External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G25160 355 / 8e-125
ER lumen protein retaining receptor family protein (.1)
AT1G19970 307 / 7e-106
ER lumen protein retaining receptor family protein (.1)
AT1G75760 307 / 9e-106
ER lumen protein retaining receptor family protein (.1)
AT4G38790 306 / 4e-105
ER lumen protein retaining receptor family protein (.1)
AT2G21190 301 / 2e-103
ER lumen protein retaining receptor family protein (.1)
AT3G25040 102 / 1e-26
ERD2B
endoplasmic reticulum retention defective 2B (.1)
AT1G29330 99 / 3e-25
ATERD2, AERD2, ERD2
ARABIDOPSIS THALIANA ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ARABIDOPSIS ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ER lumen protein retaining receptor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003146
498 / 0
AT3G25160 388 / 2e-137
ER lumen protein retaining receptor family protein (.1)
Lus10038198
381 / 9e-135
AT3G25160 362 / 4e-127
ER lumen protein retaining receptor family protein (.1)
Lus10025906
360 / 2e-120
AT3G25150 363 / 4e-118
Nuclear transport factor 2 (NTF2) family protein with RNA binding (RRM-RBD-RNP motifs) domain (.1), Nuclear transport factor 2 (NTF2) family protein with RNA binding (RRM-RBD-RNP motifs) domain (.2)
Lus10017213
302 / 1e-103
AT1G75760 481 / 5e-174
ER lumen protein retaining receptor family protein (.1)
Lus10026285
300 / 4e-103
AT4G38790 473 / 7e-171
ER lumen protein retaining receptor family protein (.1)
Lus10021097
298 / 4e-102
AT1G75760 475 / 8e-172
ER lumen protein retaining receptor family protein (.1)
Lus10007570
295 / 6e-101
AT4G38790 466 / 2e-168
ER lumen protein retaining receptor family protein (.1)
Lus10012176
260 / 8e-87
AT2G21190 397 / 2e-140
ER lumen protein retaining receptor family protein (.1)
Lus10042389
209 / 9e-68
AT2G21190 355 / 5e-125
ER lumen protein retaining receptor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G246700
357 / 3e-125
AT3G25160 400 / 5e-142
ER lumen protein retaining receptor family protein (.1)
Potri.009G128900
309 / 1e-106
AT2G21190 476 / 2e-172
ER lumen protein retaining receptor family protein (.1)
Potri.004G167400
309 / 2e-106
AT4G38790 472 / 1e-170
ER lumen protein retaining receptor family protein (.1)
Potri.002G022300
303 / 3e-104
AT1G75760 464 / 2e-167
ER lumen protein retaining receptor family protein (.1)
Potri.017G056000
114 / 7e-31
AT3G25040 383 / 3e-137
endoplasmic reticulum retention defective 2B (.1)
Potri.001G315800
112 / 3e-30
AT3G25040 355 / 3e-126
endoplasmic reticulum retention defective 2B (.1)
Potri.011G071200
79 / 1e-17
AT1G29330 341 / 1e-120
ARABIDOPSIS THALIANA ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ARABIDOPSIS ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ER lumen protein retaining receptor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0141
MtN3-like
PF00810
ER_lumen_recept
ER lumen protein retaining receptor
Representative CDS sequence
>Lus10022763 pacid=23159894 polypeptide=Lus10022763 locus=Lus10022763.g ID=Lus10022763.BGIv1.0 annot-version=v1.0
ATGGGTTCGAAGAGGAACGCTATGGGGGGCGGAGGGTCGTCGCCGCCGGCGGTGAAGGCGGTGGTTAAGCTCTGTGTTAGGAATCACAACCATTTCTTCA
TCGCCTCTGAGGCCATTCACGCCGCCGGGATCATCGTCCTTGCATACAAGCTCATCACCCAGAAAACCTGCTCAGGGCTTTCACTGAGGAGTCAGGAGCT
GACAGCATTGTTCCTAGCGGTGAGGCTGTGTTGCAGCGTGTTCATGGAGGCGGACATCCACACCATGCTCGATTTCGCCACCCTCGCTCTGACCCTGTGG
GTCATCTACATGATGAGGTTCAAGTTGAAGTCCTCCTACATCAAGGAGCTCGACAACTTGAACGTCATCTATCTGGTTGTCCCTTGTCTTGCCCTTGCCT
TTATAGTCAAGCCACATACCTTATACGGCTTGGTAAGCGAATTCCTCTGGGCGTTTTGCGCCTACCTCGAAGCTGTCTCTGTCCTGCCCCAGCTGCGAAT
GATGCAGAACGCCAAGATGATCGAGCCGTTCACGTCTCGCTACGTATTCGCGCTGGGGGTGGCTAGATTCCTGTCCTGTGCACATTGGATCATCAGGGTG
ATCGAGACTCGTGGGGCGTACCTGTACATAACGGGGAGCGGGTACTTCTGGTTCCCGGTGGCGTTCCTGGCGGAGATGGTCCAGACGTTCATCTTGGCTG
ATTTCTGCTACTATTACGTTAAGAGTTTCATGGCAGGGCAGCTTGTGATGAGGATGCCTGTTTAG
AA sequence
>Lus10022763 pacid=23159894 polypeptide=Lus10022763 locus=Lus10022763.g ID=Lus10022763.BGIv1.0 annot-version=v1.0
MGSKRNAMGGGGSSPPAVKAVVKLCVRNHNHFFIASEAIHAAGIIVLAYKLITQKTCSGLSLRSQELTALFLAVRLCCSVFMEADIHTMLDFATLALTLW
VIYMMRFKLKSSYIKELDNLNVIYLVVPCLALAFIVKPHTLYGLVSEFLWAFCAYLEAVSVLPQLRMMQNAKMIEPFTSRYVFALGVARFLSCAHWIIRV
IETRGAYLYITGSGYFWFPVAFLAEMVQTFILADFCYYYVKSFMAGQLVMRMPV
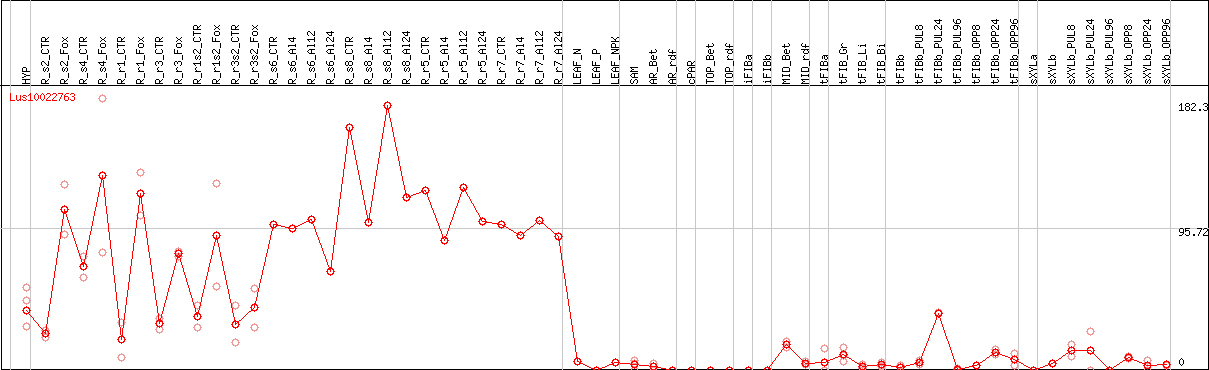
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022763 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.