External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G09890 181 / 1e-58
Ankyrin repeat family protein (.1.2)
AT2G28840 66 / 4e-13
XBAT31
XB3 ortholog 1 in Arabidopsis thaliana (.1.2)
AT5G40160 64 / 8e-13
EMB139, EMB506
embryo defective 506, EMBRYO DEFECTIVE 139, Ankyrin repeat family protein (.1)
AT2G03430 62 / 5e-12
Ankyrin repeat family protein (.1)
AT2G25600 62 / 9e-12
AKT6, SPIK
Shaker pollen inward K+ channel, Shaker pollen inward K+ channel (.1)
AT4G19150 60 / 2e-11
Ankyrin repeat family protein (.1.2)
AT4G32500 56 / 1e-09
AKT5
K+ transporter 5, K+ transporter 5, K+ transporter 5 (.1)
AT2G26650 56 / 1e-09
AKT1, ATAKT1
K+ transporter 1, K+ transporter 1, K+ transporter 1 (.1)
AT5G12320 54 / 1e-09
ankyrin repeat family protein (.1)
AT5G57740 54 / 8e-09
XBAT32
XB3 ortholog 2 in Arabidopsis thaliana (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021508
254 / 9e-88
AT3G09890 209 / 5e-69
Ankyrin repeat family protein (.1.2)
Lus10023911
226 / 2e-76
AT3G09890 219 / 6e-73
Ankyrin repeat family protein (.1.2)
Lus10014410
221 / 3e-74
AT3G09890 217 / 5e-72
Ankyrin repeat family protein (.1.2)
Lus10014522
69 / 3e-14
AT5G40160 322 / 3e-109
embryo defective 506, EMBRYO DEFECTIVE 139, Ankyrin repeat family protein (.1)
Lus10032165
65 / 9e-13
AT5G40160 293 / 3e-98
embryo defective 506, EMBRYO DEFECTIVE 139, Ankyrin repeat family protein (.1)
Lus10033052
61 / 2e-11
AT2G26650 1211 / 0.0
K+ transporter 1, K+ transporter 1, K+ transporter 1 (.1)
Lus10017765
61 / 3e-11
AT2G26650 395 / 1e-130
K+ transporter 1, K+ transporter 1, K+ transporter 1 (.1)
Lus10040829
57 / 3e-10
AT2G28840 546 / 0.0
XB3 ortholog 1 in Arabidopsis thaliana (.1.2)
Lus10016561
56 / 9e-10
AT2G28840 640 / 0.0
XB3 ortholog 1 in Arabidopsis thaliana (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0465
Ank
PF00023
Ank
Ankyrin repeat
Representative CDS sequence
>Lus10022609 pacid=23178669 polypeptide=Lus10022609 locus=Lus10022609.g ID=Lus10022609.BGIv1.0 annot-version=v1.0
ATGATTTCTAATGGAGACATCAAAATAACAAACAACATTGACGAGCCATTGGAAGATGGCGACACGGCTCTCCATCCGGCTTGTCTGTATGGATACTCGA
GCTGTGTTAGGGTCCTGTTGGAACTGGGAGCAAACATAGAGGATAAGGATGAAGATGGGGCAATCCCTCTACATGATGCTTCTGCTGGTGCTATGGCAGG
ATATACTGAGATAGTGCAGCTTCTGCTAAACCATGCTAATGATGCTGAGGCTGTGAAGAGGATGCTAGAGACAATGGATGACGAGGGTGATACTCCACTC
CACCACGCTGCGAGAGGGGAGCACGCGGAGGTCGTGAACCTGTTGCTGGCTTCTGGCGCTTCAGTAACTCGAACAAACTCGTACGGAAAGACTCCAAGTG
AGCTGGCTGATCCAGAAACAGAAGCTAGGAGAATCCTGGAAGGCAGTGCCTCATCTCAATGA
AA sequence
>Lus10022609 pacid=23178669 polypeptide=Lus10022609 locus=Lus10022609.g ID=Lus10022609.BGIv1.0 annot-version=v1.0
MISNGDIKITNNIDEPLEDGDTALHPACLYGYSSCVRVLLELGANIEDKDEDGAIPLHDASAGAMAGYTEIVQLLLNHANDAEAVKRMLETMDDEGDTPL
HHAARGEHAEVVNLLLASGASVTRTNSYGKTPSELADPETEARRILEGSASSQ
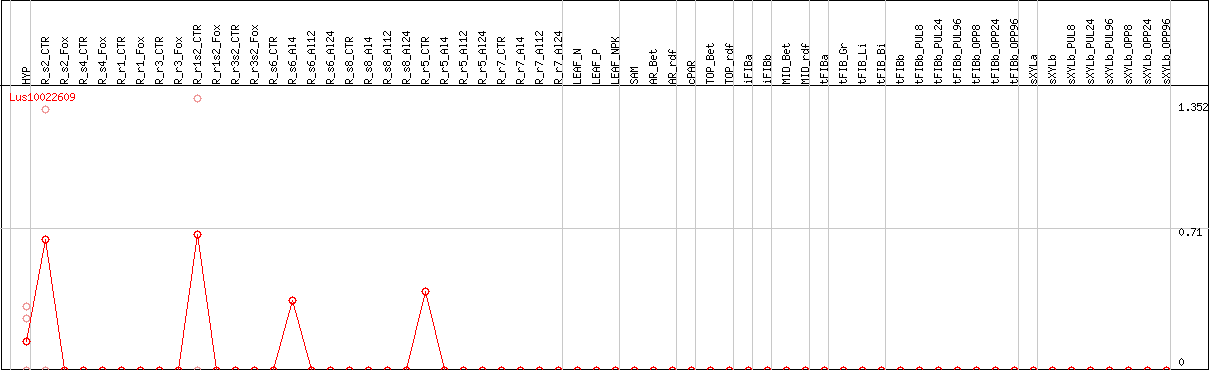
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022609 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.