External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G51550 49 / 5e-07
FER
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
AT1G30570 43 / 4e-05
HERK2
hercules receptor kinase 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032976
143 / 9e-40
AT3G51550 682 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Lus10003942
136 / 1e-37
AT3G51550 656 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Lus10032971
135 / 3e-37
AT3G51550 706 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Lus10014185
47 / 2e-06
AT3G51550 1254 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Lus10016907
45 / 9e-06
AT3G04690 967 / 0.0
ANXUR1, Malectin/receptor-like protein kinase family protein (.1)
Lus10037768
44 / 3e-05
AT3G04690 400 / 1e-147
ANXUR1, Malectin/receptor-like protein kinase family protein (.1)
Lus10029945
0 / 1
AT3G51550 424 / 1e-137
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G095566
52 / 4e-08
AT3G51550 270 / 2e-81
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.017G095900
50 / 2e-07
AT3G51550 845 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.017G096600
49 / 4e-07
AT3G51550 829 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.017G096800
48 / 7e-07
AT3G51550 823 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.017G097001
48 / 7e-07
AT3G51550 506 / 2e-168
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.017G096000
47 / 1e-06
AT3G51550 832 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.016G138700
47 / 2e-06
AT3G51550 1252 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
Potri.008G105500
45 / 7e-06
AT3G04690 999 / 0.0
ANXUR1, Malectin/receptor-like protein kinase family protein (.1)
Potri.010G145600
44 / 2e-05
AT3G04690 1027 / 0.0
ANXUR1, Malectin/receptor-like protein kinase family protein (.1)
Potri.006G110000
42 / 0.0001
AT3G51550 1256 / 0.0
FERONIA, Malectin/receptor-like protein kinase family protein (.1)
PFAM info
Representative CDS sequence
>Lus10022104 pacid=23146477 polypeptide=Lus10022104 locus=Lus10022104.g ID=Lus10022104.BGIv1.0 annot-version=v1.0
ATGGCTTTCTGGGTCAACATCATGAATCAAACGGATGGGGGTTATGTGGATGTCATATATGAAGCAGGTGAGAACGAAATCCCTCTGTTCAAAGATTACT
TGTCGAGAATGGAAGGAGCTAAGAATAAGAACCTCTTCATCGCTTTTTCTCCTTATGAGGTGAACAACGCCTACATAATTTCCATCCTCAATGGAGTTGA
GATCTACAAATTGAGCTCCTCCAACAATCTAGTAGTCAGACCCAACCCTAGCCTGCCACCACTTCAATCCCAGTCCTCCTCATCAAAATCCGGTAGAAAC
CAGACAAAATTAGTCGCTACCGTAGCCGGCAGCATTCTTTCAGGTTTGCTGATGCTCTCCCTTTTACTCTTGTTCATCATTAGGCAGAAACGGAAAGTTA
AGGGCCGAAATGTTGGAGGCTCTACTTCTCCTTCTACAGGGTTTGCAAAATTTTCTTTCATTCAATCAACGTTGCCCTCAATAAAATTTGCTGTCGATTT
TCACTAA
AA sequence
>Lus10022104 pacid=23146477 polypeptide=Lus10022104 locus=Lus10022104.g ID=Lus10022104.BGIv1.0 annot-version=v1.0
MAFWVNIMNQTDGGYVDVIYEAGENEIPLFKDYLSRMEGAKNKNLFIAFSPYEVNNAYIISILNGVEIYKLSSSNNLVVRPNPSLPPLQSQSSSSKSGRN
QTKLVATVAGSILSGLLMLSLLLLFIIRQKRKVKGRNVGGSTSPSTGFAKFSFIQSTLPSIKFAVDFH
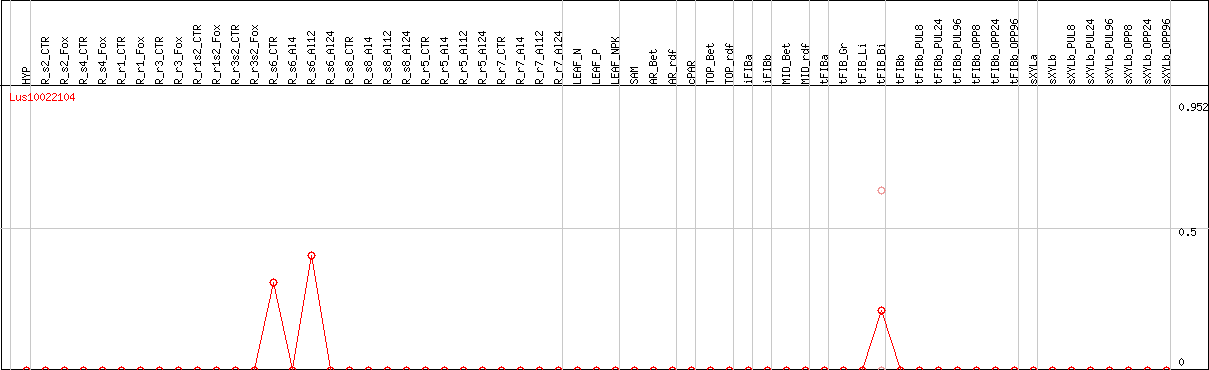
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022104 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.