Lus10023360 [FLAX]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Lus10023360 pacid=23181051 polypeptide=Lus10023360 locus=Lus10023360.g ID=Lus10023360.BGIv1.0 annot-version=v1.0
ATGTCTGAGGTTTCTGCTTCGGAGAAGGCGCAGCTAAACCCCTGCTGTGAATCTTGGAGTGGGAAGTGCTCGAAGTTGCTGGAAGGGAGGAAGGCTCTAA
GGAAGGCGGTGCAGCTCCTCAAAGAAGAGCTCGACAAGATTCAACAAGACAATGCCAATCTTAAAAAAGGTTCTTGCAGCAACTTTACTTTTCCCGTATC
TTACTGTGTAACTGAGTTTTTTCCAGAATGTGAGGAGGCACGGAACATTGCTGCCAAAGAGAGTGAAAGGAAAGAGAATGAGCTGTCAGAAAAAATGTCT
CTTGAGAATGAGGTTTCTGCTTTGAAATCTGAGATCTCGTCACTGCAACAGAGAGGAAGTTTAGATGCGAAAGGGAGGGATAGGGAAATAGGGCTACTGC
GCGAACAAGTTTCTGAGACCAACAAACTTAAAGAGCTTCTTGAGAAAGAGAAGAGAAGGGCAGATGCTGAGAAGAAGAACGCTGAGACAGAGAAAAGAAG
AGCTGCTGAGGCTCAAAAGCGTATGAAAGCAGAGAAGGTTAAGGCTGTTGAAGAAAATAAGCATGCTGATGTTGAAACAAAGAAGGCTGAGGGTTACAGG
GTGCAGTTAGAGGTACTGAAAAAAGATGTTGAGGATATGAATTCATTACTGATTTCCGAGAGGGAGGAGAAAGCAAGATACTTGGCTATTAAGACCTCGT
TGGAGAATGAGGTATCTGTTCTTCAATCTGAGATATCCTGTTTGAAGAAAGAATGTGCTGCTGGCCAGCAGAAGGAGAAGGAAACTCAGAATGCCTCTGA
AATTTTTGAAAATGCTAATGTAGAGAATGCTGGTGCAGATAGGAAGCATGGTGGCTATTGTGAAAATACAGCTGAAACTTGTCTGCTCCAGTTACAGGCA
TTGATGAAAGAAGTAGAAGAAACCAAATCAATGTTGGTTTTCGAAAAGGAGGAGCATAGGAAAGAAGTTGCCCGAAGACTCTTGTTGGAAGAAGAAATTG
TTGGGTTAAAGTCTCAAATCTGCTGGTTACAACAAAGAGGAGATGAAGATATTCATGGGGAAATAGAGCTTCTCAAAGGTTCCCTGGGCGGGAAAGAAGA
GGAAATTCGGAAACTAAAGGAACTTTTTGATAAAGTTATATCGCAGTTGAATTCTGAGAAGAAGAAAGTTGAAGTGCAGAAGAAGATTGCTGATGAAGCA
TGTGAACGGGTGAAAGCTGAAAAGTCTAAGCTGGTCACTGAAAGTAAGAATGCTAGTTATGAAGCAAAGAAGGCTGAAAAATTTCAGGTGCAGTTAGATT
ATTTGAAGAAAGACTTCCTTAAAGTGCAATCAGAATTGGAGTCTGAGAAAGAGAAGTTCTCAATGATGAAGGAAAAGCGGGAAGTGGAACGACAAAATGT
GATAAAGGCGAGACAGTCGGTGGAAGCTGAAATTGCCAAGGCAGACCACCAAAGAAAGCTTGCTCAAGCGAATGGTAAGGTGTCTGCGAAAGAAAGAGCT
CGTGCTGACGAACTTTCTGGCCAGCTAGAACTATATAAACATAAGAATGAGGAGCTGCAGAAACAGATACAACAGTTTCTTCACTGCCAAAATAGTGCAG
AACTACCTGGCACTTTGCCTGTGGGAAAAGATAAAACCAAACTCACAAAGATGAAGCTTCTCAAGAAGCAATTGAAGCTTGGAAAGATGCAGTTGAAGCA
TTCCAAGCAAGTGGCTGAGTTTGAAAAGCACCGCAATGTGATCCTACGGGAAGAGCTAGGCAACCTGAGGCATGATTTTGTTCGGATTTCTGGTCGCTTG
GATCTGCTGGCTAAATGTTTCTCTCCAAATGTTGAAGGTATAGCAGACATGAAAAAGCACAGCGTTAAACAGTTTCCTAACCTTAATTGTTACACATTCC
AGGATGGAGCTCTTGGAGAGATCCAGTATCTGATGAAGGGAAGAAAGCTCTATGATATAGAACCACATCAAATGTATACTAAAAAAGAAAGCGAAGATTT
CAACCCTGACTGCGGAGCTACAGCTACGACTAATCTTATGAAGCATGCTAATATGCTCTCACCTACATGTGACCTCAATTTATCTACCTCAGGTATTGAT
TCTCAGTTGGAGTCTTTTCAGGGAGGCTCTAACACAAAGTTGTTACTACATTCTTCAATGCATGCCAGTTCGGCGTCTTTTTCTGATGGTGAGTTGGTCC
ACTCACAGGACAGGGGTGAGTTTGTTCCATTATCAGAGAAAATGGTTGAAGATAATTCTGATTCTCAACCAACTTTTCCCTACGTGTCTGCGGAGGTTAA
GAGCTTACCAAAAAGAAGAAATCTTAGTATTGATCATTATGATCACAGAGACCAAGTGGTTTTGGATGCTGTTGAGTCTGTAAAGCATTTGTTCATGGAG
GATAAAAAGTTGCATCTACAGATGCAAGAGAAATTTCACCTCTTGCATCAAATTTTGAATGGACAACAGGATGTACAGTTGGAAGAACCAACAAATCTAG
ATCCCATCATCCAATATGGCTCATGTGATAAACTTGAGAGATTGCACAAGAGGAGAAAGCTATCCCCTGAAGATGAGTCTGTTTTGAAACCTGTTCATGA
TACGGTTAAACCTGGCAACAAGAAAATTTTTGAAGGGAAAGTTATCGGTGAGCTAGATGCAACAAGTCCAAATGATGTTTCTATCAGGGCAGAACAGGCT
TTTGAAGAGACTGGTAAGAGTTATGTGAAAGCCGTCCTGGATATTTTGCCCAATTTTGAGGATATAGCTGATGTGGATTACATGAAATTGCTGGAGTTGG
ATAACAGTGCTGAGGAGGAAATATACAAGCAAGCAATGGAAATGCCGCTGTCCCCAACACTCCCTGAAATCAGTTTTCCGGAGGCTTTAGTATTATCTGA
TGTGCCAAGAAATGCAGAGCTTGGAAGCATCCACCATGAGGAGGCTTTAGTACTTGACACCATTGATATGGAACTGCACTGCAATGAATCAAGAGGTCAT
GGCTCTAAAATCACTCATGAGGGATCTCGACATGAGAAGGAAGATCTAGCCGGGGATGTCTATTCTCAATCACTGGAAGCAGATAAAGCTTCCAGTAGAT
GGTCGGGGAACTGTGGATATGGGCCTAAGACGGCACGTATTCCCACAATTTTTGTAACTACTTATGTTGAAGATTCCAGGTCCATTTACAGGGTCTTTTC
TGCCACAAAAGCTTGCATGGATTACTGTGCTGTAAATAGTCATCCTCACTCCATGCTACAGAGAATCCTGCATGCTGTCAGAATACAGGAAAAAATTTCA
TCTATGGAGAGAAGCTGTGTACTCTTCACCTTGATGGTTCTTAACTTCTCTGAATGCAAGTTGGTGAAAACACGAAGGATTTTGTATAAGGATCTAATTG
CTTATGTAGATTCTCTGGCTAGAGACCTCCAACCAGTGTTATCTACTGCAGGAGGGAGAAGCTCATTCGAAGAAGTGTGTTCTTTATATGAACTGCTTGG
CATCATTGACGACTTTCTCATCAATGGGACAATTTTTGAATGTACCAACTTTACTTCGGATTTCCGTAGTGAGGAAGAATTGAAGTTCAGCAGAAAATCT
GCTACAGTTGAACAGATGGTGGTTGGGAGTACTTTGCTGTCATCTATTTGCACATCAACTGGATATCTAGGATTCGTTTGTGAGACTTCCTACAATCTTC
TACGGAAATCCAGATGTGATACTGCCTTCCTATTGACGATATTTCATGTCTTTGCTCATGTTGTCCGGGAACAGTATTTCTCCTCTAATGACTTCGGCTT
GACCATGATTGTGGTGAGATCAATTGTTGGTTGTCTCGAAAGGAAAATGTCACTGGTTGCTTCAAGCCCTACTTCTCGTTCATCTGAAAATACTATTGAG
CTTCAGTTTATTTGCTGCCAAACATGCCCATTTGGTGCAGTTTCCATAGATGTAGTTACGTCTGTGCTGTTGGAAAAGCTCTGGGGTTCTGAGGATCCTG
GAACTACCCGTAGAACGGAATCAACAAGTTTGGCCAACTCCAATCTATTGTCGTTTGAGCATAATCAAGTTGGGGAAGGTATTCGTGGGTCAAAGTCTAT
CATGAATGGGACGATGGGTGATACAAGCAATATTCTTTCGTTATTAGAGCTTGTCTCCTGCAAAATGGGTTGGGAATGGACGCATAGCAAAGTTATACCC
GAGTTGCTTCAAATGTTGGAGAGATCTGTGACAACCACCTTTCAGATTGCTGCTGTTCTTTTCCTCGGTGAAGTTGGAAGGTTTGGAGTCGGTGCTTCAG
GTTCTGAAGATAAAGGAGTGGAGAGTCTAACATGCAAATTATCTAGCTTCTTTTGCCAGCTAATGGCCGAGGAAGCCATCTTCTCTGTTCAAATCGCCAC
AGTGGCCTCTCTTCTTAGGCTTCTGTTTCTGGACTTGCCAAGTATCGTGGACGAGTCCTTCAAATGCTCGACCACTGCGAGTCAGTCTGCTGCGATCGAT
CTGGTGAGGGACTGGTTTTCTCGTTTGAGCAAGGATCGGCAGGCAACGTCGTTAAGCTTCCTGCAATCTCAAATGGACACACGTTGA
|
||||||||||||||||||||
|
AA sequence
|
>Lus10023360 pacid=23181051 polypeptide=Lus10023360 locus=Lus10023360.g ID=Lus10023360.BGIv1.0 annot-version=v1.0
MSEVSASEKAQLNPCCESWSGKCSKLLEGRKALRKAVQLLKEELDKIQQDNANLKKGSCSNFTFPVSYCVTEFFPECEEARNIAAKESERKENELSEKMS
LENEVSALKSEISSLQQRGSLDAKGRDREIGLLREQVSETNKLKELLEKEKRRADAEKKNAETEKRRAAEAQKRMKAEKVKAVEENKHADVETKKAEGYR
VQLEVLKKDVEDMNSLLISEREEKARYLAIKTSLENEVSVLQSEISCLKKECAAGQQKEKETQNASEIFENANVENAGADRKHGGYCENTAETCLLQLQA
LMKEVEETKSMLVFEKEEHRKEVARRLLLEEEIVGLKSQICWLQQRGDEDIHGEIELLKGSLGGKEEEIRKLKELFDKVISQLNSEKKKVEVQKKIADEA
CERVKAEKSKLVTESKNASYEAKKAEKFQVQLDYLKKDFLKVQSELESEKEKFSMMKEKREVERQNVIKARQSVEAEIAKADHQRKLAQANGKVSAKERA
RADELSGQLELYKHKNEELQKQIQQFLHCQNSAELPGTLPVGKDKTKLTKMKLLKKQLKLGKMQLKHSKQVAEFEKHRNVILREELGNLRHDFVRISGRL
DLLAKCFSPNVEGIADMKKHSVKQFPNLNCYTFQDGALGEIQYLMKGRKLYDIEPHQMYTKKESEDFNPDCGATATTNLMKHANMLSPTCDLNLSTSGID
SQLESFQGGSNTKLLLHSSMHASSASFSDGELVHSQDRGEFVPLSEKMVEDNSDSQPTFPYVSAEVKSLPKRRNLSIDHYDHRDQVVLDAVESVKHLFME
DKKLHLQMQEKFHLLHQILNGQQDVQLEEPTNLDPIIQYGSCDKLERLHKRRKLSPEDESVLKPVHDTVKPGNKKIFEGKVIGELDATSPNDVSIRAEQA
FEETGKSYVKAVLDILPNFEDIADVDYMKLLELDNSAEEEIYKQAMEMPLSPTLPEISFPEALVLSDVPRNAELGSIHHEEALVLDTIDMELHCNESRGH
GSKITHEGSRHEKEDLAGDVYSQSLEADKASSRWSGNCGYGPKTARIPTIFVTTYVEDSRSIYRVFSATKACMDYCAVNSHPHSMLQRILHAVRIQEKIS
SMERSCVLFTLMVLNFSECKLVKTRRILYKDLIAYVDSLARDLQPVLSTAGGRSSFEEVCSLYELLGIIDDFLINGTIFECTNFTSDFRSEEELKFSRKS
ATVEQMVVGSTLLSSICTSTGYLGFVCETSYNLLRKSRCDTAFLLTIFHVFAHVVREQYFSSNDFGLTMIVVRSIVGCLERKMSLVASSPTSRSSENTIE
LQFICCQTCPFGAVSIDVVTSVLLEKLWGSEDPGTTRRTESTSLANSNLLSFEHNQVGEGIRGSKSIMNGTMGDTSNILSLLELVSCKMGWEWTHSKVIP
ELLQMLERSVTTTFQIAAVLFLGEVGRFGVGASGSEDKGVESLTCKLSSFFCQLMAEEAIFSVQIATVASLLRLLFLDLPSIVDESFKCSTTASQSAAID
LVRDWFSRLSKDRQATSLSFLQSQMDTR
|
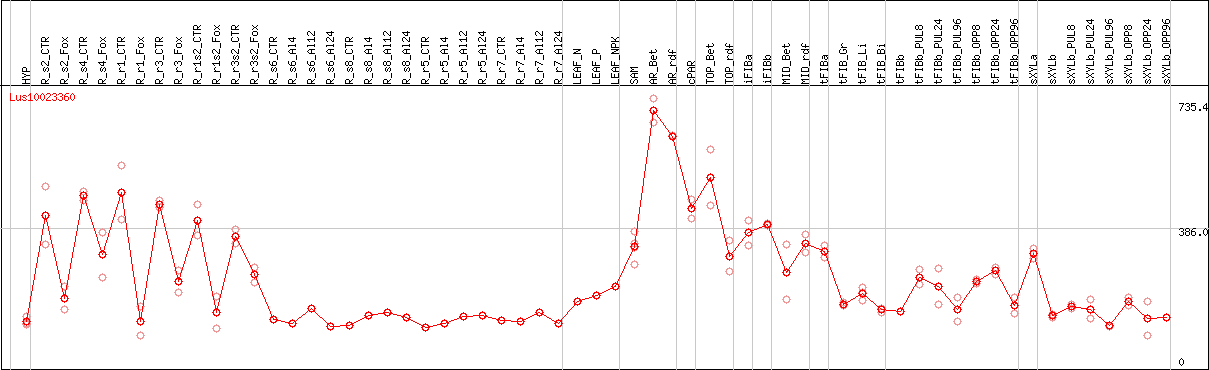
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10023360 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
