External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G60660 248 / 3e-83
PIP2F, PIP2;4
plasma membrane intrinsic protein 2;4 (.1)
AT4G35100 245 / 3e-82
SIMIP, PIP3A, PIP2;7, PIP3
PLASMA MEMBRANE INTRINSIC PROTEIN 3A, PLASMA MEMBRANE INTRINSIC PROTEIN 2;7, plasma membrane intrinsic protein 3 (.1.2)
AT2G39010 240 / 5e-80
PIP2E, PIP2;6
PLASMA MEMBRANE INTRINSIC PROTEIN 2;6, plasma membrane intrinsic protein 2E (.1)
AT2G16850 239 / 7e-80
PIP3B, PIP2;8
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
AT2G37170 239 / 9e-80
PIP2;2, PIP2B
PLASMA MEMBRANE INTRINSIC PROTEIN 2;2, plasma membrane intrinsic protein 2 (.1)
AT3G53420 239 / 1e-79
PIP2;1, PIP2A
PLASMA MEMBRANE INTRINSIC PROTEIN 2;1, plasma membrane intrinsic protein 2A (.1.2)
AT2G37180 234 / 6e-78
PIP2C, PIP2;3, RD28
RESPONSIVE TO DESICCATION 28, PLASMA MEMBRANE INTRINSIC PROTEIN 2C, PLASMA MEMBRANE INTRINSIC PROTEIN 2;3, Aquaporin-like superfamily protein (.1)
AT3G54820 234 / 9e-78
PIP2D, PIP2;5
PLASMA MEMBRANE INTRINSIC PROTEIN 2D, plasma membrane intrinsic protein 2;5 (.1)
AT1G01620 216 / 2e-71
TMP-B, PIP1C, PIP1;3
PLASMA MEMBRANE INTRINSIC PROTEIN 1;3, plasma membrane intrinsic protein 1C (.1.2)
AT4G23400 215 / 2e-70
PIP1D, PIP1;5
plasma membrane intrinsic protein 1;5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021475
355 / 7e-126
AT2G39010 265 / 4e-89
PLASMA MEMBRANE INTRINSIC PROTEIN 2;6, plasma membrane intrinsic protein 2E (.1)
Lus10015083
261 / 6e-90
AT5G60660 290 / 2e-100
plasma membrane intrinsic protein 2;4 (.1)
Lus10026504
260 / 3e-89
AT5G60660 358 / 1e-126
plasma membrane intrinsic protein 2;4 (.1)
Lus10019934
259 / 2e-87
AT2G37170 474 / 6e-171
PLASMA MEMBRANE INTRINSIC PROTEIN 2;2, plasma membrane intrinsic protein 2 (.1)
Lus10023184
254 / 2e-85
AT2G37170 490 / 3e-177
PLASMA MEMBRANE INTRINSIC PROTEIN 2;2, plasma membrane intrinsic protein 2 (.1)
Lus10039222
239 / 8e-80
AT2G16850 518 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Lus10027467
239 / 1e-79
AT2G16850 518 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Lus10014978
238 / 2e-79
AT2G16850 514 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Lus10023515
231 / 2e-76
AT3G54820 482 / 5e-174
PLASMA MEMBRANE INTRINSIC PROTEIN 2D, plasma membrane intrinsic protein 2;5 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G128200
258 / 4e-87
AT2G37170 490 / 3e-177
PLASMA MEMBRANE INTRINSIC PROTEIN 2;2, plasma membrane intrinsic protein 2 (.1)
Potri.006G128000
257 / 1e-86
AT2G37170 487 / 5e-176
PLASMA MEMBRANE INTRINSIC PROTEIN 2;2, plasma membrane intrinsic protein 2 (.1)
Potri.009G013900
255 / 4e-86
AT2G37170 458 / 8e-165
PLASMA MEMBRANE INTRINSIC PROTEIN 2;2, plasma membrane intrinsic protein 2 (.1)
Potri.008G039600
245 / 5e-82
AT3G53420 445 / 1e-159
PLASMA MEMBRANE INTRINSIC PROTEIN 2;1, plasma membrane intrinsic protein 2A (.1.2)
Potri.010G222700
244 / 1e-81
AT3G54820 484 / 6e-175
PLASMA MEMBRANE INTRINSIC PROTEIN 2D, plasma membrane intrinsic protein 2;5 (.1)
Potri.004G176300
242 / 6e-81
AT4G35100 513 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3A, PLASMA MEMBRANE INTRINSIC PROTEIN 2;7, plasma membrane intrinsic protein 3 (.1.2)
Potri.016G089500
231 / 2e-76
AT2G37180 458 / 1e-164
RESPONSIVE TO DESICCATION 28, PLASMA MEMBRANE INTRINSIC PROTEIN 2C, PLASMA MEMBRANE INTRINSIC PROTEIN 2;3, Aquaporin-like superfamily protein (.1)
Potri.009G136600
230 / 3e-76
AT4G35100 493 / 1e-178
PLASMA MEMBRANE INTRINSIC PROTEIN 3A, PLASMA MEMBRANE INTRINSIC PROTEIN 2;7, plasma membrane intrinsic protein 3 (.1.2)
Potri.005G109300
217 / 4e-71
AT2G16850 413 / 4e-147
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Potri.006G098100
215 / 4e-70
AT4G00430 482 / 4e-174
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00230
MIP
Major intrinsic protein
Representative CDS sequence
>Lus10022577 pacid=23178558 polypeptide=Lus10022577 locus=Lus10022577.g ID=Lus10022577.BGIv1.0 annot-version=v1.0
ATGGCAAAGGACGTGGAAGCCGCCGATAAACCAGCCGCCGAGTTCTCAGCCAAGGACTACCACGACCCACCGCCGGCACCCATCTTCGCTCTCGAAGAGC
TCACCAAATGGTCCTTTTACAGAGCCCTCATCGCCGAGTTCATCGCCACCCTCCTCTTCCTCTACATCACCGTCCTCACCGTCATCGGCTACAAGTATGG
TGGTGGAGCTAATGAGCTGGCTAGTGGGTATAATAAGGGAACCGGTCTGGGTGCTGAGATTATCGGAACGTTCGTTTTGGTTTATACCGTTTTCTCGGCT
ACTGACCCCAAGAGGAGCGCCAGAGACTCCCATGTTCCTGTATTGGCACCACTTCCAATTGGGTTTGCAGTGTTCATGGTGCACTTGGCAACAATCCCAA
TAACTGGGACTGGAATCAACCCTGCTAGGAGTTTTGGTGCTGCTGTTATCTACAACCAAGACAAGGCTTGGGATGATCAGTGGATATTTTGGGTGGGGCC
GTTCATTGGAGCTGCAATAGCTGCTTTGTACCACCAATACATCCTAAGAGCAGCAGCAATCAAGGCATTGGGTTCATTCAGGAGCAATGCTTGA
AA sequence
>Lus10022577 pacid=23178558 polypeptide=Lus10022577 locus=Lus10022577.g ID=Lus10022577.BGIv1.0 annot-version=v1.0
MAKDVEAADKPAAEFSAKDYHDPPPAPIFALEELTKWSFYRALIAEFIATLLFLYITVLTVIGYKYGGGANELASGYNKGTGLGAEIIGTFVLVYTVFSA
TDPKRSARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVIYNQDKAWDDQWIFWVGPFIGAAIAALYHQYILRAAAIKALGSFRSNA
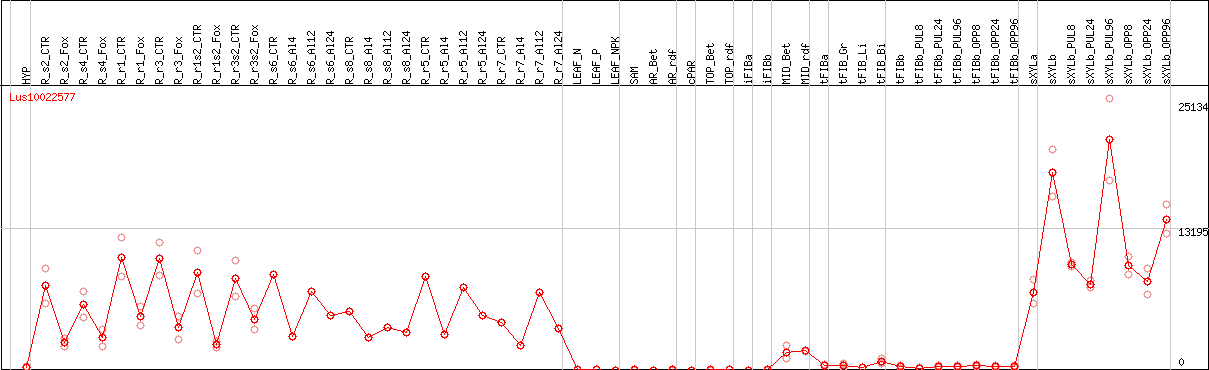
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022577 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.