External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69490 51 / 1e-08
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT1G61110 51 / 2e-08
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT3G17730 50 / 2e-08
NAC
ANAC057
NAC domain containing protein 57 (.1)
AT5G17260 49 / 1e-07
NAC
ANAC086
NAC domain containing protein 86 (.1)
AT1G77450 48 / 2e-07
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT2G18060 48 / 3e-07
NAC
ANAC037, VND1
Arabidopsis NAC domain containing protein 37, vascular related NAC-domain protein 1 (.1)
AT5G14000 47 / 4e-07
NAC
ANAC084
NAC domain containing protein 84 (.1)
AT5G13180 47 / 4e-07
NAC
VNDIP2, ANAC083, VNI2
VND-interacting 2, NAC domain containing protein 83 (.1)
AT1G54330 47 / 4e-07
NAC
ANAC020
NAC domain containing protein 20 (.1)
AT3G03200 47 / 5e-07
NAC
ANAC045
NAC domain containing protein 45 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001876
87 / 3e-21
AT1G01470 178 / 3e-54
LIGHT STRESS-REGULATED 3, LATE EMBRYOGENESIS ABUNDANT 14, Late embryogenesis abundant protein (.1)
Lus10036117
54 / 2e-09
AT3G15510 241 / 2e-78
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10007410
54 / 2e-09
AT3G15510 241 / 1e-78
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10042284
52 / 1e-08
AT3G17730 348 / 1e-121
NAC domain containing protein 57 (.1)
Lus10043095
51 / 2e-08
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10026373
51 / 2e-08
AT3G17730 348 / 2e-121
NAC domain containing protein 57 (.1)
Lus10032657
51 / 3e-08
AT3G15510 353 / 1e-120
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10031951
50 / 4e-08
AT3G04070 56 / 3e-09
NAC domain containing protein 47 (.1.2)
Lus10011215
50 / 5e-08
AT1G61110 305 / 1e-102
NAC domain containing protein 25 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10022018 pacid=23160214 polypeptide=Lus10022018 locus=Lus10022018.g ID=Lus10022018.BGIv1.0 annot-version=v1.0
ATGGAATCAACCAAAGGTTATACATATCCGGTGGGTATTCGCTTTAATCCGAAGGAGGAAGAACTCGTCAATTATTACCTCAAGGGTAAAGCCAAAGGGG
AGCCTCTGCCTTCGAACCTGATACATAAGATCGACCTCTACGGTGCTAAAACTCCTTGGGAGATCTTTCCTGGCAATAAAAGAATTAAGGATGCTAGGGA
GAGACAAATTGGAATAAAGAAGACATTTAGTTTCAAAGTGACGAGTGAAAAAGCAAAGGTGAAAGGAAAGAGTTGTGCTTCTTCTTCTTCTACTGCTGTT
GCTGAAGGTTCTTCTGGTTGGATTATGCACGAATATTCCCTATTAGGATAG
AA sequence
>Lus10022018 pacid=23160214 polypeptide=Lus10022018 locus=Lus10022018.g ID=Lus10022018.BGIv1.0 annot-version=v1.0
MESTKGYTYPVGIRFNPKEEELVNYYLKGKAKGEPLPSNLIHKIDLYGAKTPWEIFPGNKRIKDARERQIGIKKTFSFKVTSEKAKVKGKSCASSSSTAV
AEGSSGWIMHEYSLLG
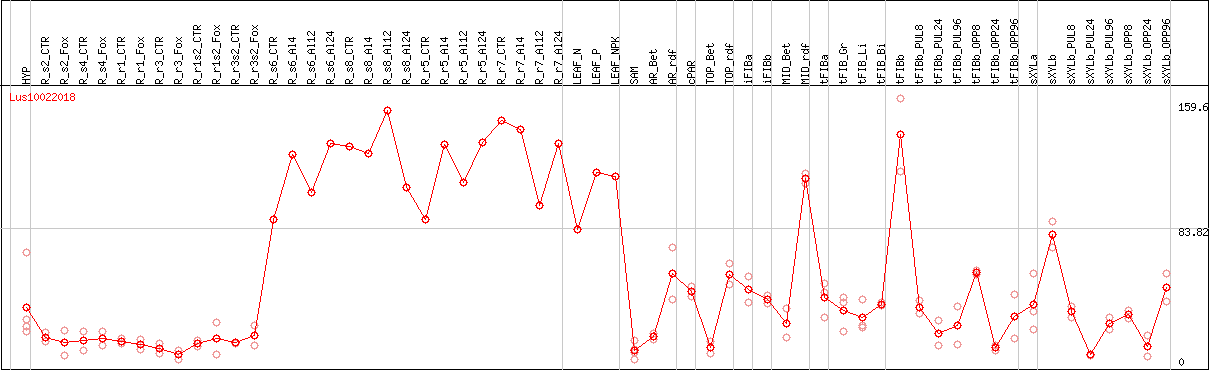
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022018 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.