External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G03210 129 / 5e-36
ATFUT2, FUT2
fucosyltransferase 2 (.1)
AT2G03220 121 / 5e-33
ATFUT1, ATFT1, FT1, MUR2
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
AT1G14070 117 / 8e-32
FUT7
fucosyltransferase 7 (.1)
AT1G14110 117 / 1e-31
ATFUT9, FUT9
fucosyltransferase 9 (.1)
AT1G14080 112 / 9e-30
ATFUT6, FUT6
fucosyltransferase 6 (.1)
AT1G14100 112 / 1e-29
FUT8
fucosyltransferase 8 (.1)
AT2G15350 109 / 6e-29
ATFUT10, FUT10
fucosyltransferase 10 (.1)
AT2G15390 109 / 1e-28
ATFUT4, FUT4
fucosyltransferase 4 (.1.2)
AT2G15370 108 / 2e-28
ATFUT5, FUT5
ARABIDOPSIS FUCOSYLTRANSFERASE 5, fucosyltransferase 5 (.1)
AT1G74420 64 / 1e-12
ATFUT3, FUT3
fucosyltransferase 3 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041641
285 / 3e-95
AT1G14100 426 / 5e-144
fucosyltransferase 8 (.1)
Lus10008696
142 / 2e-40
AT2G03220 504 / 3e-174
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Lus10037152
127 / 9e-36
AT2G03220 564 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Lus10029049
124 / 5e-34
AT2G03220 450 / 5e-153
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Lus10036776
122 / 2e-33
AT2G03220 659 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Lus10026128
104 / 7e-27
AT2G03220 395 / 4e-132
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Lus10041642
103 / 3e-26
AT2G03220 454 / 8e-155
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Lus10026125
96 / 2e-24
AT2G03210 305 / 6e-100
fucosyltransferase 2 (.1)
Lus10008693
96 / 2e-24
AT2G03210 305 / 6e-100
fucosyltransferase 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G191401
147 / 6e-42
AT2G03220 555 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Potri.003G191301
140 / 1e-39
AT2G03220 524 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Potri.003G191500
139 / 2e-39
AT2G03220 570 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Potri.001G034100
137 / 6e-39
AT2G03220 560 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Potri.008G090000
129 / 8e-36
AT2G03220 744 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Potri.003G191200
127 / 3e-35
AT2G03220 570 / 0.0
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
Potri.001G033800
127 / 7e-35
AT2G03220 524 / 3e-180
MURUS 2, ARABIDOPSIS THALIANA FUCOSYLTRANSFERASE 1, fucosyltransferase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03254
XG_FTase
Xyloglucan fucosyltransferase
Representative CDS sequence
>Lus10024079 pacid=23141520 polypeptide=Lus10024079 locus=Lus10024079.g ID=Lus10024079.BGIv1.0 annot-version=v1.0
ATGTACTCAAACAAACCAACTGTGACTGGTGAAGTCATCAGGTTTTACCAACCAAGCCATGAGGAGACACAGAAGTTTAAGGACAGTGATCACAACATCA
AGGCATTGGTTGAGATATATCTGCTTAGCATGTGCGAGAAGTTAGTGATCAGCTCGAAGTCAACATTTGGGTATGTAGCTCAGGGTTTAGGGGATACCAA
GGCATGGGTCCTGAGACGGCCGGGAGACGGTGCTTCTGATCCTTGTGTGAGAGACTTGTCACCAGATCCATGCTTTCACTATCCTCCAACGAGGAGCTGT
AATGTTGGTAGTAACAAGAACAACAACAGATATGTTGATGTTGCTGATCATGTTCCCTATATGAAGCATTGCAAGGACCAGAAGATCGGAGTGAAGCTAG
TCGATATTCATAAGGAGTTGTAG
AA sequence
>Lus10024079 pacid=23141520 polypeptide=Lus10024079 locus=Lus10024079.g ID=Lus10024079.BGIv1.0 annot-version=v1.0
MYSNKPTVTGEVIRFYQPSHEETQKFKDSDHNIKALVEIYLLSMCEKLVISSKSTFGYVAQGLGDTKAWVLRRPGDGASDPCVRDLSPDPCFHYPPTRSC
NVGSNKNNNRYVDVADHVPYMKHCKDQKIGVKLVDIHKEL
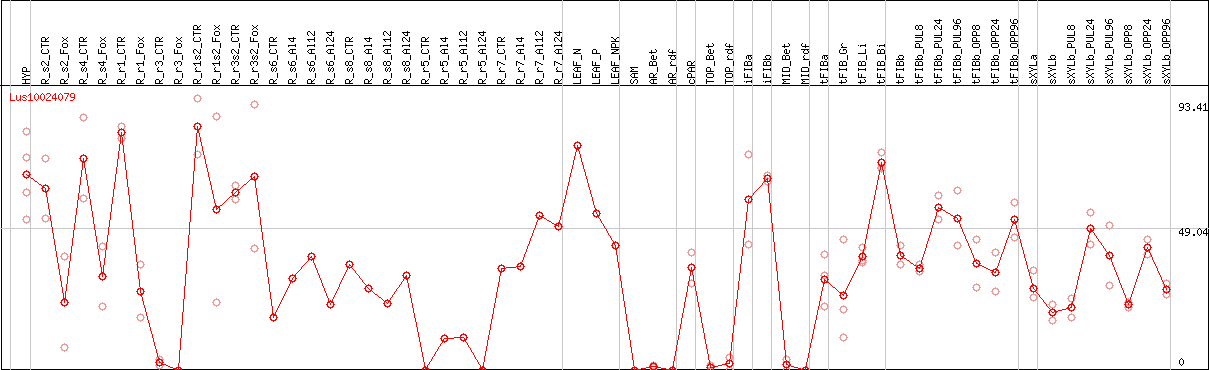
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024079 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.