External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G20330 458 / 1e-163
FRL1, CVP1, SMT2
FRILL1, COTYLEDON VASCULAR PATTERN 1, sterol methyltransferase 2 (.1)
AT1G76090 427 / 2e-151
SMT3
sterol methyltransferase 3 (.1)
AT5G13710 213 / 2e-67
CPH, SMT1
CEPHALOPOD, sterol methyltransferase 1 (.1.2)
AT1G64970 78 / 2e-16
VTE4, TMT1, G-TMT
VITAMIN E DEFICIENT 4, gamma-tocopherol methyltransferase (.1)
AT1G73600 59 / 2e-09
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT3G18000 58 / 2e-09
XPL1, NMT1, XIPOTL1, PEAMT
XIPOTL 1, N-METHYLTRANSFERASE 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G48600 58 / 2e-09
AtPMEAMT
phosphoethanolamine N-methyltransferase, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT1G78140 49 / 2e-06
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G69523 48 / 4e-06
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT2G41040 45 / 4e-05
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005625
408 / 1e-143
AT1G20330 582 / 0.0
FRILL1, COTYLEDON VASCULAR PATTERN 1, sterol methyltransferase 2 (.1)
Lus10029498
122 / 4e-33
AT5G13710 380 / 2e-133
CEPHALOPOD, sterol methyltransferase 1 (.1.2)
Lus10004158
108 / 9e-30
AT1G20330 142 / 9e-43
FRILL1, COTYLEDON VASCULAR PATTERN 1, sterol methyltransferase 2 (.1)
Lus10009537
77 / 1e-15
AT1G64970 453 / 3e-160
VITAMIN E DEFICIENT 4, gamma-tocopherol methyltransferase (.1)
Lus10020357
73 / 2e-14
AT1G64970 434 / 7e-154
VITAMIN E DEFICIENT 4, gamma-tocopherol methyltransferase (.1)
Lus10039600
64 / 2e-12
AT5G13710 174 / 2e-54
CEPHALOPOD, sterol methyltransferase 1 (.1.2)
Lus10031348
64 / 2e-11
AT3G18000 843 / 0.0
XIPOTL 1, N-METHYLTRANSFERASE 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10009056
62 / 1e-10
AT1G73600 773 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Lus10042006
56 / 1e-08
AT3G18000 841 / 0.0
XIPOTL 1, N-METHYLTRANSFERASE 1, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G245800
453 / 2e-161
AT1G20330 667 / 0.0
FRILL1, COTYLEDON VASCULAR PATTERN 1, sterol methyltransferase 2 (.1)
Potri.002G016300
452 / 2e-161
AT1G20330 647 / 0.0
FRILL1, COTYLEDON VASCULAR PATTERN 1, sterol methyltransferase 2 (.1)
Potri.001G263700
223 / 4e-71
AT5G13710 587 / 0.0
CEPHALOPOD, sterol methyltransferase 1 (.1.2)
Potri.009G058600
218 / 3e-69
AT5G13710 591 / 0.0
CEPHALOPOD, sterol methyltransferase 1 (.1.2)
Potri.016G056000
177 / 1e-53
AT5G13710 513 / 0.0
CEPHALOPOD, sterol methyltransferase 1 (.1.2)
Potri.013G077000
72 / 3e-14
AT1G64970 441 / 8e-156
VITAMIN E DEFICIENT 4, gamma-tocopherol methyltransferase (.1)
Potri.012G047400
58 / 3e-09
AT1G48600 862 / 0.0
phosphoethanolamine N-methyltransferase, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.015G039000
58 / 3e-09
AT1G48600 862 / 0.0
phosphoethanolamine N-methyltransferase, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.018G120000
49 / 1e-06
AT2G41040 418 / 6e-147
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.002G095100
47 / 8e-06
AT1G78140 442 / 2e-156
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF08241
Methyltransf_11
Methyltransferase domain
Representative CDS sequence
>Lus10023995 pacid=23160779 polypeptide=Lus10023995 locus=Lus10023995.g ID=Lus10023995.BGIv1.0 annot-version=v1.0
ATGGATTCAACCACAATCCTCTTCACACTATTCATCCCTGTCCTCTTCGCCGTCCTCTACTGGTTCGTCTGCCTCCTGGGATCGGCCGAGGTCAAGGGCA
AGCACGCCTCCCAGCTCACCGGCGGCTCAATCTCCGGCGAGAAAATCCAGGACAAGTACAACCAGTACTGGTCCTTCTTCCGATCCCCTTCCCAAATCGA
GACCGCCGACAAAGTCCCCGCCTTTGTCGACACTTTCTACAACCTCGTCACCGACATCTACGAGTGGGGGTGGGGCCAGTCTTTCCACTTCTCCCCTTCA
TTGCCGGGGAAATCTCACCGTGACGCCACGCGCCTCCACGAGGAATTCGCCGTCGATCAGCTCAATGTCAAGCCGGGGGACAAAATCCTCGACGCCGGAT
GCGGAGTCGGTGGGCCCATGAGGGCGATTGCGGCCCACTCCGGTGCGAATGTCGTCGGGATCACAATCAACGAGTATCAGGTAAATAGGGCACGGATGCA
TAACAAGAAGGCCGGATTGGACGGCCTTTGCGAGGTTGTTAAGGGTAATTTCTTGTCCATGCCGTTTGAGGATGAGAGCTTCGACGGAGCTTATTCAATT
GAAGCGACCTGCCATGCTCCCAAACTTGAAGAGGTTTACGCTGAGATTTTTAGGGTTTTGAAGCCTGGAGCTGTTTATGTTTCCTACGAGTGGGTCACCA
CTGATTTGTTCAAAGCTGAGGATCCAGTTCACGTCGAGATAATCGAAGGGATTGAGAGAGGGGATGCTTTGCCAGGGCTGAGGAGGTACAACGACATTGC
TGAAACCGCCATGAAAGATGGGTTTGAAGTGTTAGGTGATGATGGATGA
AA sequence
>Lus10023995 pacid=23160779 polypeptide=Lus10023995 locus=Lus10023995.g ID=Lus10023995.BGIv1.0 annot-version=v1.0
MDSTTILFTLFIPVLFAVLYWFVCLLGSAEVKGKHASQLTGGSISGEKIQDKYNQYWSFFRSPSQIETADKVPAFVDTFYNLVTDIYEWGWGQSFHFSPS
LPGKSHRDATRLHEEFAVDQLNVKPGDKILDAGCGVGGPMRAIAAHSGANVVGITINEYQVNRARMHNKKAGLDGLCEVVKGNFLSMPFEDESFDGAYSI
EATCHAPKLEEVYAEIFRVLKPGAVYVSYEWVTTDLFKAEDPVHVEIIEGIERGDALPGLRRYNDIAETAMKDGFEVLGDDG
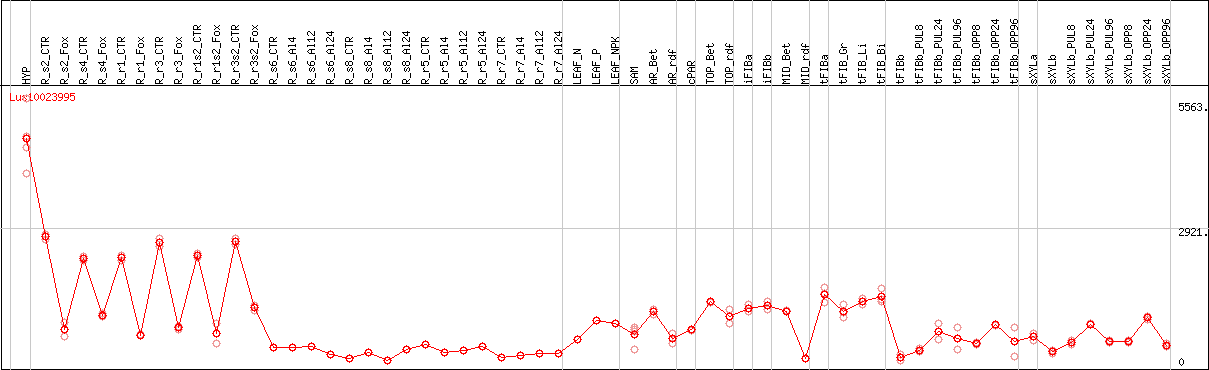
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10023995 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.