External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G03905 373 / 1e-130
ABCI19
ATP-binding cassette I19, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G44110 360 / 2e-125
ABCI21, ATPOP1, ATNAP2, POP1
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN, ATP-binding cassette A21, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
AT5G02270 257 / 2e-84
ABCI20, ATNAP9
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 9, ATP-binding cassette I20, non-intrinsic ABC protein 9 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024914
471 / 9e-169
AT1G03905 439 / 2e-156
ATP-binding cassette I19, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10010838
255 / 7e-83
AT5G02270 514 / 0.0
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 9, ATP-binding cassette I20, non-intrinsic ABC protein 9 (.1.2)
Lus10023792
253 / 1e-82
AT5G02270 530 / 0.0
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 9, ATP-binding cassette I20, non-intrinsic ABC protein 9 (.1.2)
Lus10003971
252 / 4e-78
AT5G02270 530 / 0.0
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 9, ATP-binding cassette I20, non-intrinsic ABC protein 9 (.1.2)
Lus10030674
44 / 0.0001
AT3G28860 2271 / 0.0
P-GLYCOPROTEIN 19, MULTIDRUG RESISTANCE PROTEIN 11, Arabidopsis thaliana ATP-binding cassette B19, ATP-binding cassette B19, ATP binding cassette subfamily B19 (.1)
Lus10041690
44 / 0.0001
AT4G39850 1404 / 0.0
PEROXISOME DEFECTIVE 3, COMATOSE, Arabidopsis thaliana ATP-binding cassette D1, acetate non-utilizing 2, ATP-binding cassette D1, peroxisomal ABC transporter 1 (.1.2.3)
Lus10024056
44 / 0.0002
AT4G39850 1666 / 0.0
PEROXISOME DEFECTIVE 3, COMATOSE, Arabidopsis thaliana ATP-binding cassette D1, acetate non-utilizing 2, ATP-binding cassette D1, peroxisomal ABC transporter 1 (.1.2.3)
Lus10002094
42 / 0.0002
AT4G33460 304 / 9e-105
embryo defective 2751, ATP-binding cassette I10, ABC transporter family protein (.1)
Lus10000827
42 / 0.0004
AT4G33460 362 / 5e-127
embryo defective 2751, ATP-binding cassette I10, ABC transporter family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G036300
395 / 4e-139
AT1G03905 441 / 2e-157
ATP-binding cassette I19, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.005G226600
395 / 5e-139
AT1G03905 435 / 3e-155
ATP-binding cassette I19, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.006G086000
258 / 8e-85
AT5G02270 511 / 0.0
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 9, ATP-binding cassette I20, non-intrinsic ABC protein 9 (.1.2)
Potri.003G155800
248 / 6e-81
AT5G02270 503 / 0.0
ARABIDOPSIS THALIANA NON-INTRINSIC ABC PROTEIN 9, ATP-binding cassette I20, non-intrinsic ABC protein 9 (.1.2)
Potri.017G081100
41 / 0.0009
AT3G28860 2107 / 0.0
P-GLYCOPROTEIN 19, MULTIDRUG RESISTANCE PROTEIN 11, Arabidopsis thaliana ATP-binding cassette B19, ATP-binding cassette B19, ATP binding cassette subfamily B19 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00005
ABC_tran
ABC transporter
Representative CDS sequence
>Lus10022909 pacid=23159973 polypeptide=Lus10022909 locus=Lus10022909.g ID=Lus10022909.BGIv1.0 annot-version=v1.0
ATGGCGGAAAGTGCAGCTTCGTCGACTACACCGGCGGGCGGCGGAGGAAGAGACGGAGTCGGGAATGGTAGTATCAGCGTGTCGGCGATGCAATTCTCGT
ACGACGGAAAACAGGCGCCTCTTTTCTTCGACTTCAGTCTCGACATTGCTCCTGGATCTCGATGCTTGCTCACCGGCGCCAATGGATCAGGGAAGTCGAC
TTTACTGCGGATCGTGGCGGGGAAGCACATGGTGGGAGGGAAAGATGTCGTGAGAGTACTGAATAGCTCAGCTTTCCATGATACTCAGCTCGTTTGTAGC
GGTGACTTGGCCTATTTGGGAGGTTCGTGGAGTAAAACCGTAGGATCAGCGGGAGAGATTCCGCTGCAGGGTGATTTCTCTGCTGAACATATGATATTTG
GAGTTGAGGGAGTTGATCCAGCTAGAAGAGAGCAGCTAATTGAGCTACTTGATATCGATCTTCGATGGCGTATGCATAAGGTTTCTGATGGACAGCGGCG
TCGTGTTCAAATCTGTATGGGTCTTCTTTATCCGTATAAGGTTCTGTTATTGGATGAGGTGACAGTTGATTTGGACGTTGTTGCGAGGATGGACTTGCTC
GAGTTCTTTAGGGAAGAAACTGAGCAGAGGGGAGCGACATTAATCTACGCAACTCATATTTTCGACGGGCTAGAGGGATGGGCGACACACCTTGCTTACA
TTCAAGATGGTGAACTTAACAGGTATTCAAAGCTATCCGAGCTGAACGAGTTGAAAAGCTCCGTAAACCTACTCACTGTCGTCGAGAACTGGCTACGTGC
CGAAACCAAGCAGGAGAAAAAGAAGAAGAAGGAACCCATCAAAGCCCCTCCCGCCCAAACCCAGAAGCCCACCTCGGCTTCTCCGTTCATGTCGTCTAGT
AGACACATGGCCTATTACCGTTGA
AA sequence
>Lus10022909 pacid=23159973 polypeptide=Lus10022909 locus=Lus10022909.g ID=Lus10022909.BGIv1.0 annot-version=v1.0
MAESAASSTTPAGGGGRDGVGNGSISVSAMQFSYDGKQAPLFFDFSLDIAPGSRCLLTGANGSGKSTLLRIVAGKHMVGGKDVVRVLNSSAFHDTQLVCS
GDLAYLGGSWSKTVGSAGEIPLQGDFSAEHMIFGVEGVDPARREQLIELLDIDLRWRMHKVSDGQRRRVQICMGLLYPYKVLLLDEVTVDLDVVARMDLL
EFFREETEQRGATLIYATHIFDGLEGWATHLAYIQDGELNRYSKLSELNELKSSVNLLTVVENWLRAETKQEKKKKKEPIKAPPAQTQKPTSASPFMSSS
RHMAYYR
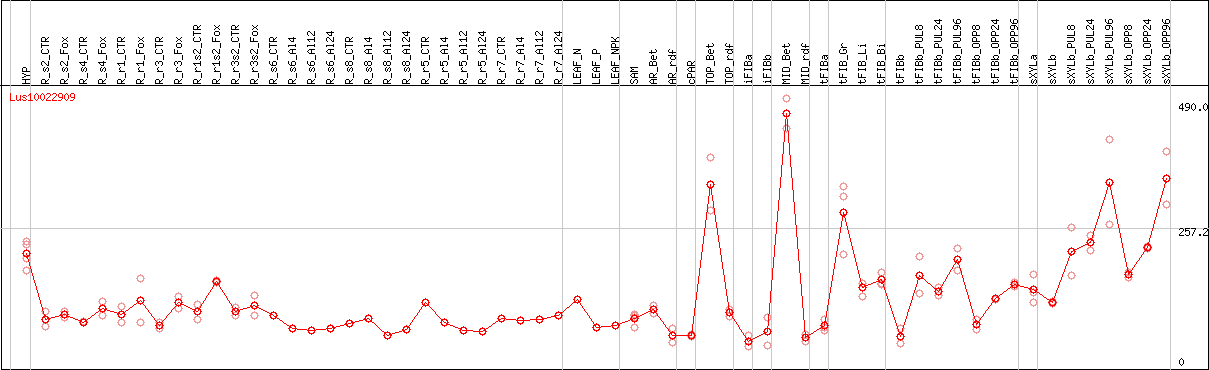
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10022909 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.