External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G09650 256 / 1e-80
CRM3, HCF152
HIGH CHLOROPHYLL FLUORESCENCE 152, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G69290 62 / 7e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G06920 59 / 6e-10
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G68980 57 / 2e-09
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G03100 54 / 3e-08
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G18940 51 / 2e-07
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G18475 51 / 2e-07
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63630 49 / 2e-07
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G62720 51 / 3e-07
AtNG1
novel gene 1, Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT2G17140 51 / 3e-07
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015059
314 / 5e-102
AT3G09650 969 / 0.0
HIGH CHLOROPHYLL FLUORESCENCE 152, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10037077
62 / 6e-11
AT1G69290 827 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10042362
57 / 4e-09
AT1G02420 613 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10043417
54 / 3e-08
AT3G06920 1358 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10028942
49 / 5e-08
AT5G01110 118 / 2e-32
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10026306
52 / 8e-08
AT1G02420 522 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10020775
51 / 2e-07
AT1G03100 961 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10031423
51 / 2e-07
AT5G66520 538 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10009201
51 / 2e-07
AT1G12700 274 / 3e-86
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G085300
266 / 2e-84
AT3G09650 961 / 0.0
HIGH CHLOROPHYLL FLUORESCENCE 152, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.008G095800
67 / 2e-12
AT1G69290 860 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G014100
57 / 2e-09
AT3G06920 1404 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G212400
56 / 7e-09
AT1G03100 1044 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G045000
55 / 1e-08
AT1G12700 502 / 4e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.008G108300
54 / 3e-08
AT3G16890 771 / 0.0
pentatricopeptide (PPR) domain protein 40 (.1)
Potri.017G091600
53 / 6e-08
AT4G37170 924 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G150100
52 / 2e-07
AT1G62930 442 / 8e-149
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G176100
51 / 2e-07
AT3G49730 783 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.009G105600
51 / 3e-07
AT5G39710 1022 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10023168 pacid=23170278 polypeptide=Lus10023168 locus=Lus10023168.g ID=Lus10023168.BGIv1.0 annot-version=v1.0
ATGCTGAAAGATTCGCGGGTCAAAGTTGATTTGATTGCCTGGAATATGTTAGTTGAAGGGTATTGTAAGTTGGGTTTGGTTCAGGAAGCGAAGAGAGTTG
TAGAGAAGATGAAGGAGAATGGGTTTTACCATGATGTGGCTACTTATGGAAGTCTAGCAAATGGAATAGCATTGGCTAAAAAACCAGGGGAAGCTCTTCT
GTTATGGAAGGAAGTAAAAGATAGATGCGAGATGAAATGGAAAGGAGAGAGCTCCGAGTCCGACCCTGATTCGCCTCCTGTCCCACCTTTGAGGCCCGAT
GACGAGCTGTTAGATACACTGGCTGATATCTGCGTCAGAGGTGCTTTCTTCCAGAAGGCTTTGGAGATCGTTGCCTGCATGGAAGAATATAGAATTCCAC
CAAACAAGTCAAAGTACAAGAAGATTTATGTGGAGATGCATTCCAGGATGTTTACCAGTAAACATGCGTCGCAAGCGAGGCAAGATCGGAGGAAAGAGAG
GAAGAGAGCAGCCGAGGCTTTCAAGTTTTGGTTAGGTTTGCCTAATGAATATTATGGAAGCGAATGGCGACTCGATCCCTTGGACACCGAGATGGATGTC
AGCACAAAGGGATTAGAGAGGCTGGCAGGAACAGTTGTTTTGCAAGCAAATGCAGCAATAGTCCTTGTATGCAAACACAAAGAACTTGATCGAACTTACG
AGTAG
AA sequence
>Lus10023168 pacid=23170278 polypeptide=Lus10023168 locus=Lus10023168.g ID=Lus10023168.BGIv1.0 annot-version=v1.0
MLKDSRVKVDLIAWNMLVEGYCKLGLVQEAKRVVEKMKENGFYHDVATYGSLANGIALAKKPGEALLLWKEVKDRCEMKWKGESSESDPDSPPVPPLRPD
DELLDTLADICVRGAFFQKALEIVACMEEYRIPPNKSKYKKIYVEMHSRMFTSKHASQARQDRRKERKRAAEAFKFWLGLPNEYYGSEWRLDPLDTEMDV
STKGLERLAGTVVLQANAAIVLVCKHKELDRTYE
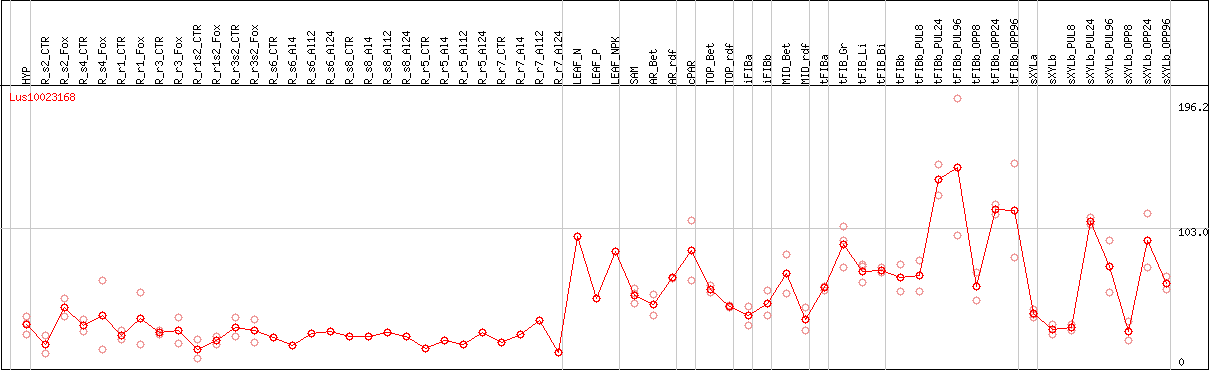
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10023168 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.