Lus10023486 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10023486 pacid=23160496 polypeptide=Lus10023486 locus=Lus10023486.g ID=Lus10023486.BGIv1.0 annot-version=v1.0
ATGGACGGGAGCGAGGATAAGGATGCCAAGGGTTCATCTAGTACAGTCACAGTTACTGAGCCAGTAGCTGCTAGTGACGCTGGTAGTAAGAAACTCTCTA
CTGCTGCTGTGGAAAACGTAGAGGAATCGAAATCGAGAGTAGTGCTTGGGTTTGATGGGCATGAAGTGGATTTGGAGAAAGATGCAGAAGAAGAAGAGGA
AGGGGAAGGCGTGTCTGGTACAGTTGATGGCTTGGGACCTGTTCAGGACACGGGTGATGAGGCATGGAGTCCGATAACTAAGAGTGTCGAGTGGAACTCG
TTGCAAGAGGATGTAGGTAAAGATGGCAAGGCCAAAGTTGATGATGAAGCTTTACCTGAAAGTGCCTCTCTAAAAGAGAGTTCGATAGATGAGGAAAGAC
TTGCAACTGATGTCTCCACGGGTGAGGGTACCGAACTTGTAGAGGGTTCGTCGGTTGTGGAGACTGTCGTCACAGAAACAGAAACAGAAACGGTTGTGGA
GACTGTAGTCACAACAACGGTTGTAGAGACTGAAGTCATGGAAACCCAAGTTGCTGATGATTCTGAAGATGGTGGGGATGGAGACACACATGTTATGGAT
AGTTTTGAGGATGGGGAACCGCCTCGGTTTGTACAGGTTTCAGAAGGCGGGGATACCCAAATCTTGGATGATTCTGGTGATAGCGATGCTGCATCTGCTG
AAGGGGAGACACAGGTTGTTCCGAGTTTAATGAATCAGGATGCCCAAGTTGCAGGGGTTCCTGGTAACAAGGACATAGATGGTTCTAGGGATGAGGAAAC
CCTTGCTACCCATGATTCCAAGGGCAGGGAAACTCAGGTTTTTGTGGATGGTTCCAAGGATGGGGAAAATGTGCTAGGTTCGGTGGCTGAAGGTACGCAG
GTCGTAGAGGATTCTATTGACAACGACATTGATTGTTGCCGGGAGGGTGAAGCCCAAGTTAATGATGGTCCCAAGCATGCCGAAGCTCAGATTTTACAAG
ATTCAGTGGGTAAGGATGTCACTGAAGTTAATGATCATACTAACTATGGAGTAACTCAGGTTACACACAGTTTATCTGATGATACCCAAGTCATAGAGGA
TCGTGATACTCATGACGTGGATGTTTCCAAGAATGTGGACACTGAAATAACTTATAGTTCCAGTGGTCAGGAGACTCAAGTTGCTGATGATTCGATGGAT
GTGGAAACTAAGGTTATCAACGGTTCAATAGATGCTGAAGAACAAGTCAAAGTTGCCGATACTCCTGCCATTTCAGAAGATGAAAATGTGGAGAGGGGTT
TAACAGATGAAGGTGCTCCTATTCCAATTGAGGATCTAAATTATAAGGAAGTTGGTGCATCGGACCAAGACATCAATGATGCAGGTATGTCAGATGATTC
GATGGATGTGGAAACTGAGGTTGTCAATGGTTCAGTACATGCTCAAGAACAAGTCACAGATGCTGATACTTCTGCCGTTTCAAAAGATGAGAATGTAGAG
AGAGGTTTAACAGAGGAAGTTGCTCCTACACCGATTGGGGATATAAATTCTCAGGAAGTCAATGCATTGGAGCATGACATCAAGGATCTAAGTACGTCTG
GTGGTTCGGAAGTCACTTGTGGGGAAAATAATCAAGTTTCTGCTGCTACTGAGAAAACCGAGTTGGTTAAAGAGCAGCCAATGGAAATTGTTGAAACTAA
ACAAGATGTTGCTCAATATTCAGGGTTTAAACTGGAATCTGCTTTCCCACGTGAAGCAACAAGTGTTGATCCCGTGGACCAGGTACCTAGAATTGATGCT
CTGGATGGAATGACATCCAGTTCTGTCAATGATTCAGTTCTAGTGGAGAAAAATGAGATTGAGTATCCCACCTGTGAAACTCTGGACAAAGTAATGCAAG
TTTCTGTTCAAGGGAACATTGCATCAGTGGACGATGGAGATATTACTTGTCCGAACATTGAAGGTATGGATGCTGATGCTTTTAATGAAAATTTTTGCTT
TTCAGTGGAAGAACTCCAGGCACCAGGTGAAACAGTCAACGGGAGCACTGAACATCACTATAATGCACTTGCAGATCTACCATCTTTCTCTCAACCAACT
GATGTGTTTATTGGAGGTGAATTTGCTGGTACCATGGAAGAGAAGGTATTTCCGTTTTCCGGAAAAGAACACTTCATACACCAGTCTGTCTCTTTTGGTC
CTATTAGGGATAACTCAAACGTAGAGCAAGTAATGATAAATGGCATACAAATTACTTATGCTGAACAAACTTGTCTGCAGAAACAGCAGGAAACTAGAGT
TGCAGCTGGCAAGCTAGTGGATGCAGATAAAGAGGGTGTAGCAAATGGTAGCATTCATTCTCCAGATTCAACTTCTTCTTTCCATTCTACCCAGGCAGTT
GCTGGAGACGATGTTGCCCAAATGGATCACATTGTCGGTTTAGATGCAAGAGCCTCTGGAGACGATCAGTCAGCTTCCTTCCACAATGAAGTTTTAGCTT
CCACCACCGGTTTCACAAAAGCCTCTGTGAACAATGTAGAGTTGGTTGTAGAGACCAGTATTCTTGATGTGAAAGAAAGGGGAGTTGATACTACTGAGGA
CAAATCTCCTCAATGGGCTAGTTTACATTGGGGAAGCTTTGCAAAGGCACAGCAAGCTAAATATCAGCTGCCATCAGACTTCGATGGTGAATTCTGTGTA
TCTGATTTGGTTTGGGGAAAAGTTAGGAGTCATCCTAGGTGGCCTGGGCAGATATTTGACCCTGCAGATGCGTCTGAGCAAGCATTGAAGTATCAGAAGG
ACAGTTTTTTGGTAGCATATTTTGGGGACCGTACATTTGCCTGGAATGAAGCACATCTTTTAAAGCCATTTAGATCCCATTTTAGCCAGTCAGAGAAGCA
AAGCAATTCAGAAGCATTCCAGAATGCTGTTGATTCTGCTTTGGAAGAAGTTTCTAGGCGAGTCGAAATGGGTTTGGCTTGCTCCTGTGTGCCAAATGAA
GATCGATACAAAATCGAGTCAGAGCTTGTTGGTAATGCCGGAATCCAAGACGAAGCCATCACAAGAGGTGACTTGAACGTATCTAGAACTGCAGGGATGT
TTGAACCTGTCAATTTGATTAATTATGTGAAATCTCTGGCGCAATCCCCAGCTTCCGGGGCTGATCGTGTAGAGCTCGCCATTCTAAGGTCTCAGTTGCT
CTCCTTCTATCGTTTAAAAGGCTATTCTTCCCTGCCTGAGTTTCAGTGCTTTGGTGAGTCGTTGGAGGATGGCGAGGCAATTAATCATGCTTGTGAAGAT
GACTTGCAACCTTCCACTTTGCCAAATTCCCATACTCAAAGACCTGCTTATCATAAACGTAAACATAACTTGAAGGATGGTGTTTATCGTGGTAAAAAGG
TTAAAAGCTCGACCCTTTTGATGGATGACTCGTGGGATTCCTTAGAGGTTGAAGTGGGAGCCGAGGCAAAGAGCCGCAAGGGTCCCGAAGAGGTAAAAAC
GATATCTCTCACTAAAATATCGACTTCTCCGAAGCCTTCCTTTAGGATCGGCGAATGTATTCGGCGAGCAGCTAGTCAAATGACGGGATCGCCCTCAATC
CTGAAGTCAAACAGCCAGAAGTTCGAGTTCGATGATGAATTTGACCATAGCATGCCACAGTGTGACGAGGGAGAAAATAACAGCGTGGATGCATCTGTAG
ACGAGTTGCTAGTGCAGCTTCAGTCAGCAGCACTGCAAAGATGTAGTTCTTCGAATGGTATTGTTGGGTTCTTCTCTGATTATAGAAACTCTGTGGTTGT
TGACCAACAAGATGTTGTGCTGGTTGATAAGAGGAGGAAAGTGTCTCATTCGGTTGGCGATTCAGAAACGTACGAGTTTGTGGATATGAACGACACTTAC
TGGACCGACAGGGTGATTGAAAACGGTTCAGAAGAGAAGCCTACTCAGACCGACCGGGTAATTGAAAATGGTGGTTCGGAAGAGAAGCCCACTCGAAAGA
GCAAAAAACGGGAGAATCTATTCGTACCGGTCAGTCTCGATAAACCGGTTCAGAAGTCGAATGCCAGGAAACGATATTCTGACGGCAACAACTTCAACGT
GTCGCAGGAGAAACCACCCGGTTATGTGGACCCCAAGGCTCCGGCGGAGCTTGTGATGCATTTCCCAGTGGTGGATTCGGTTCCTTCCGAGATGAAACTG
AACAAGATGTTCAAATGTTTTGGACCGTTAAAAGAATCCGGGACAGAGGTTGATCGGGATACTAACCGTGCCAGGGTGGTGTTTAGGCATTCGTCTGATG
CAGAGGCTGCTTATGGTAGTGCATCAAAGTTCAACATTTTCGGGTCGTTGCTTGTAAATTACCAGCTCAACTATACCATATCCGTGCCGTTCAAATCTGA
ACCGATCTCCATTACTCATGGAGACGAAGATGGAAATGTCTTTCTCCAATACTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10023486 pacid=23160496 polypeptide=Lus10023486 locus=Lus10023486.g ID=Lus10023486.BGIv1.0 annot-version=v1.0
MDGSEDKDAKGSSSTVTVTEPVAASDAGSKKLSTAAVENVEESKSRVVLGFDGHEVDLEKDAEEEEEGEGVSGTVDGLGPVQDTGDEAWSPITKSVEWNS
LQEDVGKDGKAKVDDEALPESASLKESSIDEERLATDVSTGEGTELVEGSSVVETVVTETETETVVETVVTTTVVETEVMETQVADDSEDGGDGDTHVMD
SFEDGEPPRFVQVSEGGDTQILDDSGDSDAASAEGETQVVPSLMNQDAQVAGVPGNKDIDGSRDEETLATHDSKGRETQVFVDGSKDGENVLGSVAEGTQ
VVEDSIDNDIDCCREGEAQVNDGPKHAEAQILQDSVGKDVTEVNDHTNYGVTQVTHSLSDDTQVIEDRDTHDVDVSKNVDTEITYSSSGQETQVADDSMD
VETKVINGSIDAEEQVKVADTPAISEDENVERGLTDEGAPIPIEDLNYKEVGASDQDINDAGMSDDSMDVETEVVNGSVHAQEQVTDADTSAVSKDENVE
RGLTEEVAPTPIGDINSQEVNALEHDIKDLSTSGGSEVTCGENNQVSAATEKTELVKEQPMEIVETKQDVAQYSGFKLESAFPREATSVDPVDQVPRIDA
LDGMTSSSVNDSVLVEKNEIEYPTCETLDKVMQVSVQGNIASVDDGDITCPNIEGMDADAFNENFCFSVEELQAPGETVNGSTEHHYNALADLPSFSQPT
DVFIGGEFAGTMEEKVFPFSGKEHFIHQSVSFGPIRDNSNVEQVMINGIQITYAEQTCLQKQQETRVAAGKLVDADKEGVANGSIHSPDSTSSFHSTQAV
AGDDVAQMDHIVGLDARASGDDQSASFHNEVLASTTGFTKASVNNVELVVETSILDVKERGVDTTEDKSPQWASLHWGSFAKAQQAKYQLPSDFDGEFCV
SDLVWGKVRSHPRWPGQIFDPADASEQALKYQKDSFLVAYFGDRTFAWNEAHLLKPFRSHFSQSEKQSNSEAFQNAVDSALEEVSRRVEMGLACSCVPNE
DRYKIESELVGNAGIQDEAITRGDLNVSRTAGMFEPVNLINYVKSLAQSPASGADRVELAILRSQLLSFYRLKGYSSLPEFQCFGESLEDGEAINHACED
DLQPSTLPNSHTQRPAYHKRKHNLKDGVYRGKKVKSSTLLMDDSWDSLEVEVGAEAKSRKGPEEVKTISLTKISTSPKPSFRIGECIRRAASQMTGSPSI
LKSNSQKFEFDDEFDHSMPQCDEGENNSVDASVDELLVQLQSAALQRCSSSNGIVGFFSDYRNSVVVDQQDVVLVDKRRKVSHSVGDSETYEFVDMNDTY
WTDRVIENGSEEKPTQTDRVIENGGSEEKPTRKSKKRENLFVPVSLDKPVQKSNARKRYSDGNNFNVSQEKPPGYVDPKAPAELVMHFPVVDSVPSEMKL
NKMFKCFGPLKESGTEVDRDTNRARVVFRHSSDAEAAYGSASKFNIFGSLLVNYQLNYTISVPFKSEPISITHGDEDGNVFLQY
|
DESeq2's median of ratios [FLAX]
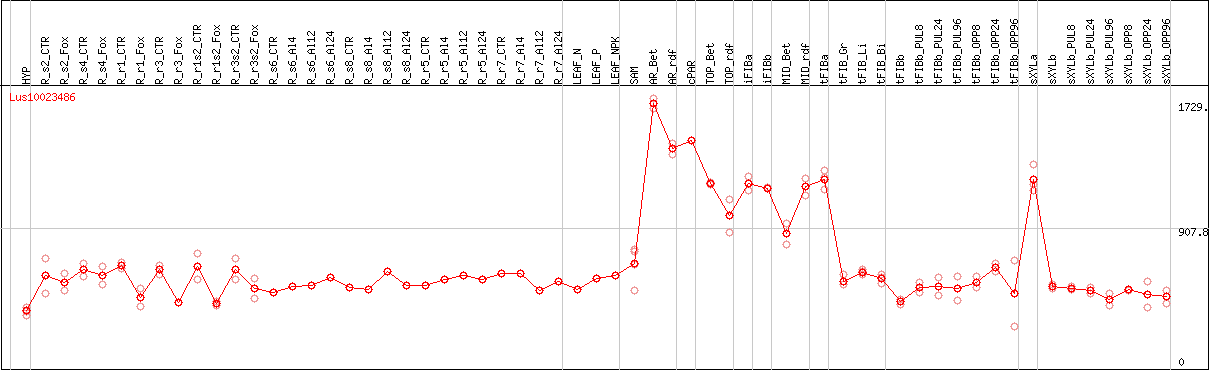
Coexpressed genes
Lus10023486 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
