External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G02450 322 / 7e-109
NAC
LOV1, ANAC034, ANAC035
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
AT5G39820 190 / 6e-58
NAC
ANAC094
NAC domain containing protein 94 (.1)
AT1G26870 190 / 5e-57
NAC
FEZ, ANAC009
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT1G77450 184 / 2e-56
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT2G17040 184 / 4e-56
NAC
ANAC036
NAC domain containing protein 36 (.1)
AT1G01720 181 / 8e-55
NAC
ATAF1, ANAC002
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT4G27410 180 / 1e-54
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT3G15500 180 / 2e-54
NAC
ATNAC3, ANAC055
NAC domain containing protein 55, NAC domain containing protein 3 (.1)
AT1G52890 180 / 3e-54
NAC
ANAC019
NAC domain containing protein 19 (.1)
AT3G04070 179 / 1e-53
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008897
433 / 4e-153
AT2G02450 315 / 5e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10008419
339 / 2e-114
AT2G02450 327 / 7e-109
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10003367
321 / 2e-106
AT2G02450 333 / 3e-110
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10008420
307 / 1e-102
AT2G02450 320 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10030478
194 / 4e-58
AT5G39820 297 / 4e-98
NAC domain containing protein 94 (.1)
Lus10003269
187 / 3e-56
AT3G04070 324 / 3e-109
NAC domain containing protein 47 (.1.2)
Lus10015367
186 / 3e-56
AT1G26870 298 / 8e-99
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10022636
185 / 5e-56
AT2G17040 287 / 6e-97
NAC domain containing protein 36 (.1)
Lus10037178
189 / 6e-56
AT5G39820 306 / 3e-101
NAC domain containing protein 94 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G046700
322 / 1e-108
AT2G02450 318 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Potri.004G230800
315 / 8e-106
AT2G02450 311 / 2e-103
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Potri.019G031400
186 / 7e-56
AT3G04070 332 / 1e-112
NAC domain containing protein 47 (.1.2)
Potri.011G123300
184 / 1e-55
AT4G27410 354 / 2e-122
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Potri.001G404100
184 / 1e-55
AT4G27410 370 / 8e-129
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Potri.009G141600
181 / 4e-55
AT2G17040 314 / 6e-108
NAC domain containing protein 36 (.1)
Potri.017G082000
183 / 2e-54
AT1G26870 299 / 3e-98
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.011G046700
181 / 4e-54
AT1G61110 331 / 8e-113
NAC domain containing protein 25 (.1)
Potri.002G081000
179 / 4e-54
AT1G01720 416 / 5e-148
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.007G066300
179 / 4e-54
AT3G12910 269 / 2e-89
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10023208 pacid=23180999 polypeptide=Lus10023208 locus=Lus10023208.g ID=Lus10023208.BGIv1.0 annot-version=v1.0
ATGAAGGAAAACAATGGCGATCGTGAAGTGGTCGACGAAGAAGAAGAAGATGAGCATGATCATGACATGGTGATGCCAGGGTTCAGGTTCCATCCCACAG
AGGAAGAGCTTATCCAATTCTACCTCCGCCGTAAGGTTGAGGGCAAACCCTTCAACGTCGAACTCATCACTTTCCTCGACCTTTATCGCTACGACCCTTG
GCACCTCCCTGCTATGGCGGCGATCGGGGAGAAAGAGTGGTTCTTCTATGTGCCTAGGGACAGGAAGTACAGGAACGGTGATAGGCCGAACCGGGTGACC
ACTTCTGGTTACTGGAAGGCTACTGGGGCGGACCGGATGATTCGAGGTGAGAACTCGAGGTCTATCGGCCTGAAGAAAACCCTAGTGTTTTACTCAGGGA
AGGCTCCCAAGGGGGTTCGGTCTAGCTGGATTATGAATGAGTATCGTCTTCCTCACCATGATACCCAACGATATCAGAAGACGGAGATATCGCTATGTCG
AGTATACAAGAGACCAGGAGTCGAAGATCATCGCGGTGTACCATCGACATCGACTCTTCCGCGCTCTATTCCTTCGTCATTTAGGGCACGGCATCATCAG
CAACATCCAGGTGCATCTTCTGCTGTGGGACTAATCTCCAACCAGCTCCCCACACTAACTCACCATCCACTACAGCTTCCTCCACAACAACAACAAGTAC
TGGATCATGCTCAAGTAGAAGAAGATCAGCTCATGCTCCTAAACGGTACTAGTACTAACAACAATAATAATTTGGATGATTTGCATCGGCTAGTGAACTA
CCAACAGAATCACGATCACCTTAGTTACCATCCACACCATCAGCTGCATATGTGGGAGTGGGACCGCTTTGGGGGGGGGGGGGCCCCCCCCCCGCCTCCG
CGGCCACCTCAACATGATCATCAGAATCATGATCTTCAAGGTGGCCATAGATTGGCAGGTGATCATCATCACTACGAGACCAACAACCAAACTAATCTTT
TCAAGTAA
AA sequence
>Lus10023208 pacid=23180999 polypeptide=Lus10023208 locus=Lus10023208.g ID=Lus10023208.BGIv1.0 annot-version=v1.0
MKENNGDREVVDEEEEDEHDHDMVMPGFRFHPTEEELIQFYLRRKVEGKPFNVELITFLDLYRYDPWHLPAMAAIGEKEWFFYVPRDRKYRNGDRPNRVT
TSGYWKATGADRMIRGENSRSIGLKKTLVFYSGKAPKGVRSSWIMNEYRLPHHDTQRYQKTEISLCRVYKRPGVEDHRGVPSTSTLPRSIPSSFRARHHQ
QHPGASSAVGLISNQLPTLTHHPLQLPPQQQQVLDHAQVEEDQLMLLNGTSTNNNNNLDDLHRLVNYQQNHDHLSYHPHHQLHMWEWDRFGGGGAPPPPP
RPPQHDHQNHDLQGGHRLAGDHHHYETNNQTNLFK
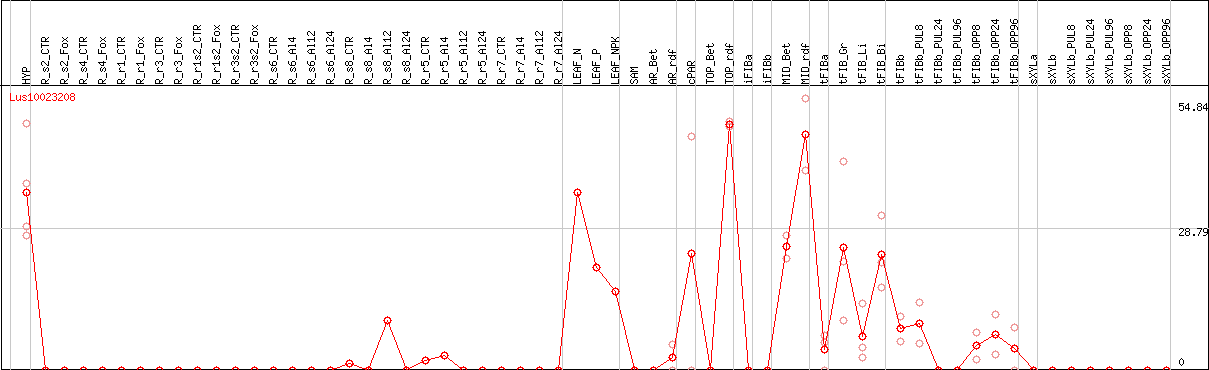
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10023208 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.