External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G39510 322 / 9e-113
ZIG1, SGR4, ATVTI1A, ATVTI11, ZIG, VTI11
SHOOT GRAVITROPSIM 4, VESICLE TRANSPORT V-SNARE 11, Vesicle transport v-SNARE family protein (.1)
AT1G26670 258 / 9e-88
VTI1B, ATVTI12, VTI12
VESICAL TRANSPORT V-SNARE 12, Vesicle transport v-SNARE family protein (.1)
AT3G29100 242 / 1e-81
ATVTI13, VTI13
vesicle transport V-snare 13 (.1.2)
AT5G39630 176 / 2e-55
Vesicle transport v-SNARE family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034696
398 / 1e-141
AT5G39510 318 / 5e-110
SHOOT GRAVITROPSIM 4, VESICLE TRANSPORT V-SNARE 11, Vesicle transport v-SNARE family protein (.1)
Lus10026134
259 / 5e-89
AT5G39510 255 / 3e-87
SHOOT GRAVITROPSIM 4, VESICLE TRANSPORT V-SNARE 11, Vesicle transport v-SNARE family protein (.1)
Lus10037141
246 / 4e-83
AT1G26670 324 / 1e-113
VESICAL TRANSPORT V-SNARE 12, Vesicle transport v-SNARE family protein (.1)
Lus10036788
208 / 1e-67
AT1G26670 243 / 3e-81
VESICAL TRANSPORT V-SNARE 12, Vesicle transport v-SNARE family protein (.1)
Lus10000565
183 / 3e-58
AT5G39510 178 / 3e-56
SHOOT GRAVITROPSIM 4, VESICLE TRANSPORT V-SNARE 11, Vesicle transport v-SNARE family protein (.1)
Lus10021629
81 / 2e-17
AT5G61380 326 / 4e-103
PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, CCT motif -containing response regulator protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G115700
331 / 1e-116
AT5G39510 325 / 5e-114
SHOOT GRAVITROPSIM 4, VESICLE TRANSPORT V-SNARE 11, Vesicle transport v-SNARE family protein (.1)
Potri.017G087700
330 / 3e-116
AT5G39510 321 / 1e-112
SHOOT GRAVITROPSIM 4, VESICLE TRANSPORT V-SNARE 11, Vesicle transport v-SNARE family protein (.1)
Potri.010G164300
275 / 3e-94
AT1G26670 324 / 9e-114
VESICAL TRANSPORT V-SNARE 12, Vesicle transport v-SNARE family protein (.1)
Potri.008G091000
273 / 1e-93
AT1G26670 327 / 5e-115
VESICAL TRANSPORT V-SNARE 12, Vesicle transport v-SNARE family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0147
Traffic
PF12352
V-SNARE_C
Snare region anchored in the vesicle membrane C-terminus
CL0147
Traffic
PF05008
V-SNARE
Vesicle transport v-SNARE protein N-terminus
Representative CDS sequence
>Lus10021624 pacid=23180302 polypeptide=Lus10021624 locus=Lus10021624.g ID=Lus10021624.BGIv1.0 annot-version=v1.0
ATGAGCGGGGTATTTGAAGGATACGAACGGCAGTATTGTGAGCTTTCTGCAAATCTATCCAAGAGATGCAATTCAGCGTCTGCACTTGATGGAGAGCAAA
AGAAGCAAAAACTTAGTGAAATAAAAGCCGGTTTGGAAGATGCAGAGACTCTGATTCGCAAAATGGACCTCGAGGCAAGAAGTTTGCCCCCAAATGTCAA
GGCTGTGCTTCTGGCGAAGTTAAGGGAATACAAGTCTGATCTCAATAATCTGAAAAGTGAAGTAAAACGGATAGCATCCGGTAACCTCAATGCCGCTGCC
AGAGATGAACTTCTAGAATCAGGCTTCTCTGATGCACTGACGGGTTCAGCCGATCACAGGTCAAGGTTAATGACCTCAACAGCGAGATTGAATAAGACTG
ATGAGAGAATCAAGGATAGTAGGAGAACAATGCTGGAAACCGAAGAGCTTGGTGTTTCACTTCTTCAAGATTTGCATCAGCAACGACAGTCCCTTCTGCG
TGGCCATGACACGCTTCATGGAGTGGATGACAACGTTGGGAGGAGCAAGCGAGTGTTGGGTAACATGAGTAGAAGGATGAACAGAAACAAATGGGTCATT
GGTTCGGTCGTCGCTGTCCTAGTTGTTGCCATCGGCGTGATTCTGTATTTTAAACTCAGATAA
AA sequence
>Lus10021624 pacid=23180302 polypeptide=Lus10021624 locus=Lus10021624.g ID=Lus10021624.BGIv1.0 annot-version=v1.0
MSGVFEGYERQYCELSANLSKRCNSASALDGEQKKQKLSEIKAGLEDAETLIRKMDLEARSLPPNVKAVLLAKLREYKSDLNNLKSEVKRIASGNLNAAA
RDELLESGFSDALTGSADHRSRLMTSTARLNKTDERIKDSRRTMLETEELGVSLLQDLHQQRQSLLRGHDTLHGVDDNVGRSKRVLGNMSRRMNRNKWVI
GSVVAVLVVAIGVILYFKLR
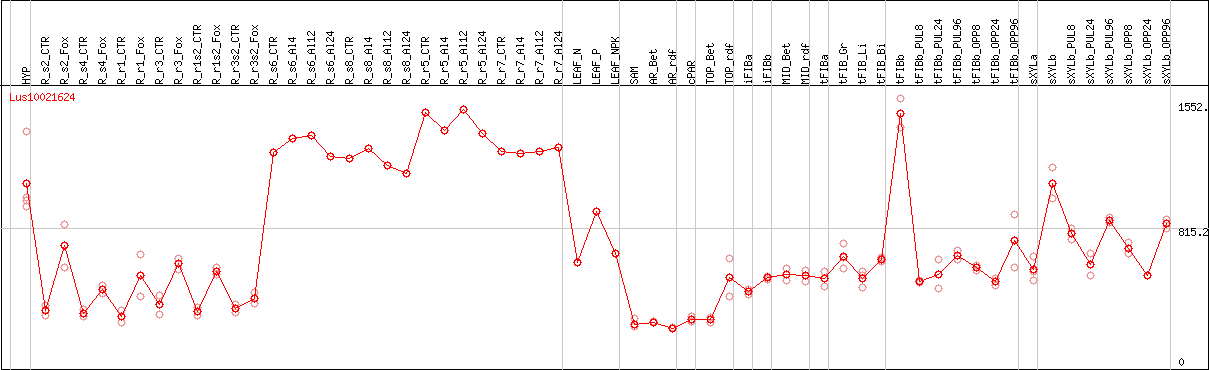
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021624 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.