External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G26570 145 / 7e-43
UGD1, ATUGD1
UDP-glucose dehydrogenase 1 (.1)
AT5G15490 136 / 1e-39
UGD3
UDP-glucose dehydrogenase 3, UDP-glucose 6-dehydrogenase family protein (.1)
AT3G29360 132 / 5e-38
UGD2
UDP-glucose dehydrogenase 2, UDP-glucose 6-dehydrogenase family protein (.1.2)
AT5G39320 129 / 5e-37
UDP-glucose 6-dehydrogenase family protein (.1)
AT3G01010 119 / 8e-36
UDP-glucose/GDP-mannose dehydrogenase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034703
167 / 2e-51
AT5G15490 882 / 0.0
UDP-glucose dehydrogenase 3, UDP-glucose 6-dehydrogenase family protein (.1)
Lus10021626
166 / 1e-50
AT5G15490 875 / 0.0
UDP-glucose dehydrogenase 3, UDP-glucose 6-dehydrogenase family protein (.1)
Lus10026656
139 / 1e-40
AT1G26570 868 / 0.0
UDP-glucose dehydrogenase 1 (.1)
Lus10004656
137 / 1e-39
AT1G26570 864 / 0.0
UDP-glucose dehydrogenase 1 (.1)
Lus10036887
131 / 1e-39
AT3G29360 444 / 5e-157
UDP-glucose dehydrogenase 2, UDP-glucose 6-dehydrogenase family protein (.1.2)
Lus10037096
132 / 3e-38
AT1G26570 815 / 0.0
UDP-glucose dehydrogenase 1 (.1)
Lus10026657
132 / 3e-37
AT1G26570 856 / 0.0
UDP-glucose dehydrogenase 1 (.1)
Lus10004657
58 / 3e-12
AT5G15490 133 / 4e-38
UDP-glucose dehydrogenase 3, UDP-glucose 6-dehydrogenase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G118600
142 / 5e-42
AT5G15490 879 / 0.0
UDP-glucose dehydrogenase 3, UDP-glucose 6-dehydrogenase family protein (.1)
Potri.010G159800
132 / 3e-38
AT5G15490 853 / 0.0
UDP-glucose dehydrogenase 3, UDP-glucose 6-dehydrogenase family protein (.1)
Potri.008G094300
130 / 3e-37
AT3G29360 858 / 0.0
UDP-glucose dehydrogenase 2, UDP-glucose 6-dehydrogenase family protein (.1.2)
Potri.017G092000
129 / 6e-37
AT3G29360 898 / 0.0
UDP-glucose dehydrogenase 2, UDP-glucose 6-dehydrogenase family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0039
HUP
PF03720
UDPG_MGDP_dh_C
UDP-glucose/GDP-mannose dehydrogenase family, UDP binding domain
Representative CDS sequence
>Lus10021639 pacid=23180319 polypeptide=Lus10021639 locus=Lus10021639.g ID=Lus10021639.BGIv1.0 annot-version=v1.0
ATGAGCTGCAACTGGACAAGCGTAAAAGAACAAGTGAGCATAGTGTCGGATGCGTACGAGGCTACGAAGGATGCTCACGGTGTGTACATTCTGACCGAGT
GGGATGAGTTCAAGAAACTATACTACAAAAAGATATACGACAAGATGCAAAAGCCAGCATTTGTGTTTCATGGGAGGAAGGTGGTTGATGCTGCAAAGTT
GAGGGAGATTGGTTTCATTGTTTACTCCATTGGGAAGCCTTTGGATAATTGGCTCCAGGACTTGCCTGGTTTGGCTTGA
AA sequence
>Lus10021639 pacid=23180319 polypeptide=Lus10021639 locus=Lus10021639.g ID=Lus10021639.BGIv1.0 annot-version=v1.0
MSCNWTSVKEQVSIVSDAYEATKDAHGVYILTEWDEFKKLYYKKIYDKMQKPAFVFHGRKVVDAAKLREIGFIVYSIGKPLDNWLQDLPGLA
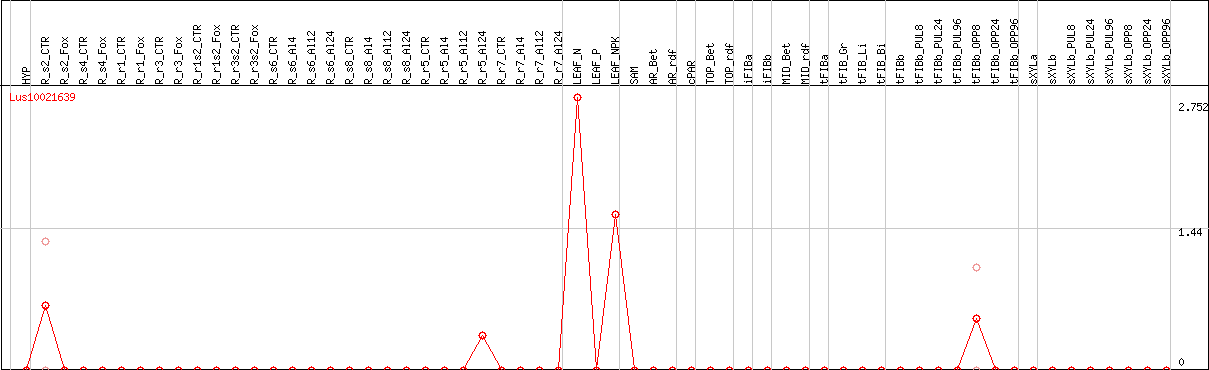
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021639 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.