (.1)
Potri.008G088700
81 / 7e-19
AT1G14160 163 / 3e-51
Uncharacterised protein family (UPF0497) (.1)
Potri.004G043300
47 / 1e-06
AT4G03540 103 / 2e-28
Uncharacterised protein family (UPF0497) (.1)
Potri.011G052100
44 / 2e-05
AT4G03540 100 / 2e-27
Uncharacterised protein family (UPF0497) (.1)
Potri.001G436400
42 / 8e-05
AT1G17200 245 / 1e-82
Uncharacterised protein family (UPF0497) (.1), Uncharacterised protein family (UPF0497) (.2)
Potri.013G131700
40 / 0.0004
AT1G03700 159 / 9e-50
Uncharacterised protein family (UPF0497) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0396
Marvel-like
PF04535
DUF588
Domain of unknown function (DUF588)
Representative CDS sequence
>Lus10021332 pacid=23178236 polypeptide=Lus10021332 locus=Lus10021332.g ID=Lus10021332.BGIv1.0 annot-version=v1.0
ATGTCGACTTCAATTGAGATGACAGAGGAAAGCAGCGCTGTGGCGAAGGGAAAAGCGCCTCTGATCACCACAACTACGTACAGCCACGACGACGGTCATG
CAAAGGGAAGAAACGATAGCAATAATTATAAGACTGTTGGTGGGTACAAGAAGGGAATTGCCATTTTCGACTTCGTCCTAAGGCTAGCAGCCGTCGTTGC
TGCTCTGGCTGCTGCTGGTACAATGGGGAGCAGCGAGCAAACCCTCCCCTTCTTCACCCAGTTCTTCCAGTTCCAGGCCAGTTACGACGACCTCCCTACC
TTCCAGTTCTTTGTGATCGCAATGGCAATCGTGGGAGGCTACTTGGTTCTGTCTCTCCCGTTCTCAATCGTCCTCATCATCCGCCCACAAGCCGTAGCTC
CGAGGCTGCTACTCGTGCTCCTTGACACGGTGGCTCTGACTCTGAACACGTCAGCAGCAGGGGCAGCGGCAGCGATAGTGTACTTGGCTCACAACGGGAA
CTCTAGCGCCAACTGGCTCGCAATCTGCCAGCAGTTCGGCGACTTCTGTCAGAAAGTCAGTGGAGCAGTTGTCGCTTCGTTTGTGGCAGCTGTGGCCTTG
ATGTTCTTGGTTGTCCTCTCTGGCTTGGCTCTCCGAAGGCATTAG
AA sequence
>Lus10021332 pacid=23178236 polypeptide=Lus10021332 locus=Lus10021332.g ID=Lus10021332.BGIv1.0 annot-version=v1.0
MSTSIEMTEESSAVAKGKAPLITTTTYSHDDGHAKGRNDSNNYKTVGGYKKGIAIFDFVLRLAAVVAALAAAGTMGSSEQTLPFFTQFFQFQASYDDLPT
FQFFVIAMAIVGGYLVLSLPFSIVLIIRPQAVAPRLLLVLLDTVALTLNTSAAGAAAAIVYLAHNGNSSANWLAICQQFGDFCQKVSGAVVASFVAAVAL
MFLVVLSGLALRRH
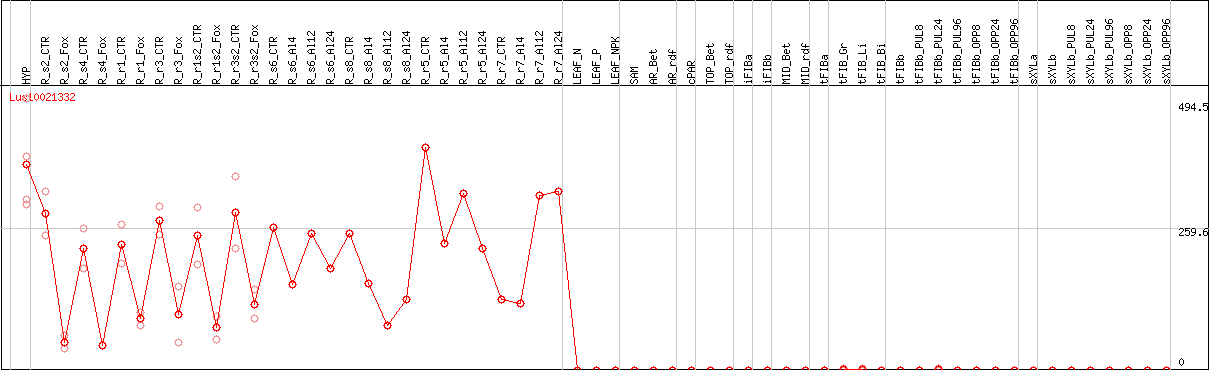
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021332 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.