External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G43540 66 / 6e-15
C2H2ZnF
C2H2 and C2HC zinc fingers superfamily protein (.1)
AT3G09290 65 / 3e-14
C2H2ZnF
TAC1
telomerase activator1 (.1)
AT3G53820 59 / 3e-12
C2H2ZnF
C2H2 and C2HC zinc fingers superfamily protein (.1)
AT3G23130 59 / 5e-12
C2H2ZnF
FLO10, FON1, SUP
SUPERMAN, FLORAL ORGAN NUMBER 1, FLORAL DEFECTIVE 10, C2H2 and C2HC zinc fingers superfamily protein (.1)
AT4G17810 59 / 8e-12
C2H2ZnF
ZFP12
C2H2 and C2HC zinc fingers superfamily protein (.1)
AT2G37740 59 / 1e-11
C2H2ZnF
ATZFP10, ZFP10
zinc-finger protein 10 (.1)
AT2G42410 59 / 1e-11
C2H2ZnF
ATZFP11, ZFP11
zinc finger protein 11 (.1)
AT5G06070 58 / 2e-11
C2H2ZnF
RAB, RBE
RABBIT EARS, C2H2 and C2HC zinc fingers superfamily protein (.1)
AT3G23140 57 / 4e-11
C2H2ZnF
URO
UPRIGHT ROSETTE, C2H2 and C2HC zinc fingers superfamily protein (.1)
AT5G27880 47 / 1e-07
C2H2ZnF
C2H2 and C2HC zinc fingers superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021213
107 / 8e-31
AT3G09290 73 / 2e-16
telomerase activator1 (.1)
Lus10041160
74 / 1e-17
AT5G43540 69 / 8e-15
C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10021881
72 / 5e-17
AT5G43540 67 / 3e-14
C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10040767
66 / 2e-14
AT3G09290 71 / 5e-15
telomerase activator1 (.1)
Lus10033867
64 / 6e-14
AT5G43540 65 / 1e-13
C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10014755
63 / 1e-13
AT5G43540 63 / 5e-13
C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10024363
61 / 8e-13
AT3G53820 69 / 3e-15
C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10021215
60 / 7e-12
AT3G23130 102 / 1e-25
SUPERMAN, FLORAL ORGAN NUMBER 1, FLORAL DEFECTIVE 10, C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10030967
59 / 1e-11
AT4G17810 112 / 1e-30
C2H2 and C2HC zinc fingers superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10022204 pacid=23139418 polypeptide=Lus10022204 locus=Lus10022204.g ID=Lus10022204.BGIv1.0 annot-version=v1.0
ATGGCGCCGAAAAAGCAACGCAGCTCATCGGACTCATCCAGCGAAGACACCGTAAACGACACGGTGGTTTCGCTGCCGAAACCCAAACGCTCGTACGAGT
GCAGTTTCTGCAAGAGAGGGTTCACCAACGCCCAGGCACTAGGCGGCCACATGAACATCCACCGCCGCGACCGCAAAAAGTCGACAGCTTCTGCCGCCGC
CGCCAACGTCCGCCAGCCGGTTGGCCCNNNNNNCCCCCCAGCCGGTTGGCCCCTTCGCCGTTTGGGATGCTCAATATCGCCACTTGTACTTTGA
AA sequence
>Lus10022204 pacid=23139418 polypeptide=Lus10022204 locus=Lus10022204.g ID=Lus10022204.BGIv1.0 annot-version=v1.0
MAPKKQRSSSDSSSEDTVNDTVVSLPKPKRSYECSFCKRGFTNAQALGGHMNIHRRDRKKSTASAAAANVRQPVGXXXPPAGWPLRRLGCSISPLVL
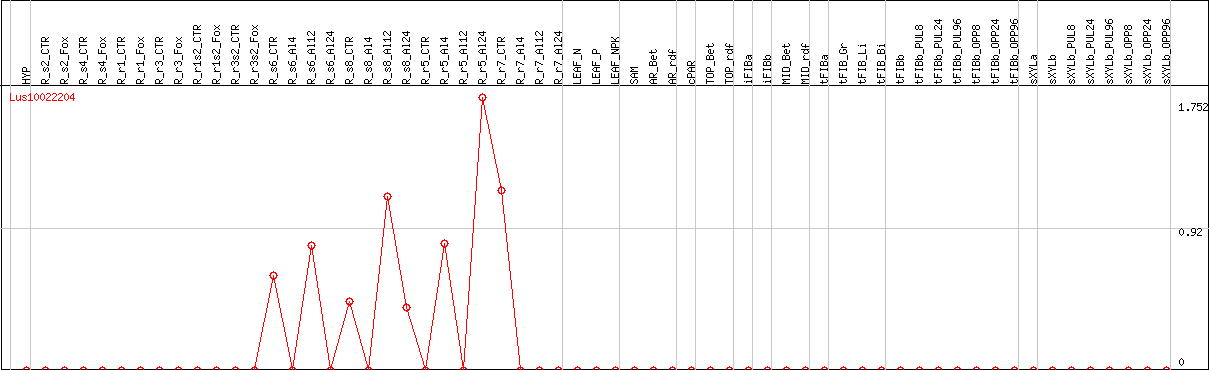
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022204 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.