External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G18430 257 / 4e-89
Calcium-binding EF-hand family protein (.1.2)
AT3G51920 60 / 8e-12
CML9, CAM9, ATCML9
CALMODULIN LIKE PROTEIN 9, calmodulin 9 (.1)
AT3G22930 55 / 8e-10
CML11
calmodulin-like 11 (.1)
AT4G16350 50 / 1e-07
SCABP2, CBL6
SOS3-LIKE CALCIUM BINDING PROTEIN 2, calcineurin B-like protein 6 (.1)
AT1G24620 47 / 7e-07
EF hand calcium-binding protein family (.1)
AT4G01420 45 / 4e-06
CBL5
calcineurin B-like protein 5 (.1)
AT1G64480 45 / 5e-06
CBL8
calcineurin B-like protein 8 (.1)
AT4G33000 43 / 3e-05
SCABP8, ATCBL10, CBL10
SOS3-LIKE CALCIUM BINDING PROTEIN 8, calcineurin B-like protein 10 (.1.2.3)
AT5G24270 42 / 6e-05
ATSOS3, CBL4, SOS3
CALCINEURIN B-LIKE PROTEIN 4, SALT OVERLY SENSITIVE 3, Calcium-binding EF-hand family protein (.1.2)
AT5G37770 41 / 0.0001
CML24, TCH2
TOUCH 2, CALMODULIN-LIKE 24, EF hand calcium-binding protein family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010775
322 / 1e-114
AT3G18430 273 / 6e-95
Calcium-binding EF-hand family protein (.1.2)
Lus10029670
244 / 1e-83
AT3G18430 277 / 1e-96
Calcium-binding EF-hand family protein (.1.2)
Lus10042708
236 / 8e-81
AT3G18430 271 / 4e-94
Calcium-binding EF-hand family protein (.1.2)
Lus10011028
50 / 6e-08
AT4G17615 359 / 5e-128
SOS3-LIKE CALCIUM BINDING PROTEIN 5, ARABIDOPSIS THALIANA CALCINEURIN B-LIKE PROTEIN, calcineurin B-like protein 1 (.1.2)
Lus10001816
49 / 3e-07
AT1G64480 271 / 3e-93
calcineurin B-like protein 8 (.1)
Lus10038764
48 / 6e-07
AT5G55990 375 / 5e-134
calcineurin B-like protein 2 (.1)
Lus10003191
47 / 1e-06
AT1G64480 264 / 2e-90
calcineurin B-like protein 8 (.1)
Lus10039098
47 / 2e-06
AT5G55990 374 / 1e-128
calcineurin B-like protein 2 (.1)
Lus10037648
45 / 9e-06
AT4G33000 328 / 2e-114
SOS3-LIKE CALCIUM BINDING PROTEIN 8, calcineurin B-like protein 10 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0220
EF_hand
PF00036
EF-hand_1
EF hand
Representative CDS sequence
>Lus10022173 pacid=23156289 polypeptide=Lus10022173 locus=Lus10022173.g ID=Lus10022173.BGIv1.0 annot-version=v1.0
ATGGGGAATTCGTCCTCGATGCTGACTCAGTACGACATCGAAGAGGTCCAGGAGCACTGTTATCACGCATTTTCGCAGCAAGAGATCGTCTCGCTGTACC
AGAGGTTTTGCCAACTCGATCGCAGCAGTGGTGGGTTTATATCCGCCGACGAGTTCCTCTCCGTGCCCGAGTTCGCCGTCAATCCTCTCGCTCAGAGACT
GCTGAGAACGGTTGATGGATTGAATTTCAAAGAATTCGTTGCTTTCCTGTCTGCATTCAGCCCTGGCGCTACTTTACAACAGAAAATCGAATTCATCTTT
AAGGTATATGATTCGGATGGAAATGGCTCAGTTTCGTTCAAAGAAATATTGGATGTGTTGCGCGATTTGACTGGACATTTTATATCTGAAGAACAGCGGG
AGCAAGTGCTGAATCAGGTCCTTGAAGAAGCAGGTTACACAAAGGACTCGTTATTAGTTTTATCTGACTTTGTAAAGGTACTGTGA
AA sequence
>Lus10022173 pacid=23156289 polypeptide=Lus10022173 locus=Lus10022173.g ID=Lus10022173.BGIv1.0 annot-version=v1.0
MGNSSSMLTQYDIEEVQEHCYHAFSQQEIVSLYQRFCQLDRSSGGFISADEFLSVPEFAVNPLAQRLLRTVDGLNFKEFVAFLSAFSPGATLQQKIEFIF
KVYDSDGNGSVSFKEILDVLRDLTGHFISEEQREQVLNQVLEEAGYTKDSLLVLSDFVKVL
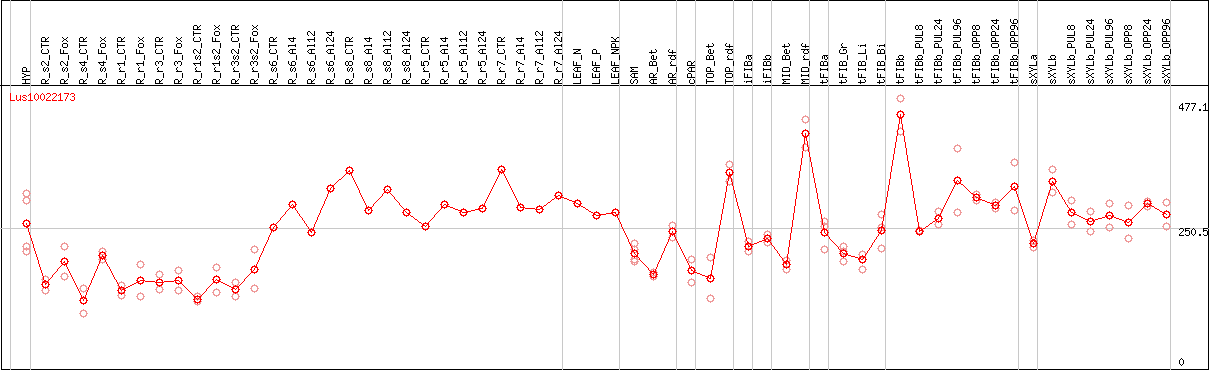
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022173 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.