External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G52720 220 / 1e-70
CAH1, ATACA1, ACA1
A. THALIANA ALPHA CARBONIC ANHYDRASE 1, alpha carbonic anhydrase 1 (.1.2)
AT1G08080 199 / 1e-62
ATACA7, ACA7
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
AT1G08065 165 / 2e-49
ATACA5, ACA5
alpha carbonic anhydrase 5 (.1)
AT4G20990 154 / 3e-45
ATACA4, ACA4
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
AT2G28210 151 / 1e-44
ATACA2
alpha carbonic anhydrase 2 (.1)
AT5G04180 152 / 2e-44
ATACA3, ACA3
alpha carbonic anhydrase 3 (.1)
AT4G21000 127 / 5e-35
ATACA6, ACA6
A. THALIANA ALPHA CARBONIC ANHYDRASE 6, alpha carbonic anhydrase 6 (.1)
AT5G56330 119 / 2e-31
ATACA8, ACA8
A. THALIANA ALPHA CARBONIC ANHYDRASE 8, alpha carbonic anhydrase 8 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017033
521 / 0
AT3G52720 231 / 4e-75
A. THALIANA ALPHA CARBONIC ANHYDRASE 1, alpha carbonic anhydrase 1 (.1.2)
Lus10022710
286 / 1e-96
AT3G52720 218 / 4e-70
A. THALIANA ALPHA CARBONIC ANHYDRASE 1, alpha carbonic anhydrase 1 (.1.2)
Lus10021455
195 / 7e-61
AT1G08080 343 / 1e-119
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Lus10016104
192 / 1e-59
AT1G08080 344 / 7e-120
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Lus10042030
192 / 5e-59
AT4G20990 313 / 9e-107
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
Lus10018017
191 / 2e-58
AT4G20990 240 / 4e-78
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
Lus10021454
187 / 5e-58
AT1G08080 300 / 1e-102
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Lus10016105
183 / 2e-56
AT1G08080 333 / 1e-115
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Lus10042013
172 / 1e-51
AT4G20990 224 / 2e-72
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G079600
238 / 1e-77
AT3G52720 271 / 8e-91
A. THALIANA ALPHA CARBONIC ANHYDRASE 1, alpha carbonic anhydrase 1 (.1.2)
Potri.006G212900
218 / 6e-70
AT3G52720 261 / 5e-87
A. THALIANA ALPHA CARBONIC ANHYDRASE 1, alpha carbonic anhydrase 1 (.1.2)
Potri.004G212800
189 / 1e-58
AT1G08080 347 / 3e-121
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Potri.006G047500
182 / 9e-56
AT4G20990 276 / 5e-93
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
Potri.009G010200
180 / 3e-55
AT1G08080 342 / 3e-119
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Potri.016G043700
176 / 2e-53
AT4G20990 306 / 4e-105
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
Potri.006G121900
173 / 2e-52
AT1G08080 289 / 4e-98
A. THALIANA ALPHA CARBONIC ANHYDRASE 7, alpha carbonic anhydrase 7 (.1)
Potri.006G047400
169 / 6e-51
AT4G20990 289 / 4e-98
A. THALIANA ALPHA CARBONIC ANHYDRASE 4, alpha carbonic anhydrase 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00194
Carb_anhydrase
Eukaryotic-type carbonic anhydrase
Representative CDS sequence
>Lus10021356 pacid=23178168 polypeptide=Lus10021356 locus=Lus10021356.g ID=Lus10021356.BGIv1.0 annot-version=v1.0
ATGGGTACAACAACTGATCAAACAGCATCATTCATCACCTTCTCTGTTCTAGCAACCATTATTGCTCTGTCTCTAGCTCTCTGTTCTTGTCTTGAAGCCG
CTCCTCCTGCTCCTTCAGGGTTTGTGAACTTTGGTTACAGGGGGCCAAACGGACCTCAGAGATGGGGGAGCTTGACCCCCAATTACAAAGAGTGCGATAT
TGGGAAATTGCAGTCTCCTATCGACATTAACAGGCTGGATGTTGTGGAGAACAAGAAGCTTGAGAGATTGAAAGGCAATTACATTGCTGTTAACGCCACT
CTTATTAACACTGGCCGTAGCATCATGGTTCATTACGACGATATGCACGCTGGAGAGCTGGAAGTAGATGGGAAACAATATAAATTGCAGCAAATGCATT
GGCATTCTCCTTCAGAGCACACTTTTGATGGACACCAGTATCCATTGGAGCTCCACTTAGTTCATCAAGCAGACGATGGTATTTTCACAGTTGTATCAGT
CCTCCATCAGTACGGACCCAACGAAGATTCACTCATCAAGAAGATTAGAGTGGAGTTGGAGGAGCTAGCTGAGGAGAAATCGTCGTCGCAGAAAGATGGA
GGAACAGTAGTAGCAATACCACCACAAGTGAAGCTGAGGAAGAAAATAGAGTACAAGATGCTGAAGAAGAAAACACACAAGTACTACAGATACACTGGCT
CCCTCACTTCTCCTCCTTGCACTCAAAACGTCACCTGGAACGTCCTTGCTAAGGTGAGGAGTGTGTCAAGGGAACAGGTGAAAGCTCTGAGAGATCTGTT
ATCGGCAGATTGCAAGGCGAATGCTAGGCCGTTACAGCCTCTTAATGGCAGGAAAGTTGAGGTCTACAAGGAACCTAACGATCTCTGA
AA sequence
>Lus10021356 pacid=23178168 polypeptide=Lus10021356 locus=Lus10021356.g ID=Lus10021356.BGIv1.0 annot-version=v1.0
MGTTTDQTASFITFSVLATIIALSLALCSCLEAAPPAPSGFVNFGYRGPNGPQRWGSLTPNYKECDIGKLQSPIDINRLDVVENKKLERLKGNYIAVNAT
LINTGRSIMVHYDDMHAGELEVDGKQYKLQQMHWHSPSEHTFDGHQYPLELHLVHQADDGIFTVVSVLHQYGPNEDSLIKKIRVELEELAEEKSSSQKDG
GTVVAIPPQVKLRKKIEYKMLKKKTHKYYRYTGSLTSPPCTQNVTWNVLAKVRSVSREQVKALRDLLSADCKANARPLQPLNGRKVEVYKEPNDL
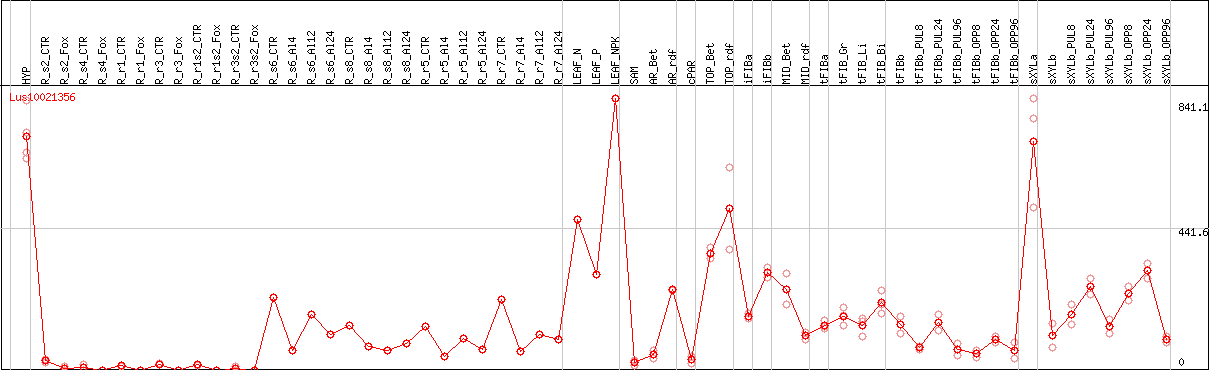
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021356 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.