Lus10023270 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10023270 pacid=23180945 polypeptide=Lus10023270 locus=Lus10023270.g ID=Lus10023270.BGIv1.0 annot-version=v1.0
ATGGAGGAACCACCGCCTGTGGTATCATCCGGTGAAGACCCAGTTCAGATGCCGCCTGTGGAGGTTACTGTAAGTCCTAATGAGACAACCAAGAAGGTTG
TAAAAAGACCCACCAAGCCTATCATTTCTCGGACCTCAACACTGTTGGCACCTAAAGGTTCAGCTAGAAAGAGGGCTGAATCCAAAGAAAGTCCGGATTC
AAGCACAAATGCAGGGAAGCAAAGCACTATTGGTAATACTTCCTCCTCCTGTACTTCCACTGCAGTTCCAGCCATTAGACGGAACAGTACTGGTGGAGTG
TCTGCTGAGAAACGTGGGCAGAATGGGGCTTCTGTTGTCAGTGCTGGGAAAAATAGTAGTGCTGTGGTGGACCCGGTAAGGAAGTCCCTTCCAGAGCTGA
GAAGGAGTTCCTTACCTTCTGTGGCTGCAAAACCCACAAGAGGATCGAGTGTTTCAGGGATTAGGAAATCAGTTCACATGTCACCTTCGGAAAAGAGTCT
GAGGACTTCAACTGCTTCTGATATCAGTATCCCGGGGACAGCCCGTAAGCCTTCAGTTAAGCCAACATTGTCGGCCTCTTCTTCTTCATCTTCTGTGAAG
AGAGTTCCATCTACATCACTTGAGAGCAGTGCCAGTAGTGTGACTAGGAAGACAGTATCAAAAGTTTCATCGTCTTCGGCTCGTTCCCCGTCAGTTTCTA
GTGGATTACGAAGTGGGTCATTGTCAACATCCTTGGATACGTGTTCTAGCTTGTCTGCAAGGAGGAGAGTGGGCACGCCTGAGAGTCGGGATTCTCGGTT
CATTGCACTGCCACAAAATGAGCTTAAAGGTGGGGGGGGTGGGGGGGGTTTTCTGGGTCACAGAGTTCGCACCCTCAATGCTAATGGATTAAACTTGTCA
GCCACTTTAGAGGTATTCTATTTCAGATACATCATAAATTATAGCTTATACTGTCTGCTGCTGAGAACTACTGCCAGTAATTCCTCTAACTACTATTTTG
TTGATTTGAGTTTCAATGAGTTTAAGGGTCCTGGATTTGAGCCTCTTGAGAATTGTACTGCCTTACAGCAACTGTATCTTGCTGGAAATCAGATCACATC
ACTTATGAGCCTTCCGCAGCTACCTAACTTGGAGTTCCTATCTGTTGCTCAAAACAAGCTGAAGTCACTTGCAATGTCAAGTCAACCTCGATTACAGGTT
TTAGCTGCAAGCAAAAATAAAATATCAACTCTTAAGGGTTTCCCGTATCTTCCTGCCCTTGAGCATTTACGAGTTGAGGAAAATCCGATACTTAAGGTGC
CCTATTTAGAAGCTGCTTCTATATTGCTTGTTGGTCCTACATTGAAAAAGTTCAATGATAGAGGTTTCACACGTGAGGCGATAACAATAGCCAAGCGGTA
TCCAGCATGCACGGCTTTGTGTATTAGAGATGGCTGGGAATTTCGTCGTCCCGAGAATGTCTCAGAATCAATTTTTCACTTTCTTTTGGAGAAATGGAAG
GACCACTTGCCTGCTGGTTATCTTTTAAAAGATGCCCGTGTTGATCAACCATTTGAAGATGACATCTGCAGATGCCACTTCAACTTTCACCAAGATGAAG
TTCCAATCGATGAGCCACCATTAGTCTTGAAATATCAATGGTTTGTGGGGGAGAGACCACTTTCAGACTTAATTCCAATTGCTGATGCCAACAAAGAGGT
GTACTGGCCAACACGTGATGACATCCGTAAATTTCTGAAAGTTGAGTGTATTCCTGTAATTGGAGAGACTGAGTATCCAGCTATATTTGCCATTTCACCC
CCAGTTTCACGAGGAAATGGCATCCCGAAAGTAGTTAACCTTGAAGTGTGTGGGGAGTTGGTGGAAGGAAGTGTTATCAAGGGCCAGACAAAGGTTGCAT
GGTGTGGTGGGACTCCTGGTAAAGGCGTTTCCAGTTGGCTTAGGAAAAAGTGGAACAGCAGCCCTGTGGTTATTGCTGGAGCAGAAGATGAAGAGTACCA
GTTGACTCTAGATGATGTTGATTCAAGTCTTGTCTTTATGTATACACCAGTCACAGAGGAAGGTGCAAAAGGGGAACCTCAGTATAAGTACACGGATTTC
GTTAAAGCAGCTCCTCCTTCAGTCAACAATGTCCAAATTGTTGGAGATGTGGTTGAAGGAAATATAATGAAAGGTGTTGGTGATTACTTTGGAGGAAAAG
AGGGTCCTAGCAAGTTTGAATGGCTACGTGAGAACAAGGACACTGGGGAGTTTTTAGTAGTATCATCTGGTACATCGGAATATACTCTGACAAAGGGAGA
TGTTGGTCACCGTGTCGCTTTTGTATATATCCCCACTAATTTTGAAGGACAAGAAGGAGAATCTATGTCAGTCATGACTCATGCAGTTAAGCGAGCTCCT
CCGAAGGTATCAAATGTTAAGATAATTGGCGAATTAAGAGAAAACAGTAAGATTACTGTTACTGGCATTGTCACCGGAGGAACTGAAGGCTCTAGCAGGG
TACAATGGTTCAGAATGACTTCTTCAACATTTGATGATGAGAATGGTCTTGAAGCACTGAGCACGTCCAAGATTGCCAAGGCATTTCGGATTCCTCTTGG
AGCGGTTGGCTATTACATTGTTGCAAAATATACTCCGATGACTTCAGATGGGGAATCTGGTCAACCAGCATATGCAATGTCAGAAGCTATCATCGAAACC
CTTCCTCCGAGCCTAAACTTCTTGTCTATAACTGGTGACTACACTGAAGGTGGAATATTGACCGCATCATATGGCTACGTGGGGGGTCATGAAGGGAAAA
GTATATATAATTGGTATCTTCATGAGGTAGAAACAGACTCTGGTATCTTGATACCTGAGGGTTGTGGAGTCCTTCAATATCAGGTTAACAAAGATGCAAT
TGGGAAATTTGTATCCTTCGAATGTGTCCCGGTGCGTGATGATGATATTGTAGGTGAACCTAGGAGTTGCATGGGCCAGGAACGTCTTCAACCAGGAAGC
CCACGATTGCTCGCTCTGCAAATTGTTGGAAGTGCTGTTGAAGGAACTCAACTAAGTGTCAACAAAAAGTACTGGGGTGGTGAAGAGGGAGAATCAGTTT
TTCGTTGGTTACGGATTGCATCAGATGGTACCCAATTTAAAATTCCTGGTGCCAGTGCTTCTTCATACACCATGGTGTTTGATGATATTGGTTACTATGT
ATCTGTTTCATGCGAACCTGTCAGAAGCGATTGGGCACGTGGTCCTGTTGTCCTCTCTGAACAAGTTGGACCAATCGTACCTGGCCCTCCAACTTGCCAA
TCCTTAGAGTTCCTTGGATCTATGATTGAGGGGCAGCGAGTCAGCTTCATAGCTACTTACACAGGAGGGGACAGGGGAAATTGCTGCCATGACTGGTTTA
GGGTGAGAAGTAATGGTGTCAGGGAGAAACTTAGTGATGATGACTTTCTGGATTTGACGTTAGAAGATGTTGGAAAACATATCGAATTAATCTACTTTCC
AATTCGCAAAGATGGTGCACGGGGAAGTCCTAAAACCATTAAGTCTAATATCGTTGTTCCCGCCTACACTCCCTCACTTGAAGATGTGGGATCTTTTTTG
GCTTTATACTGGGTGCCCGTACGTGCTGATGGAAAACGTGGAAAGCCATTGGTTGCTATTTCAAGCTCTCCTGTTAACCCAGGGCTTCCGAGCGTTTCCG
AGGTTCATGTCAAGGAGTTATCATCAGGTGCCTATTGTGGGGAAGGGGTATATTTTGGTGGGTATGAGGGGTCCAGCCAGTTTAGCTGGTATAGAGAAAC
CAACGAGGGAAAAATGATCCTCATTGATGGAGCTAACTCTGTAACTTATGAAGTCACGGACTCTGATTACACTTGCCGTTTGTTATTTGGATACACCCCA
GTTCGCTCAGATTCGGTTGTGGGAGAACTTGTGTTGTCAGCCCGAACTGACATTATCCTCCCTGAACTTCCAAGAGTCGAAATGCTCGCTCTCACTGGAA
AGGCAATAGAAGGGGACATACTTACTGCGGTTGAGGTCATTCCAGAGAATTTGACTCAACAATCCGTTTGGAGCAAATACAAAAAGAATGTCAAATATCA
ATGGTTCCGTTCCTCTGAGTCAGGGGAGAGCAACTGTTTTGAGCCTTTAACTACTCAGGTGTCATGCTCTTACAAGGTGCGATTGGAGGACATTGGTAGA
TGGTTGAGATGCGAATGTATCATTTATGATGTCTTTGGAAGGTTAAGCGAACCAGCATATGCTGTGACTGAAGCTGTATTACCAGGAATTCCTAGAATGC
ATAAGTTGGAGATCGAGGGAAGGGGTTACCACACTAACCTATATGCTGTTCGGGGTGTATATACTGGTGGAAAAGAAGGAAAAAGTAAAATCCAGTGGCT
AAGATCAATGGTTGGGAGTCCTGATCTCATCTCCATCCAAGGAGAGGTAGGAAGGATGTACGAAGCTAATGTTGATGATGTAGGGTATAGGCTAGTGGCT
ATTTATACCCCAATTAGGGAAGATGGCGTAGAGGGGCAACCTGTATCTGCATCAACCGAGCCGATTGCTGTTGAGCCTGATGTTTTAAGCGAAGTAAAAC
AGAAACTTGACGTCGGGTCTGTGAAGTTTGAGGTATTATGCGACAAGGACCGTTCATCCAAGAAGTTTGCGGGGGAAGGAAGTCTGGAGAGAAGGGTTTT
GGAGATCAACAGAAAGAGAGTGAAGGTTATGAAGCCTGGCTCCAAGACTTCATTTCCAGCTACCGAAATTCGTGGAACTTATACTCCTCCTTTCCATGTG
GAGCTGTTCAGAAACGACCAACACCGGCTAAGAATCGTAGTGGACAGCGAGAACGTAGCAGACTTGATGGTGCACTCGCGGCACTTGAGGGACATAGTTG
TTCTTGTGATCAGAGGGCTGGCTCAGAGATTCAACAGTACATCTCTCAATTCTCTCCTCAAGATAGACACGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10023270 pacid=23180945 polypeptide=Lus10023270 locus=Lus10023270.g ID=Lus10023270.BGIv1.0 annot-version=v1.0
MEEPPPVVSSGEDPVQMPPVEVTVSPNETTKKVVKRPTKPIISRTSTLLAPKGSARKRAESKESPDSSTNAGKQSTIGNTSSSCTSTAVPAIRRNSTGGV
SAEKRGQNGASVVSAGKNSSAVVDPVRKSLPELRRSSLPSVAAKPTRGSSVSGIRKSVHMSPSEKSLRTSTASDISIPGTARKPSVKPTLSASSSSSSVK
RVPSTSLESSASSVTRKTVSKVSSSSARSPSVSSGLRSGSLSTSLDTCSSLSARRRVGTPESRDSRFIALPQNELKGGGGGGGFLGHRVRTLNANGLNLS
ATLEVFYFRYIINYSLYCLLLRTTASNSSNYYFVDLSFNEFKGPGFEPLENCTALQQLYLAGNQITSLMSLPQLPNLEFLSVAQNKLKSLAMSSQPRLQV
LAASKNKISTLKGFPYLPALEHLRVEENPILKVPYLEAASILLVGPTLKKFNDRGFTREAITIAKRYPACTALCIRDGWEFRRPENVSESIFHFLLEKWK
DHLPAGYLLKDARVDQPFEDDICRCHFNFHQDEVPIDEPPLVLKYQWFVGERPLSDLIPIADANKEVYWPTRDDIRKFLKVECIPVIGETEYPAIFAISP
PVSRGNGIPKVVNLEVCGELVEGSVIKGQTKVAWCGGTPGKGVSSWLRKKWNSSPVVIAGAEDEEYQLTLDDVDSSLVFMYTPVTEEGAKGEPQYKYTDF
VKAAPPSVNNVQIVGDVVEGNIMKGVGDYFGGKEGPSKFEWLRENKDTGEFLVVSSGTSEYTLTKGDVGHRVAFVYIPTNFEGQEGESMSVMTHAVKRAP
PKVSNVKIIGELRENSKITVTGIVTGGTEGSSRVQWFRMTSSTFDDENGLEALSTSKIAKAFRIPLGAVGYYIVAKYTPMTSDGESGQPAYAMSEAIIET
LPPSLNFLSITGDYTEGGILTASYGYVGGHEGKSIYNWYLHEVETDSGILIPEGCGVLQYQVNKDAIGKFVSFECVPVRDDDIVGEPRSCMGQERLQPGS
PRLLALQIVGSAVEGTQLSVNKKYWGGEEGESVFRWLRIASDGTQFKIPGASASSYTMVFDDIGYYVSVSCEPVRSDWARGPVVLSEQVGPIVPGPPTCQ
SLEFLGSMIEGQRVSFIATYTGGDRGNCCHDWFRVRSNGVREKLSDDDFLDLTLEDVGKHIELIYFPIRKDGARGSPKTIKSNIVVPAYTPSLEDVGSFL
ALYWVPVRADGKRGKPLVAISSSPVNPGLPSVSEVHVKELSSGAYCGEGVYFGGYEGSSQFSWYRETNEGKMILIDGANSVTYEVTDSDYTCRLLFGYTP
VRSDSVVGELVLSARTDIILPELPRVEMLALTGKAIEGDILTAVEVIPENLTQQSVWSKYKKNVKYQWFRSSESGESNCFEPLTTQVSCSYKVRLEDIGR
WLRCECIIYDVFGRLSEPAYAVTEAVLPGIPRMHKLEIEGRGYHTNLYAVRGVYTGGKEGKSKIQWLRSMVGSPDLISIQGEVGRMYEANVDDVGYRLVA
IYTPIREDGVEGQPVSASTEPIAVEPDVLSEVKQKLDVGSVKFEVLCDKDRSSKKFAGEGSLERRVLEINRKRVKVMKPGSKTSFPATEIRGTYTPPFHV
ELFRNDQHRLRIVVDSENVADLMVHSRHLRDIVVLVIRGLAQRFNSTSLNSLLKIDT
|
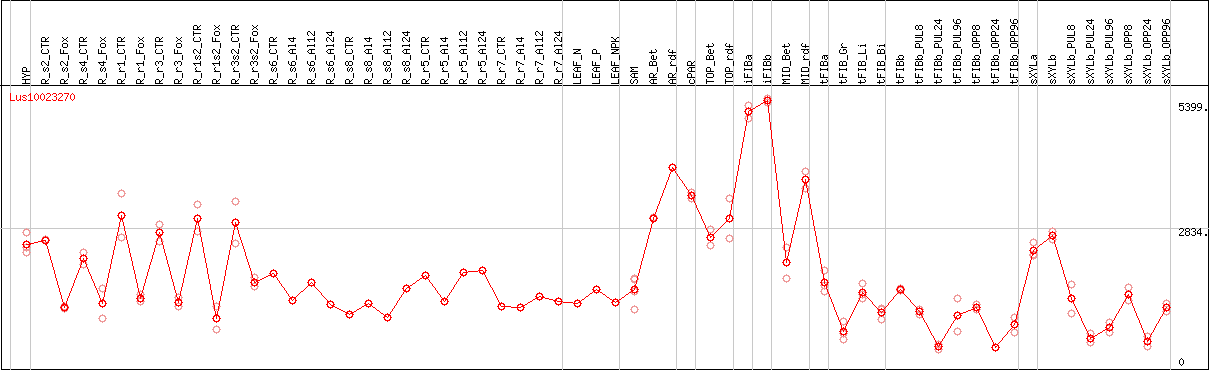
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10023270 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
