Lus10024041 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10024041 pacid=23141572 polypeptide=Lus10024041 locus=Lus10024041.g ID=Lus10024041.BGIv1.0 annot-version=v1.0
ATGGAAGACGTCTGCGAAGGAAAGGACTTTTCCTTCCCGAAGGAGGAGGAAAAAATTCTCGCCTTCTGGTCGCAAATCCAGGCCTTCAAGACTCAATTGT
CACGCTCAGAGTCGCTGCCGGAGTACATTTTCTATGACGGTCCTCCCTTTGCTACCGGCCTTCCTCACTACGGCCACATCCTTGCTGGAACTATCAAGGA
TATCGTGACCCGGTACCAGACCACGACCGGCCACCACGTCACGCGGCGGTTCGGATGGGATTGCCATGGCTTGCCGGTAGAGAACGAGATTGATAGGAAG
CTTGGGATTAAGAGAAGGGAGGAAGTGTTGCAAATGGGGATTGGCAAGTATAATGAGGAGTGTAGGAGCATCGTCACTCGTTATGTTGAGGAGTGGGAGA
AGATTATCACGAGAACTGGGCGCTGGATTGATTTCGATGATGGCTATAAAACTATGGACTTGAAGTTTATGGAGAGTGTTTGGTGGGTTTTCGGTCAGCT
TTTCGAGAAGGGTCTCGTTTACAAAGGTTTCAAGGTCATGCCGTATAGCACTGGTTGCAAAACTGTACTGTCGAATTTCGAAGCTGGCCAGAATTTTAAG
GATGTACCTGATCCTGAAATTATGGTGGCCTTTCCTATTGTTGATGATCTCCATAATGCTGCGTTTGTGGCCTGGACAACTACGCCCTGGACTCTTCCTA
GTAATCTTGCACTTTGTGTCAACCCTACCTTTGATTATGTTAAGGTACGCAACAAGTATTCTGGGAAAGTCTATGTGATTGCAGAATGTCTACTGTCCTC
CATTCCCACCGAGAAGCCAAAATCAAGTGATTCAAAGACATCAAGTTCAAAGGCGAAGGGTGCCAAGACTGAAAACCCTATGGATTCATATGAATTACTA
GAGAAAGTCAAGGGCAAAGAACTTGTGAATAAAAAGTATGTACCACTATTCAATTATTTCCAGGAGTTCTCTGGCACAGCATTCAGAGTTGTCGCTGATA
GTTATGTGACATCTGACCGGGGTACTGGAATTGTTCATTGTGCTCCAGCATTTGGAGAGGATGATTACCGTGTTTGCATTGAGAATGAAATAATCAGCAA
GGGGGAGAATTTGATTGTTGCGGTTGATGATGATGGGTGCTTTACTGATAGAATTTCTGATTTCTGTGGGCGCTATGTCAAAGATGCGGACAAAGATATT
ATCGAGGCAGTGAAGGGAAATGGCAGACTTGTCAAAACTGGAAGCTTTATGCACAGCTACCCATTCTGTTGGAGATCTGACACTCCACTTCTTTATAGAG
CTGTTCCAAGCTGGTTTGTTAAAGTGGAAAGTTTGAAAGATCAATTATTGGAAAGCAACAGTCAGACATATTGGGTACCTGGTTATGTGAAGGTTGGGAT
TCTTAAGTCCACTCCTTTCTATATGAGAACAGAAGATAAGAGATTCCATAATTGGTTGGAAAATGCAAGAGACTGGGCTATTAGTCGCAGCCGATTTTGG
GGTACTCCTCTCCCTGTATGGACCAGTGCAGATGGCGAAGAAATAGTAGTTGTTGATTCTGTCGCTAAACTTGAAAGTCTTTCTGGAGCCAAGGTGTTTG
ATCTGCATCGTCACAATATTGATCACATCACAATTCCTTCTAGTCGTGGCCCTGAATTTGGGGTGCTTCGACGCGTTGATGATGTCTTTGATTGTTGGTT
TGAGAGCGGGTCGATGCCTTATGCGTACATCCATTATCCTTTTGAGAATGTTGAGTTGTTTGAGAAGAATTTTCCTGGGCATTTTGTGGCTGAAGGATTA
GATCAAACTCGTGGATGGTTCTATACTCTCATGGTGTTGTCAACTGCATTATTTGGAAAGCCGGCATTTAGGAATCTTGTATGCAATGGGCTGGTCTTAG
CCGAGGATGGTAAAAAGATGAGTAAAAAGTTGAGAAACTACCCATCACCAATGGACGTGATCAATGATTATGGAGCGGATGCTCTAAGATTGTACCTCAT
AAATTCCCCAGTTGTCCGAGCTGAAACCTTGCGGTTCAAGAAGGATGGAGTATATAATGTTGTGAAAGATGTCTTTCTTCCGTGGTACAATGCTTACAGA
TTTCTTGTTCAGAATGCTAAGAGACTTGAGATTGAAGGACTTGCAGTATTTCGTCCTCTTAGTTCATCAGATCTGCAGAAATCTTCTAATGTGCTTGACC
AGTGGATCAACTCTGCTACTCAGAGTCTTGTTCATTTTGTTCGCCAAGAAATGGATGCATACAGACTTTACACGGTGGTTCCGTATCTTCTGTCATTTCT
TGACAACCTAACCAATGTATATGTCCGTTTCAATCGCAAGAGATTGAAAGGTCGTACGGGGGAAGATGATTGCCGAATTGCTCTTTCCACTCTCTACAAT
GTGCTTCTAACTTCTTGCAAAGTGATGTCATCATTCACGCCCTTCTTCACAGAGGTTCTCTATCAAAATTTGCGGAAAGTTAGTGCCAACTCTGAAGAAA
GCATTCACTACTGCAGTTATCCTGCGGAAGAGGGAGAGAGAGATGAACGCATTGAGCAAAGTGTCACAAGGATGACGACCATTATCGATCTTGCTCGGAA
TATCCGTGAGAGACATAACAAACCTCTCAAGTCACCACTTAGGGAGATGGTGATTGTTCATCCTGATGCAGATTTCCTTGATGATATTGCTGGAAAACTA
AGAGAGTATGTGCTAGAGGAACTCAATGTACGATCTCTTGTCCCATGTAAAGACACTTTGAAGTATGCCTCTTTACGTGCGGAGCCCGAGTTCAGTGTGT
TGGGTAAACGACTTGGCAAGGATATGGGAGTTGTCTCAAAAGTAATAAAATCGATGTCACAGAAAGATATTCTGGAATTTGAGAAAGCAGGAGAGGTCAC
CATTAAGTCGCACGTCCTAAAATCAGACGATATTAAGGTTCTCCGAGACTTCAGATGTCCTGATGGATTGACAGACAAGGAGATTGATGCTGCTGGAGAT
GGTGATGTTTTGGTGATTTTGGATTTACGTCCTGATGAGTCGTTGTTTGAGGCTGGTCTTGCTCGTGAGGTGGTGAATAGAATTCAAAAATTGCGGAAAA
AGATTGCCCTTGAACCGACTGATGCTGTGGAGGTATACATCGGCTCCTTGGATGACAATAAGCCAAGGTTGGAAAGAGCTCTTAATTCTCAGGAACTCTA
CATCAGAAATGCAATTGGTTCTCCGCTGCTTCCATATACTGCAATCCCCCCGCATGCTGTGGTGATTGGAGAAGAGAGTTTCCATGGAATTTCTGGGTTA
TCGTTCACTATTTACTTAACAAGGCCAACGCTGGTGTTCAAGTCAGATGCTTGTGCTGAAGTATATGAAGGAAATACAAGATTTGCCCAGATGCTGCAAG
TGTACTTGCTATCTAGGGATCATAGCAGTTTGAAATTGGAATTTCAACAAGGAAATGGCAAGGTTGATCAAGGTAGATTCACTCGAGAAGTTGCCTGCCG
TGGAATTGACGTTGGAGAAGCATGTGTACTTGAGTGTCGGAGACCAACTTTTGGGAACAACAAGACAATCAGACTTGAAGGGATGCTAATCCTAACAGAA
TCAGGGATGGCAAAGAGCCCCACCAGCTCCTTTCCTAGCACTCCTGATAACTCTTCCTCCAGGGGTGCTTCAAGTACTGCTTCTGACAAGAAGAAGAAGT
CTATAATTCCCAAGATCTTTAGCTCAAAGAAAATCAACAGAGGAGGATCCGGCGACGATGTCCTAATTGCATCAGATGGAGATTGTCCTGCCCCTGCTGA
AAGAGGGACAAGTCCTGGAATTGAAGGCCTCAATCTCTCCAACTTTGATCACTCCATGGCATCATCCACCACAGAAATCAAAGAATTCCGGATTTACGTA
GCGACATGGAACGTCGGAGGAAGGACTCCTGATGCCAGCATGAATCTCGAAGATTTCCTGCAAGTGGACGAATCCCCCGATTTATACATTTGCGGGTTTC
AGGAAATTGTTCCACTCAGTGCCGGAAATGTGCTGGTGATAGAAGACAGCGAGCCGGCAGCCAAATGGCTAGCCCTCATAAGCCAAACCATCAACAGAAC
AAACTACAACTACTTCACCCCTACCTCTCCGAATCCGGCACATAGATCTTCCAACAAGTCTCAAAACAGTAAGGAAACAAAAAACCACAATAACTTCTTC
CACAAGCCATCCGTCAGAGTCCTGACGAAAAACCTGAGGGCGGACAGCAGTAGCCTTCTGAAGATCTGCAATTGTGCGGGAGATTTCCCAACGCGAGACA
GGCAGCATCGAGGGGGAGAATTCAGCGGTCCGCTTCCGATGGGAACCAGCGCAGGTCAGAATTTCGGGCTCGACAATGACTCGCATAATTCTCCTGCAGC
TAATAGCCCCGGAGGAGCCTCCCCTGGATCCACCACCAACCAAATGGTGTACAACCTCATCTCTTGCAAGCAAATGGTCGGGATTTTCCTTTCGGTTTGG
GCTCGACAGGAATTGGTCCCCCACATTGGTCACCTCAGAGCATCCTCTTGTGGGACCGGCATCATGGGATGCCTCGGGGAGAAAGAAGGGGACGAGCTGA
AGAGGAATTCTGATGTTGCTGAAATCCTGAAGAGCATTCAGTTCCCCAAGATTTGCAAGAATCAAATCAGCCGAGCTCCAGAAAGGATAGCCGATCACGA
TCGAGTGCTTTGGTTCGGAGATTTGAACTATCGGATCTCTCTCAGCTACGAAGAAACTAAAGTCCTGCTTGAAGAAAACGATTGGGACACCCTTCTGGAC
AGGGATCAGCTGAATATGGAAAGAGATGCAGGGAGGGTATTCGACGAGTTTTACGAAGGCAGGATTCTATTCGCTCCCACTTACAAATACTCACATAACT
CCGATTCCTACGCTGGAGAGACCGTAAAGAACAAGAAGAAACGTCGAACACCAGCATGGTGCGATCGAGTATTATGGAAAGGAAGAGGAATGGAGCAAAT
GTCGTACGGGAGAGTGGAGTCGAGATTCTCGGATCACAGACCGGTTTGCGCAATATTTGCGGTGGATGTGGAAATGAGTGCGACGAGAAGCAGCAGGTTC
AGGAAAGGCTACTCTTGTACAGCTCCAAGGATTCAATATGACAACGACTGTTTGCCGCATCAAAGGCAAAACAACTTCTATGGCTACTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10024041 pacid=23141572 polypeptide=Lus10024041 locus=Lus10024041.g ID=Lus10024041.BGIv1.0 annot-version=v1.0
MEDVCEGKDFSFPKEEEKILAFWSQIQAFKTQLSRSESLPEYIFYDGPPFATGLPHYGHILAGTIKDIVTRYQTTTGHHVTRRFGWDCHGLPVENEIDRK
LGIKRREEVLQMGIGKYNEECRSIVTRYVEEWEKIITRTGRWIDFDDGYKTMDLKFMESVWWVFGQLFEKGLVYKGFKVMPYSTGCKTVLSNFEAGQNFK
DVPDPEIMVAFPIVDDLHNAAFVAWTTTPWTLPSNLALCVNPTFDYVKVRNKYSGKVYVIAECLLSSIPTEKPKSSDSKTSSSKAKGAKTENPMDSYELL
EKVKGKELVNKKYVPLFNYFQEFSGTAFRVVADSYVTSDRGTGIVHCAPAFGEDDYRVCIENEIISKGENLIVAVDDDGCFTDRISDFCGRYVKDADKDI
IEAVKGNGRLVKTGSFMHSYPFCWRSDTPLLYRAVPSWFVKVESLKDQLLESNSQTYWVPGYVKVGILKSTPFYMRTEDKRFHNWLENARDWAISRSRFW
GTPLPVWTSADGEEIVVVDSVAKLESLSGAKVFDLHRHNIDHITIPSSRGPEFGVLRRVDDVFDCWFESGSMPYAYIHYPFENVELFEKNFPGHFVAEGL
DQTRGWFYTLMVLSTALFGKPAFRNLVCNGLVLAEDGKKMSKKLRNYPSPMDVINDYGADALRLYLINSPVVRAETLRFKKDGVYNVVKDVFLPWYNAYR
FLVQNAKRLEIEGLAVFRPLSSSDLQKSSNVLDQWINSATQSLVHFVRQEMDAYRLYTVVPYLLSFLDNLTNVYVRFNRKRLKGRTGEDDCRIALSTLYN
VLLTSCKVMSSFTPFFTEVLYQNLRKVSANSEESIHYCSYPAEEGERDERIEQSVTRMTTIIDLARNIRERHNKPLKSPLREMVIVHPDADFLDDIAGKL
REYVLEELNVRSLVPCKDTLKYASLRAEPEFSVLGKRLGKDMGVVSKVIKSMSQKDILEFEKAGEVTIKSHVLKSDDIKVLRDFRCPDGLTDKEIDAAGD
GDVLVILDLRPDESLFEAGLAREVVNRIQKLRKKIALEPTDAVEVYIGSLDDNKPRLERALNSQELYIRNAIGSPLLPYTAIPPHAVVIGEESFHGISGL
SFTIYLTRPTLVFKSDACAEVYEGNTRFAQMLQVYLLSRDHSSLKLEFQQGNGKVDQGRFTREVACRGIDVGEACVLECRRPTFGNNKTIRLEGMLILTE
SGMAKSPTSSFPSTPDNSSSRGASSTASDKKKKSIIPKIFSSKKINRGGSGDDVLIASDGDCPAPAERGTSPGIEGLNLSNFDHSMASSTTEIKEFRIYV
ATWNVGGRTPDASMNLEDFLQVDESPDLYICGFQEIVPLSAGNVLVIEDSEPAAKWLALISQTINRTNYNYFTPTSPNPAHRSSNKSQNSKETKNHNNFF
HKPSVRVLTKNLRADSSSLLKICNCAGDFPTRDRQHRGGEFSGPLPMGTSAGQNFGLDNDSHNSPAANSPGGASPGSTTNQMVYNLISCKQMVGIFLSVW
ARQELVPHIGHLRASSCGTGIMGCLGEKEGDELKRNSDVAEILKSIQFPKICKNQISRAPERIADHDRVLWFGDLNYRISLSYEETKVLLEENDWDTLLD
RDQLNMERDAGRVFDEFYEGRILFAPTYKYSHNSDSYAGETVKNKKKRRTPAWCDRVLWKGRGMEQMSYGRVESRFSDHRPVCAIFAVDVEMSATRSSRF
RKGYSCTAPRIQYDNDCLPHQRQNNFYGY
|
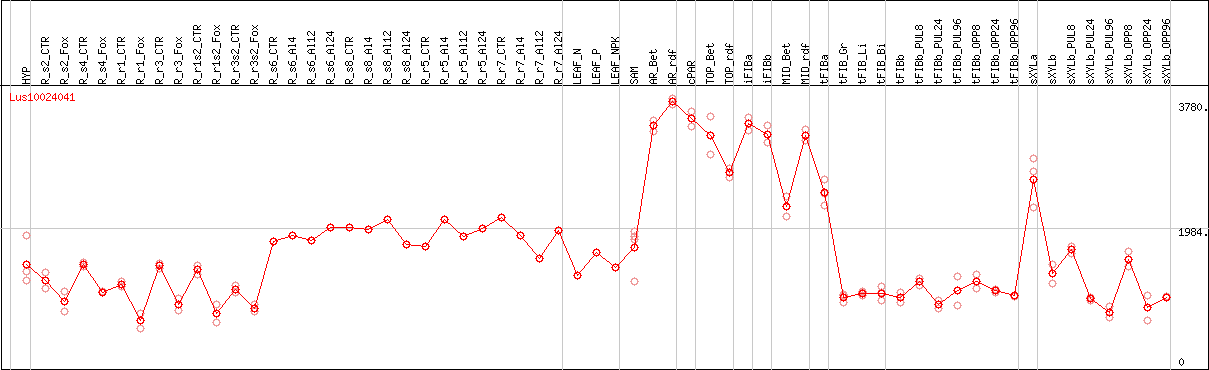
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024041 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
