External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G14080 55 / 1e-08
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G18350 52 / 8e-08
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT3G44630 51 / 2e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2), Disease resistance protein (TIR-NBS-LRR class) family (.3)
AT5G49140 51 / 2e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT3G44480 50 / 3e-07
COG1, RPP10, RPP1
recognition of peronospora parasitica 1, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G69550 48 / 2e-06
disease resistance protein (TIR-NBS-LRR class) (.1)
AT4G19050 47 / 3e-06
NB-ARC domain-containing disease resistance protein (.1)
AT3G15410 47 / 3e-06
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
AT4G16890 46 / 9e-06
BAL, SNC1
SUPPRESSOR OF NPR1-1, CONSTITUTIVE 1, BALL, disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT2G25790 46 / 1e-05
Leucine-rich receptor-like protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038482
312 / 3e-99
AT1G69550 386 / 3e-113
disease resistance protein (TIR-NBS-LRR class) (.1)
Lus10023272
179 / 1e-51
AT5G36930 423 / 4e-126
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10000945
147 / 7e-41
AT1G69550 112 / 3e-25
disease resistance protein (TIR-NBS-LRR class) (.1)
Lus10038533
143 / 3e-39
AT1G27170 386 / 7e-114
transmembrane receptors;ATP binding (.1.2)
Lus10020534
137 / 6e-37
AT5G36930 414 / 1e-124
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020533
133 / 1e-35
AT1G27170 405 / 1e-122
transmembrane receptors;ATP binding (.1.2)
Lus10004911
133 / 1e-35
AT1G27170 387 / 5e-115
transmembrane receptors;ATP binding (.1.2)
Lus10023487
132 / 3e-35
AT5G36930 284 / 4e-78
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10004115
130 / 9e-35
AT5G27970 689 / 0.0
ARM repeat superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G068200
70 / 9e-14
AT5G17680 629 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.T005501
61 / 8e-11
AT5G36930 612 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.006G282300
52 / 6e-08
AT5G36930 413 / 3e-125
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G003285
51 / 2e-07
AT5G36930 640 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.011G009400
51 / 2e-07
AT5G36930 504 / 4e-162
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.017G104301
50 / 5e-07
AT5G17680 566 / 1e-178
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.002G258200
50 / 5e-07
AT4G08850 674 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.002G258000
49 / 8e-07
AT4G08850 695 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.007G016700
49 / 1e-06
AT5G46330 286 / 5e-84
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.013G097832
49 / 1e-06
AT4G12010 148 / 8e-38
Disease resistance protein (TIR-NBS-LRR class) family (.1)
PFAM info
Representative CDS sequence
>Lus10023330 pacid=23180973 polypeptide=Lus10023330 locus=Lus10023330.g ID=Lus10023330.BGIv1.0 annot-version=v1.0
ATGAGAGGTGAAGATTTTGAGCTGACAGACACTGAATTCAAGAAATTATCTGGCTTAGAATATCTAGATGTGAGGAATGGAAGAGTAACTGGAGATTTTA
AAGACATTCTTCCACATGTTTCACCCAAGCTTAAAGTTGTGGATGTATGGTCTTGTAAGGATTTAGAGACGGCTCCCAACTTGTCTCAGTGTCAAAGTGT
AGAGCGTTTAAAACTCCAGGACTGTCAGAAGATGAGAGGGGAACTGCATATTGGTAATTTTAAGAAACTAAAAGTGCTGAAGCTCGGGTATACTAAAATA
ACGAAGTTAACAGGCGGCATGGGAGTTCTTAAGAATCTCCAGGAGATTGATGCAGGGAACTCCTATTTGACAGAAGTGCCTTCTGGTATTGCCGAATTAT
CCTCTCTTAAAACCTTGGATCTAAGGTCGCATAAACTGGGATTGAAGGAATTACCAACACTCCCTAAAAGTTTAAAGCGCCTATATCTCTCATCGCCATG
GGTGCCTAACCTTTTGGCGCTCAAGGACTTGAAAGAGCTAGGGTTTTCAAATTGTGAAGCTCCTGAAATTCCAGGGGATGTATGGATGCTATCCAAGTTG
AAGTCTTTGAGTCTGTCGGACTGCACCTGTGAAAGTCTCTAG
AA sequence
>Lus10023330 pacid=23180973 polypeptide=Lus10023330 locus=Lus10023330.g ID=Lus10023330.BGIv1.0 annot-version=v1.0
MRGEDFELTDTEFKKLSGLEYLDVRNGRVTGDFKDILPHVSPKLKVVDVWSCKDLETAPNLSQCQSVERLKLQDCQKMRGELHIGNFKKLKVLKLGYTKI
TKLTGGMGVLKNLQEIDAGNSYLTEVPSGIAELSSLKTLDLRSHKLGLKELPTLPKSLKRLYLSSPWVPNLLALKDLKELGFSNCEAPEIPGDVWMLSKL
KSLSLSDCTCESL
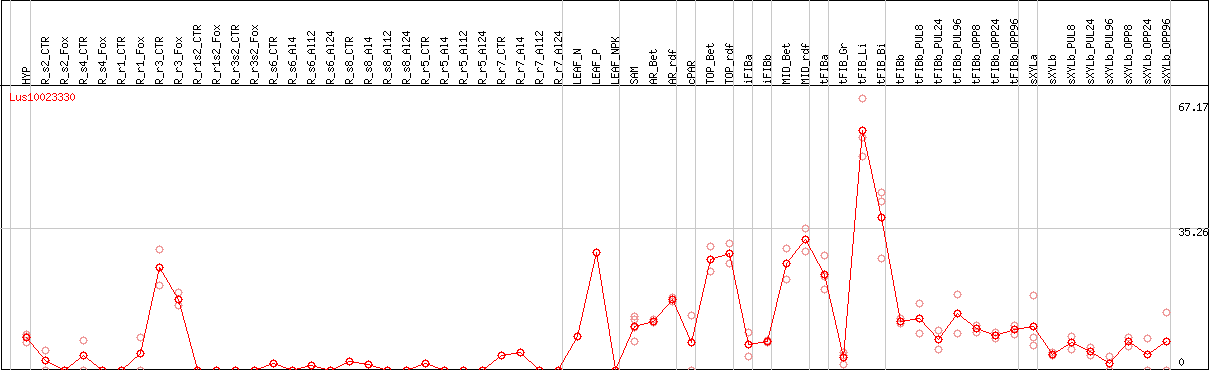
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10023330 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.