External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G26060 461 / 3e-164
EMB1345
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
AT4G32990 347 / 3e-119
Transducin/WD40 repeat-like superfamily protein (.1)
AT4G02730 95 / 4e-22
AtWDR5b
human WDR5 \(WD40 repeat\) homolog b, Transducin/WD40 repeat-like superfamily protein (.1)
AT2G41500 82 / 4e-17
EMB2776, LIS
LACHESIS, WD-40 repeat family protein / small nuclear ribonucleoprotein Prp4p-related (.1)
AT3G49660 79 / 1e-16
AtWDR5a
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G13480 80 / 3e-16
FY
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT5G67320 80 / 3e-16
HOS15
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
AT5G52820 79 / 3e-16
WD-40 repeat family protein / notchless protein, putative (.1)
AT2G21390 78 / 1e-15
Coatomer, alpha subunit (.1)
AT1G62020 77 / 2e-15
Coatomer, alpha subunit (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017045
546 / 0
AT2G26060 486 / 5e-174
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10024447
107 / 2e-25
AT5G25150 1006 / 0.0
TBP-associated factor 5 (.1)
Lus10004167
100 / 2e-23
AT2G41500 708 / 0.0
LACHESIS, WD-40 repeat family protein / small nuclear ribonucleoprotein Prp4p-related (.1)
Lus10021059
100 / 3e-23
AT2G41500 706 / 0.0
LACHESIS, WD-40 repeat family protein / small nuclear ribonucleoprotein Prp4p-related (.1)
Lus10007449
94 / 9e-21
AT5G25150 957 / 0.0
TBP-associated factor 5 (.1)
Lus10014674
91 / 1e-20
AT4G02730 508 / 0.0
human WDR5 \(WD40 repeat\) homolog b, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10006916
89 / 4e-20
AT4G02730 506 / 0.0
human WDR5 \(WD40 repeat\) homolog b, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10024972
86 / 2e-18
AT5G67320 819 / 0.0
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
Lus10002324
83 / 1e-17
AT3G15980 397 / 7e-132
Coatomer, beta' subunit (.1.2.3.4.5)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G212800
511 / 0
AT2G26060 530 / 0.0
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
Potri.001G221000
510 / 0
AT2G26060 523 / 0.0
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
Potri.014G136700
168 / 6e-52
AT2G26060 156 / 2e-47
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
Potri.006G045800
95 / 2e-21
AT2G41500 614 / 0.0
LACHESIS, WD-40 repeat family protein / small nuclear ribonucleoprotein Prp4p-related (.1)
Potri.018G020200
94 / 3e-21
AT5G25150 984 / 0.0
TBP-associated factor 5 (.1)
Potri.002G051900
89 / 4e-20
AT4G02730 524 / 0.0
human WDR5 \(WD40 repeat\) homolog b, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.006G263400
91 / 8e-20
AT5G25150 1018 / 0.0
TBP-associated factor 5 (.1)
Potri.007G050100
89 / 2e-19
AT5G67320 645 / 0.0
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
Potri.005G148500
86 / 7e-19
AT3G49660 503 / 0.0
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.005G144400
86 / 1e-18
AT5G67320 671 / 0.0
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Lus10021365 pacid=23178180 polypeptide=Lus10021365 locus=Lus10021365.g ID=Lus10021365.BGIv1.0 annot-version=v1.0
ATGGACTTCCTCGATGACGGTGGCGGCAACTTGGAGCTCAAGGAGATTCAGACGCTCCGAGGTCACACCGATAGGGTTTGGAGCCTGGCGTGGAATCCGG
CCACCGGAACTGGTGACACCCAGCCAATCATTGCTTCCTGCAGCGGCGATAAGACTGTTCGAATCTGGGAACGTAGTGCCTTGACCCGTGCTTGGGACTG
CAAGGCAGTTTTAGAAGAAACTCATACCAGGACTGTTCGGTCGTGCGCCTGGTCACCGTCTGGGAAATTATTGGCTACTGCCAGCTTTGATGCTACCACT
GCCATTTGGGAGAATGTTGCGGGTGATTTTCAATGTGTTTCCACGTTAGAGGGTCATGAAAATGAAGTCAAGAGCGTGTCATGGAATGCATCCGAGTCTC
TCTTGGCAACTTGTAGTCGAGACAAAACTGTTTGGATTTGGGAAGTGATGCCTGGGAATGAATACGAATGTGCTTCAGTATTGCAGGGCCACACACAAGA
TGTTAAGATGGTTAAGTGGCACCCAACCATGGACATACTGTTTTCCTGTAGCTATGATAATACCATTAAGGTCTGGGCTGAGGATGGTACTGGTGATTGG
GACTGCATTCAAACGATAGATGAATCGAACAATGGCCACTCTTCTACTGTTTGGGCCATTTCTTTCAATGCCGAAGGAGACAAAATGGTTACATGTAGTG
ATGATCTTACCTTAAAGGTATGGGGAACAGATCTCGAGGGAATGAAATCTGGTGATGATCAAGTACCCTGGAAACATCTTTGCACTCTATCGGGTTATCA
TGATCGCACCATATTTTCAGTTGACTGGTCAAGGGAAGGTATTATAGCCAGTGGAGCAGCAGATGATGCTATACGGTTTTTTGCTGAAACCAAAGACAGT
GCGGAAAATAGGCTGTTGGCTTCTGCAAGTGATGATGGAACAATAAAGATATGGGAATTGGCAACTCGCACCTGA
AA sequence
>Lus10021365 pacid=23178180 polypeptide=Lus10021365 locus=Lus10021365.g ID=Lus10021365.BGIv1.0 annot-version=v1.0
MDFLDDGGGNLELKEIQTLRGHTDRVWSLAWNPATGTGDTQPIIASCSGDKTVRIWERSALTRAWDCKAVLEETHTRTVRSCAWSPSGKLLATASFDATT
AIWENVAGDFQCVSTLEGHENEVKSVSWNASESLLATCSRDKTVWIWEVMPGNEYECASVLQGHTQDVKMVKWHPTMDILFSCSYDNTIKVWAEDGTGDW
DCIQTIDESNNGHSSTVWAISFNAEGDKMVTCSDDLTLKVWGTDLEGMKSGDDQVPWKHLCTLSGYHDRTIFSVDWSREGIIASGAADDAIRFFAETKDS
AENRLLASASDDGTIKIWELATRT
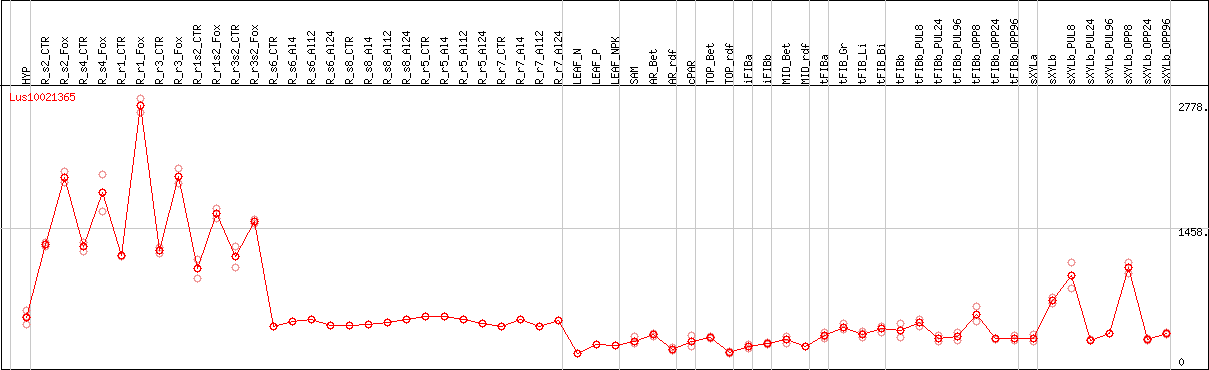
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021365 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.