External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G13180 291 / 2e-95
NOL1/NOP2/sun family protein / antitermination NusB domain-containing protein (.1)
AT5G55920 76 / 4e-15
OLI2
OLIGOCELLULA 2, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT4G26600 73 / 4e-14
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G26180 61 / 4e-10
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT1G06560 50 / 8e-07
NOL1/NOP2/sun family protein (.1)
AT4G40000 49 / 3e-06
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT2G22400 47 / 9e-06
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G66180 42 / 0.0002
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016622
382 / 1e-130
AT3G13180 759 / 0.0
NOL1/NOP2/sun family protein / antitermination NusB domain-containing protein (.1)
Lus10032368
75 / 1e-14
AT4G26600 736 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10023095
75 / 1e-14
AT4G26600 736 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10025004
74 / 2e-14
AT5G26180 566 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10026452
70 / 5e-13
AT5G26180 567 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10020761
52 / 4e-07
AT1G06560 714 / 0.0
NOL1/NOP2/sun family protein (.1)
Lus10007337
50 / 1e-06
AT1G06560 725 / 0.0
NOL1/NOP2/sun family protein (.1)
Lus10026840
45 / 4e-05
AT4G40000 954 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10020217
45 / 5e-05
AT2G22400 975 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G368500
303 / 7e-100
AT3G13180 770 / 0.0
NOL1/NOP2/sun family protein / antitermination NusB domain-containing protein (.1)
Potri.001G369600
79 / 3e-16
AT4G26600 696 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.006G224000
73 / 3e-14
AT5G26180 585 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.005G204900
52 / 3e-07
AT1G06560 758 / 0.0
NOL1/NOP2/sun family protein (.1)
Potri.007G096100
48 / 7e-06
AT2G22400 1012 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01189
Methyltr_RsmB-F
16S rRNA methyltransferase RsmB/F
Representative CDS sequence
>Lus10022523 pacid=23166914 polypeptide=Lus10022523 locus=Lus10022523.g ID=Lus10022523.BGIv1.0 annot-version=v1.0
ATGGCGCGTCCGCTTTCCCTCCATGCCTGTCTCATTGCTGAGACGCGGGACGCCGCAGCCCCAAGACCGCAGAGATTCTCAAAGACTGCGCACAAGCTCA
AGGCATCGGTTTCCATCCCTGAAAACGGTCACCGATGTGGTTGGAGGTACCATTCGTTGGAGGAAAACGTTAGACTAGCAAAGCTTGCCCTTAGACCCGG
TGCTGGTAATCTGGTGAATGGAATTCTTAGAAAGCTATTGTTGCTCAAGGAAAGTGACTCCCTTCCTCTACCCAAAGTTGAAGGTGATGATCGGGCACAA
GCACGTGCTCTTGCCACTATTCATTCTCATCCTGTGTGGATGGTTAGGCGTTGGACTAAGTATATTGGCCAAGAAGATACAGTTAGGTTACTGGTTTGGA
ACAACAGTGACCCCAGTTTCAGCTTAAGGGCAAACTGTGGAAAAGAAGTTGCAAGAGCTGAGCTTGTGAAGCAACTTCATCTGTTGAAGCTACAAGGTCT
ATGTGCAGTTCAGGACGAGAGTGCAGGCATGGTAGTTTCTGTTGTGGATCCTCAACCAAATGAGACCATTATCGACTGTTGTGCGGCTCCTGGAGGGAAG
ACACTATATATAGCATCTTTGATGAAAAATAAAGGCAAGGTAAATGCAATTGATGTGAACAAAGGCTGGTTGCGAATTCTTAAGGAGACCGCCAAGGTGC
ATCAAGTTGATAGGATGATCACCACAGTAGCTGCTGATCTTCGTGAATTTTCCGAGCAAACTAAAGAGAAATATGATAAAGTTTTGCTCGATGTTCCATG
TTCTGGGCTTGGTGTGCTCTCCAAGATTCTGCAGGAGAGAAAGTTTGAAGGTTCCTCGTCGACAAGCAGTAAGGAGAAGCGACGTGGGATGGTGCTGGTG
CTTGCTAAATGA
AA sequence
>Lus10022523 pacid=23166914 polypeptide=Lus10022523 locus=Lus10022523.g ID=Lus10022523.BGIv1.0 annot-version=v1.0
MARPLSLHACLIAETRDAAAPRPQRFSKTAHKLKASVSIPENGHRCGWRYHSLEENVRLAKLALRPGAGNLVNGILRKLLLLKESDSLPLPKVEGDDRAQ
ARALATIHSHPVWMVRRWTKYIGQEDTVRLLVWNNSDPSFSLRANCGKEVARAELVKQLHLLKLQGLCAVQDESAGMVVSVVDPQPNETIIDCCAAPGGK
TLYIASLMKNKGKVNAIDVNKGWLRILKETAKVHQVDRMITTVAADLREFSEQTKEKYDKVLLDVPCSGLGVLSKILQERKFEGSSSTSSKEKRRGMVLV
LAK
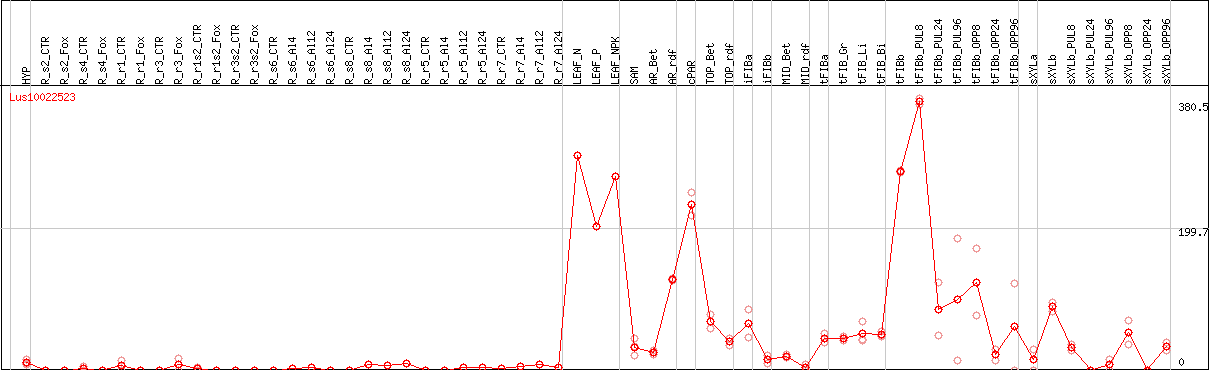
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022523 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.