Lus10021733 [FLAX]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Lus10021733 pacid=23154331 polypeptide=Lus10021733 locus=Lus10021733.g ID=Lus10021733.BGIv1.0 annot-version=v1.0
ATGTATGCCGTGGGGCGAATAGGAAGTTATATAACCAAGGGAGTATCCACGGTGTCTGGCCCATTCCATCCTTTTGGGGGAGCTGTAGACATAGTTGTAG
TGGAGCAAGCAGATGGGAGTTTAAAGTCTTCACCTTGGTATGTCCGATTTGGGAAATTTCAAGGTGTTCTTAAGGCCAAAGAGAAAGTGGTAATCATCAA
CGTTAATGGAGTCGATGCTAGTTTCAACATGTACCTTGATCAAAGAGGCGAGGCTTTCTTTATCCGAGATGAGGATGATGATGATGATGATGATGGAGGA
AATGATTCTGGGTCACATCATCCTATATTTTGTGATGGTATGAATAGACATGGACAAAATGGTTGGAGGCAGAGGCACACCCATAGTCTAGATTTTGATC
ATAGTTTTCCCAATTCAGGAGACTATGAGTTGGCAAGCAATGAGAAGTATATGCTTAGAGCTCGTTCAGGAGGACCAAGGATAATGGGGATTGTTTTTGG
TAGAAGGACAGTCAAGGCAGATAATTATCAACAGGGTGTTGGCCATTCTAGAGTTAGCTCATTGGAGCGTGCTGAGATGGCAGCTGATCTTTTGGAAGTT
AGATGGTCTACTACAGGTATTTCACCTCTCAAGACATCGAATGATAATGATAATAACATAGTGGATATTGTTAAGAGTCGCGACAATGACGAAAGGACCA
ACCCATCTTTGTCAAATCTTGATACTTGTGGAAGTAGCTTGGCTAAAGAAACTACTAGCACGAGCAATGCAGGAACACACACAAGCTCTCAAAATGCCTT
TAAGGAAGCGTTTGTAGATAAAGTTAGTGTAGATGCAAGAGGACAAGTTGTTGAAACTTTCACACAAGTTGTACTGGAAATGCCACAAAATGTCCATGAA
TGTAGGATACGAAATGCCGACCTTGAAGGTAATGATGATGATATCATGTCCAATGGCATCAATCCCGACTCGATAGATTTAGAGTCATCAAAATCACTCC
GTATGGACACAAAAAATAGTGAAGTTCAATCATTTACCATTTGTGAGGCATCTGAGAGCTCATTTGCAACTTTGAAGGGTTCAAATGAAGAAACACATGA
AACTATGTGCATTTCAAATGGGGAGCCCGGTGAAGCAACTTTGCTCGTTGAGGCTGTGCAAATGGTGTCTGAGCAGCTACCTGAGGTAATAGAGATTTGT
CATGGGGAGATGGTTAATGAAGAAACTGGAGAAACTAAGTCAGAGGCGGGCTGTGCCAAGACTTATCATGATGGTTGCTACGAGCAACAACTTTTGTCAC
AATCTTTCTTGTGCAACCATGATAAAGTCCAATCTTCAACAAAGTTCTTTGATGAACCTGACCAGCTTGCTCAATGTTGCGTGAAAAAGAATTCTAGTGT
TCCAGCCTCGGAGTGTTCTGAAGGGGAACAATTTCTCTTCAGTGATCTTGATGAACCAAGTTGCAGGGATATAGGAAATGGCTTAAGTGTCGAAGAACAT
AGTAATTGTCAATCGTATTCAACTGTAGACCAGTTTACTAAAGAGAATCAAGTAATTGATAATGGGAACTCAACAGGAGCGTCCAGTCCCATTTGCATCT
CTAGAGTTCATAATATCGATGATGTGGAAAGTGGTCACTTGGCTGGATCATTACCTAGTGCTTGGTCAGATGTTGAAAGCCGTGATGCTATGACACAACA
TCCATTAAGCCATTCTCTGGAATCGAAATCCACACCTCATTGGAAAGGTAATGAGAATGATGGGTTGGGTTGTGTAGATGTGGATAAAAAAAAGGAAGGT
TCTTTGCAAGAGGATCATCATCACAGAGATACAAATCCAGCCAGTACTGGAAATCTGTCTCAATCTTCTGTTGCTGCTACTACTAATGGAAGCTGGAAAT
TTTGGCCATTTTCTTCTAAGAAGTCAAAACAGAGAAAGATCACCAACACTGATTCAGGACATTCTAATTCTCAAAATTCTTCAGACAGTAAGATTGATAA
TGGAAAGGCAGATGATATTTCTGAGATTAAAATTAAAAATAGGAGTTCCGAGACTACATCAGATGATAAAATCAACGGTATGACTGAGAAAGAAAAGTTA
GAGAATACTTCAAATCGTAAAACTGAGAGTGAGACTTCTCAGAAAGTACCCGATAAGGAAATTGGTAGTAAAACACCACAAGATGCTTCAGAGAGCAAAG
TTGTCATGGGGAAGTCTGATGGTATTTTAGACCATGAAACTGACATTAAAATGGTTAAACCTGACGTTAGGAAAAACAAAATAAGGGCACTCAATCCAAC
GTCCGAACAATTGGAATCATTAAATCTAAACGAGGGCACAAATGTGGTGACATTCACATTCTCAACATCAATGTTGGGGAAACAGAAGGTTGAGGCCAGA
ATTTTTCTATGGAAATGGGATGATCGCATTGTAATCTCAGACGTTGATGGGACAATTACCAAATCTGATGTTCTTGGACAGTTTATGCCTATGGTTGGTA
TGGATTGGTCACAAACGGGAGTTACACGATTGTTTTCTGCGATTAAGGACAATGGGTACAAACTACTTTTTCTTAGTGCCCGGTCGATTTCACAAGCTTA
TACCACTAGACAATTTCTATTCAATCTTAAGCAGGATGGGATGGCATTACCAGACGGTCCTGTTGTTATATCGCCTGATGGACTCTTTCCTTCTTTATAT
CGTGAAGTGATTAGGAGGGCACCTCATGAATTCAAGATTGGGTGTTTAGAGGACATCAAAGCATTGTTTCCTACAAATCACAGCCCATTTTATGCTGGGT
TTGGAAACAGAGACACCGATGAGCTTAGCTATCTCAAAGTTGGAATTCCAAGGGGTAAAATCTTCATCATCAATCCAAAGGGTGAAGTTTCTGTTCACCG
GCGGGTCGACACAAAATCTTACACTTCACTCCACGATCTTGTTGATGAAATGTTCCCGCATCCTGTAACAACTATGGAGCCGGAGGATTTCAACTCATGG
AATTACTGGAGGTTGCCACCTCTTGGCATTAATACGTGA
|
||||||||||||||||||||
|
AA sequence
|
>Lus10021733 pacid=23154331 polypeptide=Lus10021733 locus=Lus10021733.g ID=Lus10021733.BGIv1.0 annot-version=v1.0
MYAVGRIGSYITKGVSTVSGPFHPFGGAVDIVVVEQADGSLKSSPWYVRFGKFQGVLKAKEKVVIINVNGVDASFNMYLDQRGEAFFIRDEDDDDDDDGG
NDSGSHHPIFCDGMNRHGQNGWRQRHTHSLDFDHSFPNSGDYELASNEKYMLRARSGGPRIMGIVFGRRTVKADNYQQGVGHSRVSSLERAEMAADLLEV
RWSTTGISPLKTSNDNDNNIVDIVKSRDNDERTNPSLSNLDTCGSSLAKETTSTSNAGTHTSSQNAFKEAFVDKVSVDARGQVVETFTQVVLEMPQNVHE
CRIRNADLEGNDDDIMSNGINPDSIDLESSKSLRMDTKNSEVQSFTICEASESSFATLKGSNEETHETMCISNGEPGEATLLVEAVQMVSEQLPEVIEIC
HGEMVNEETGETKSEAGCAKTYHDGCYEQQLLSQSFLCNHDKVQSSTKFFDEPDQLAQCCVKKNSSVPASECSEGEQFLFSDLDEPSCRDIGNGLSVEEH
SNCQSYSTVDQFTKENQVIDNGNSTGASSPICISRVHNIDDVESGHLAGSLPSAWSDVESRDAMTQHPLSHSLESKSTPHWKGNENDGLGCVDVDKKKEG
SLQEDHHHRDTNPASTGNLSQSSVAATTNGSWKFWPFSSKKSKQRKITNTDSGHSNSQNSSDSKIDNGKADDISEIKIKNRSSETTSDDKINGMTEKEKL
ENTSNRKTESETSQKVPDKEIGSKTPQDASESKVVMGKSDGILDHETDIKMVKPDVRKNKIRALNPTSEQLESLNLNEGTNVVTFTFSTSMLGKQKVEAR
IFLWKWDDRIVISDVDGTITKSDVLGQFMPMVGMDWSQTGVTRLFSAIKDNGYKLLFLSARSISQAYTTRQFLFNLKQDGMALPDGPVVISPDGLFPSLY
REVIRRAPHEFKIGCLEDIKALFPTNHSPFYAGFGNRDTDELSYLKVGIPRGKIFIINPKGEVSVHRRVDTKSYTSLHDLVDEMFPHPVTTMEPEDFNSW
NYWRLPPLGINT
|
DESeq2's median of ratios [FLAX]
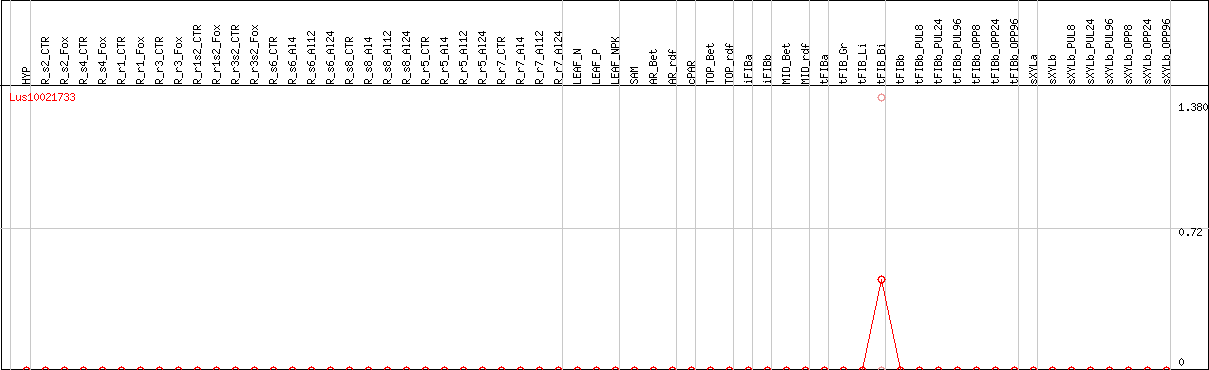
Coexpressed genes
Lus10021733 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
