External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G23020 281 / 6e-88
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G73710 144 / 6e-38
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G46580 83 / 3e-17
pentatricopeptide (PPR) repeat-containing protein (.1)
AT4G16390 73 / 6e-14
SVR7
suppressor of variegation 7, pentatricopeptide (PPR) repeat-containing protein (.1)
AT5G39980 72 / 1e-13
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G48730 71 / 3e-13
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G74750 69 / 1e-12
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G35130 64 / 6e-11
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G18900 64 / 7e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2), Pentatricopeptide repeat (PPR) superfamily protein (.3)
AT5G12100 63 / 1e-10
pentatricopeptide (PPR) repeat-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041172
598 / 0
AT3G23020 953 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10004555
86 / 5e-18
AT5G46580 960 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10000897
83 / 3e-17
AT5G46580 945 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10039071
74 / 4e-14
AT4G16390 808 / 0.0
suppressor of variegation 7, pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10038788
72 / 1e-13
AT4G16390 802 / 0.0
suppressor of variegation 7, pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10041429
72 / 2e-13
AT5G39980 976 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10034342
71 / 3e-13
AT5G39980 970 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10013894
69 / 1e-12
AT5G02860 1047 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10035735
69 / 2e-12
AT4G11690 526 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G078900
338 / 2e-109
AT3G23020 1046 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.013G144800
149 / 9e-40
AT1G73710 1206 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G089600
84 / 1e-17
AT5G46580 986 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.016G006800
75 / 9e-15
AT4G16390 916 / 0.0
suppressor of variegation 7, pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.017G075700
71 / 3e-13
AT5G39980 1045 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G005100
70 / 5e-13
AT4G30825 1121 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G019000
70 / 6e-13
AT4G16390 939 / 0.0
suppressor of variegation 7, pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.004G013300
67 / 4e-12
AT3G53700 1092 / 0.0
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.012G123600
67 / 5e-12
AT2G35130 852 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.014G118500
65 / 2e-11
AT2G31400 1213 / 0.0
genomes uncoupled 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10021891 pacid=23158072 polypeptide=Lus10021891 locus=Lus10021891.g ID=Lus10021891.BGIv1.0 annot-version=v1.0
ATGTTTGCGAAGCTCCAGTTAGATTCCAACTGCTTTCCTGGAGTTACTTCTCTGTCCAACTCTTCCTCACCAACTCAAGGAGTTTCTATCCCCACCAAGT
CTGGAGCTAAGGCCGTTGTAGTGAAGCTCCGAGAACAGAGTATTCTCGAAAATTTTCAAGGAGTTGCAGAAGAGCACAATGGGGATAGGAGAACCAAAGG
GAATCAGAGACCAGGTATTGTATTCGGTCCTGAATTGGAATTCCCAGATAGGAGGAGAACTACGCCGAACCGTTCTGGAACTCGTATGGGGAAGAATCTG
GGAGAAGAATTGAGTGAAGTGCACACCAAGTGTTCGACGAAATGGGCCTTTTACGGAGGTTCTATTCCGGCCATTTTACAAGCGTTGGACACGGTGAAGG
ACTTGGACGAGGCGTTGCATCCATGGGAGCATACTCTCAATAACAAAGTGAGGAGCATAATCTTGAAGGAGCAGTTGTGCTGGGAAAGGGCTCTTGAGAT
CTTTGAGTGGTTTAAGAACAAGAGATGTTATGAGTTGAATGTTATCCACTACAACATTATGCTAAAGATTATGGGGAAAGCGAAAAATTGGAGCCTTGTT
GAATATCTTTGTCAGGAAATGGCGGCCAAACGAATTTCTTTCATCAATTCAACTTACGGCACTCTTATTGATGTATGTAGTAAAGGTGGCCTTAAAGAGA
AGGCACTTTGGTGGCTGGAGAAGATGAACCAGCAAGGGATCCAACCCGATGAGGTGACGGCAGGGATCGTTATGCAAATGTACAAGAAAGCTGGCGAGTT
TCGGAAAGTTGAAGACTTTTTCAAGCAATGGACGTCAATGGATGATGGGTTGCGAGAGGATTATTCGTTAAGTTCGTACACGTACAATACTTTGATAGAC
ACATATGGAAAATCAGGGATTGTGCCAACAACTGTGACCATGAAAAGACACTGTAAATTATCATCTTCCATGAAACCCTTATGTGTTAATTAA
AA sequence
>Lus10021891 pacid=23158072 polypeptide=Lus10021891 locus=Lus10021891.g ID=Lus10021891.BGIv1.0 annot-version=v1.0
MFAKLQLDSNCFPGVTSLSNSSSPTQGVSIPTKSGAKAVVVKLREQSILENFQGVAEEHNGDRRTKGNQRPGIVFGPELEFPDRRRTTPNRSGTRMGKNL
GEELSEVHTKCSTKWAFYGGSIPAILQALDTVKDLDEALHPWEHTLNNKVRSIILKEQLCWERALEIFEWFKNKRCYELNVIHYNIMLKIMGKAKNWSLV
EYLCQEMAAKRISFINSTYGTLIDVCSKGGLKEKALWWLEKMNQQGIQPDEVTAGIVMQMYKKAGEFRKVEDFFKQWTSMDDGLREDYSLSSYTYNTLID
TYGKSGIVPTTVTMKRHCKLSSSMKPLCVN
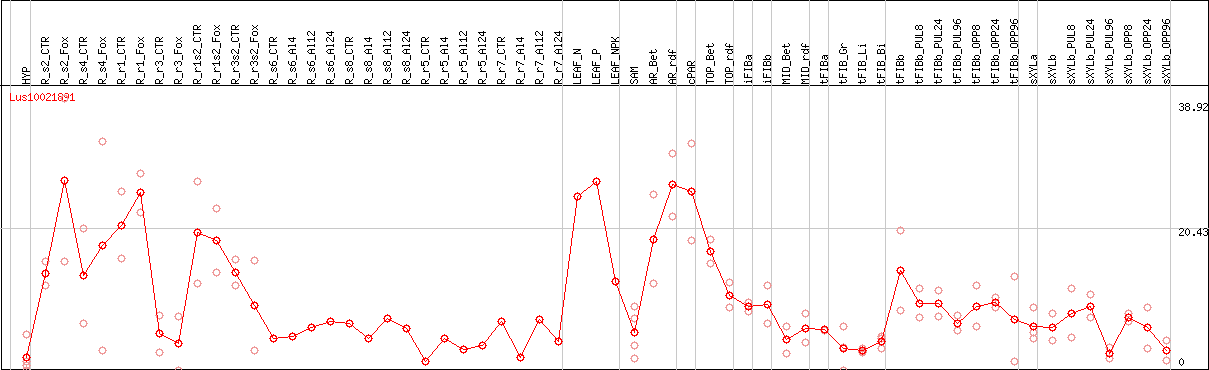
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021891 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.