External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G22700 298 / 2e-102
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
AT3G11540 61 / 1e-10
SPY
SPINDLY, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT3G04240 45 / 3e-05
SEC
secret agent, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041265
400 / 2e-141
AT1G22700 303 / 5e-102
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Lus10021336
60 / 4e-10
AT3G11540 1509 / 0.0
SPINDLY, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10017009
59 / 5e-10
AT3G11540 1356 / 0.0
SPINDLY, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10013661
47 / 6e-06
AT5G63200 801 / 0.0
tetratricopeptide repeat (TPR)-containing protein (.1)
Lus10035117
44 / 5e-05
AT4G30480 291 / 3e-99
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Lus10031983
43 / 0.0001
AT4G30480 222 / 9e-76
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Lus10003576
42 / 0.0002
AT3G04240 1606 / 0.0
secret agent, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10023748
41 / 0.0004
AT4G28600 753 / 0.0
no pollen germination related 2 (.1)
Lus10003113
41 / 0.0006
AT3G04240 1687 / 0.0
secret agent, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G114500
347 / 5e-121
AT1G22700 321 / 7e-110
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Potri.019G085200
343 / 7e-120
AT1G22700 324 / 6e-111
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Potri.016G075700
60 / 2e-10
AT3G11540 1505 / 0.0
SPINDLY, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.006G208900
59 / 5e-10
AT3G11540 1506 / 0.0
SPINDLY, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.005G054800
43 / 0.0001
AT3G04710 611 / 0.0
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
Potri.012G092000
43 / 0.0001
AT5G63200 759 / 0.0
tetratricopeptide repeat (TPR)-containing protein (.1)
Potri.006G178900
42 / 0.0002
AT4G30480 275 / 1e-92
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Potri.010G068700
42 / 0.0002
AT4G28600 708 / 0.0
no pollen germination related 2 (.1)
Potri.018G100700
40 / 0.0006
AT4G30480 303 / 8e-104
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Potri.001G473100
40 / 0.0007
AT5G56290 978 / 0.0
EMBRYO DEFECTIVE 2790, ARABIDOPSIS PEROXIN 5, peroxin 5 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF00515
TPR_1
Tetratricopeptide repeat
Representative CDS sequence
>Lus10021972 pacid=23158033 polypeptide=Lus10021972 locus=Lus10021972.g ID=Lus10021972.BGIv1.0 annot-version=v1.0
ATGAGAAAGTTAATACTTTGGATATCTGCCCTCTCTGTTCAACAAGTTTCGTTCTTGATACCTGCACAAGCAGCAAATGCGAGTGAAAGTATCAGACCAG
GTGCACTATATGATGTCGGCGAGTTATTTGAACTGGGAATCCAGCTATCGTATCTGCTATTGCTGCTGGCGCTGCTTGGTGTTGGAACGTTCTTCGTGAT
TCGCCAAGTTCTTGTTCGTAGAGAACTCGATCTTTCTGCTAAGGAACTACAGGCAGTAAGGAGTGGTGATGCAAGTGCTACCGAGTTGTTTGAACTTGGA
GCTGTGATGTTGAGGAGGAAGTTCTACCCTGCTGCCACCAAATATTTGCTTCAAGCGATTGAGAAATGGGATGGTGACGATCAGGATCTAGCGCAGGTGT
ACAATGCACTTGGGGTGAGTTATATCCGAGACGGAAAGACAGATAAAGGTGTAGCTATGCTTGAGAAGGCAGTGAAGCTTCAGCCTGGGTATGTAACTGC
ATGGAACAACCTGGGAGATGCTTACGAGAAGAAGAATGACATGAAAGGAGCCCTCAAGGCATTCGAGGAAGCGCTGTTGTTTGATCCCAATAATAAGGTG
GCACGGCCACGTAGAGATGGACTTAGGGATCGTGTGCAGATGTACAAAGGCGTGCCTGTGAAATCAAAGGATCGATAA
AA sequence
>Lus10021972 pacid=23158033 polypeptide=Lus10021972 locus=Lus10021972.g ID=Lus10021972.BGIv1.0 annot-version=v1.0
MRKLILWISALSVQQVSFLIPAQAANASESIRPGALYDVGELFELGIQLSYLLLLLALLGVGTFFVIRQVLVRRELDLSAKELQAVRSGDASATELFELG
AVMLRRKFYPAATKYLLQAIEKWDGDDQDLAQVYNALGVSYIRDGKTDKGVAMLEKAVKLQPGYVTAWNNLGDAYEKKNDMKGALKAFEEALLFDPNNKV
ARPRRDGLRDRVQMYKGVPVKSKDR
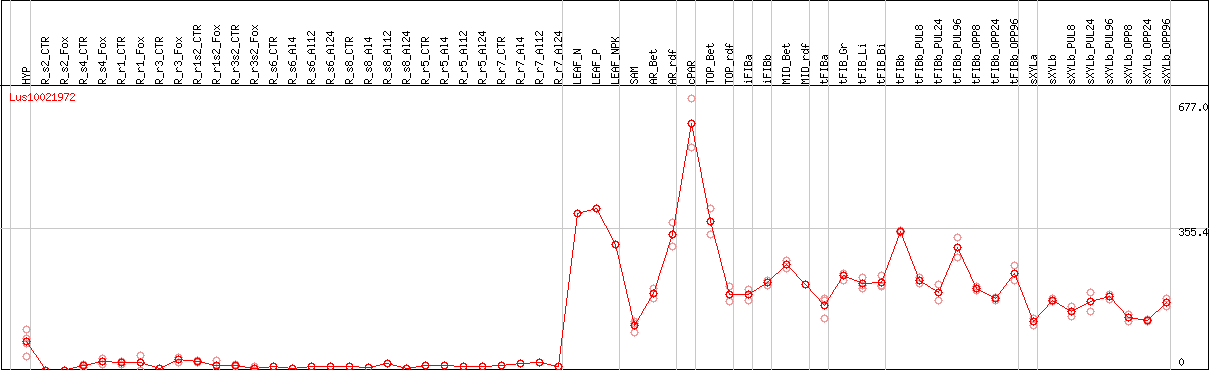
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10021972 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.